- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
RNA Degradation Assay Using RNA Exosome Complexes, Affinity-purified from HEK-293 Cells
Published: Vol 7, Iss 8, Apr 20, 2017 DOI: 10.21769/BioProtoc.2239 Views: 9919
Reviewed by: Gal HaimovichAntoine de MorreeVaibhav B Shah

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
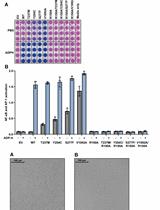

Measurement of the Activity of Wildtype and Disease-Causing ALPK1 Mutants in Transfected Cells With a 96-Well Format NF-κB/AP-1 Reporter Assay
Tom Snelling
Nov 20, 2024 1644 Views
Fluorescence Polarization-Based High-Throughput Screening Assay for Inhibitors Targeting Cathepsin L
Keyu Guo [...] Shuyi Si
Jul 20, 2025 2333 Views
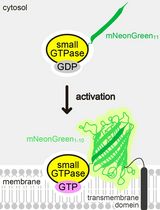
Detecting the Activation of Endogenous Small GTPases via Fluorescent Signals Utilizing a Split mNeonGreen: Small GTPase ActIvitY ANalyzing (SAIYAN) System
Miharu Maeda and Kota Saito
Jan 5, 2026 556 Views
Abstract
The RNA exosome complex plays a central role in RNA processing and regulated turnover. Present both in cytoplasm and nucleus, the exosome functions through associations with ribonucleases and various adapter proteins (reviewed in [Kilchert et al., 2016]). The RNA exosome-associated EXOSC10 protein is a distributive, 3’-5’ exoribonuclease. The following protocol describes an approach to monitor the ribonucleolytic activity of affinity-purified EXOSC10-containing RNA exosomes, originating from HEK-293 cells, as reported in (Domanski et al., 2016) and further detailed in the companion bio-protocol to this one (Domanski and LaCava, 2017).
Keywords: RNA exosomeBackground
In our previous work, we established an isogenic HEK-293 cell line expressing C-terminally 3xFLAG-tagged exosome component EXOSC10 (RRP6), under the control of a tetracycline-inducible CMV promoter (HEK-293 Flp-In T-REx – Thermo Fisher Scientific). This system permitted us to express the tagged EXOSC10 protein at a level comparable to the endogenous WT protein, and to explore exosome purification protocols using a magnetic anti-FLAG affinity medium and protein extracts derived from cryomilled cell powder (Domanski et al., 2012). Building on this, we developed protocols for further purifying RNA exosomes by rate-zonal centrifugation, using glycerol density gradients, and assaying their ribonuclease (RNase) activity (Domanski et al., 2016). EXOSC10-containing exosome fractions exhibited apparent exoribonucleolytic activity, consistent with distributive 3’-5’ hydrolysis; the same assay permitted the detection and monitoring of the processive RNase activity of affinity purified DIS3-3xFLAG ([Wasmuth and Lima, 2012] and references therein). The protocol presented here describes the RNase assay. Although this protocol presumes glycerol gradient purified EXOSC10-3xFLAG-containing exosomes as the point of entry into the assay (Domanski and LaCava, 2017), the method should be applicable to any sufficiently pure and concentrated samples.
Materials and Reagents
Note: Catalog numbers are given for most of the reagents listed below; an equivalent quality reagent from an alternative supplier can typically be substituted with comparable results. Due to the potential for artifacts introduced by contaminating RNases, care should be taken to follow best practices, such as the use of RNase-free solutions and reagents and/or using DEPC-treatment where appropriate (Farrell, 2010). Standard materials and reagents for urea-polyacrylamide gel electrophoresis are required; we use the National Diagnostics system but such gels can be prepared using standard methods (Sambrook and Russell, 2006).
- Pipette tips
- 1.5 ml microcentrifuge tubes (e.g., Eppendorf, catalog number: 022363204 )
- 10-well Gel Combs, 1.5 mm (Thermo Fisher Scientific, NovexTM, catalog number: NC3510 )
- Empty Gel Cassettes, mini, 1.5 mm (Thermo Fisher Scientific, NovexTM, catalog number: NC2015 )
- Syringe with a bent needle (to wash the residual urea out of the gel wells before sample loading)
- HEK-293 Flp-In T-REx EXOSC10-3xFLAG cells ([Domanski et al., 2016]; available upon request) Parental HEK-293 Flp-In T-REx cell line (Thermo Fisher Scientific, InvitrogenTM, catalog number: R78007 )
- Nuclease-free water (Thermo Fisher Scientific, AmbionTM, catalog number: AM9932 )
- Sodium chloride (NaCl) (Sigma-Aldrich, catalog number: S3014 )
- 2-[4-(2-hydroxyethyl)piperazin-1-yl]ethanesulfonic acid (HEPES) (Sigma-Aldrich, catalog number: 54457 )
- Triton X-100 (Sigma-Aldrich, catalog number: T8787 )
- Anti-FLAG M2 (Sigma-Aldrich, catalog number: F3165 ) antibody conjugated Dynabeads M-270 Epoxy (Thermo Fisher Scientific, InvitrogenTM, catalog number: 14302D ) (Domanski and LaCava, 2017)
- Dithiothreitol (DTT) (Sigma-Aldrich, catalog number: 43815 )
- Magnesium chloride (MgCl2) (Sigma-Aldrich, catalog number: M8266 )
- RNA substrate with 6-carboxyfluorescein (6-FAM) at the 5’-end (Reconstitute in 10 mM HEPES ~pH 7-7.5, at 0.5 nmol/µl. Store aliquots at -80 °C)
- Generic substrate: 5’-(6-FAM)-CCUAU UCUAU AGUGU CACCU AAAUG CUAGA GCU modC(2’-O-Me)-3’
- Blocked substrate: 5’-(6-FAM)-CCUAU UCUAU AGUGU CACCU AAAUG CUAGA GCU modC(2’-O-Me, 3’-PO4)-3’
- Generic substrate: 5’-(6-FAM)-CCUAU UCUAU AGUGU CACCU AAAUG CUAGA GCU modC(2’-O-Me)-3’
- RNasin® ribonuclease inhibitors (Promega, catalog number: N2515 )
- Formamide (Sigma-Aldrich, catalog number: 47671 )
- DNA loading dye (Thermo Fisher Scientific, catalog number: R0611 )
Note: This consists of 10 mM Tris-HCl (pH 7.6), 0.03% bromophenol blue, 0.03% xylene cyanol FF, 60% glycerol, 60 mM EDTA–and so, can be prepared rather than purchased. - Ethylenediaminetetraacetic acid (EDTA) (Sigma-Aldrich, catalog number: E6758 )
- Tris base (Sigma-Aldrich, catalog number: 93362 )
- Boric acid (H3BO3) (Sigma-Aldrich, catalog number: B7901 )
- N,N,N’,N’-tetramethylethylenediamine (TEMED) (Sigma-Aldrich, catalog number: T7024 )
- Ammonium persulfate (APS) (Sigma-Aldrich, catalog number: A3678 )
Note: Prepare 10% solution in H2O and store at 4 °C for several weeks. - UreaGel 29:1 Denaturing Gel System (National Diagnostics, catalog number: EC-829 )
- Recapture solution (see Recipes)
- Wash solution (see Recipes)
- 2x reaction solution (see Recipes)
- 2x RNA loading solution (see Recipes)
- 1x TBE (5x or 10x stock can be prepared) (see Recipes)
Equipment
Note: Standard equipment for urea-polyacrylamide gel electrophoresis is required, as well as an imager capable of fluorescein detection (absorption λmax = 494 nm, emission λmax = 518 nm).
- Pipettes
- Vortexer
- Benchtop mini-centrifuge
- Neodymium magnet microfuge tube rack (Thermo Fisher Scientific, catalog number: 12321D )
- Thermomixer (e.g., Eppendorf, model: ThermoMixer® F , catalog number: 5355000.011; or equivalent)
- XCell SureLock Mini (Thermo Fisher Scientific, model: SureLock® Mini-Cell , catalog number: EI0001)
- Electrophoresis power supply
- Imager with blue light (460 nm) epi-illumination and a Y515-Di (long-pass) filter–i.e., SYBR green settings. E.g., Fujifilm LAS-3000 series or newer (Fujifilm, model: LAS-3000 Series ; or equivalent)
Procedure
Note: Purify EXOSC10-containing RNA exosomes as described in the complementary protocol, Affinity purification of RNA exosomes from HEK-293 cells (Domanski and LaCava, 2017), or by other means. This assay may also be applied effectively to affinity purified DIS3-3xFLAG as previously described (Domanski et al., 2016). Examples of the manipulations associated with the affinity capture aspects of the following protocol can be viewed in our online video protocol (LaCava et al., 2016).
- Re-capture of RNA exosomes from the glycerol gradient
- After retrieving the fractions from the gradient, pool together up to two fractions constituting the peak of intact EXOSC10-containing exosomes (up to ~450 μl total volume).
Note: If desired, save an aliquot of up to 50 μl (input) for other analyses. - Add an equal volume of the recapture solution (see Recipe 1).
- Pre-wash 10 µl of the anti-FLAG magnetic medium (slurry) twice with 1 ml of recapture solution.
Note: Combine 1 ml of recapture solution with the beads slurry and briefly vortex. Pulse-spin the tube in a mini-centrifuge to collect all the solution at the bottom and then place the tube on a magnetic tube rack. Wait until beads are collected at the side of the tube and remove the supernatant (using a pipet or an aspirator). Add additional recapture solution to the beads in the tube and repeat the process for the second wash; after removing the supernatant the beads may be held on ice and are ready for use. Both washing steps can be carried out at room temperature. - Transfer diluted fractions from step A2 into a 1.5 ml tube with pre-washed beads.
- Incubate for 30 min with rotation at 4 °C (cold room).
- Place on a magnet, wait until the beads are collected at the side of the tube, and remove the supernatant.
Note: The supernatant may be compared to the input by protein staining or Western blot to monitor the extent of exosome depletion. - Wash once with 1 ml wash solution (see Recipe 2).
- Resuspend the affinity medium in 25 µl wash solution.
- Add 25 µl of the 2x reaction solution (see Recipe 3).
- Incubate at 37 °C with mixing at 1,000 rpm.
- Produce a time-course for the reaction by taking 10 µl aliquots at different time points e.g., 0, 2.5, 5, 15 and 30 min. Stop the reactions by adding 10 µl 2x RNA loading buffer (see Recipe 4).
Note: This assay should ideally include two controls set up as independent reactions: a) as above, but using the blocked substrate, and b) a mock reaction with affinity medium and reaction solution containing the generic substrate, but no RNA exosomes. Both controls should reveal the presence of intact substrate across the time course, indicating that the preparation and solutions are free from interfering RNase contamination. DIS3 is capable of degrading the 3’-PO4 blocked substrate. - Place on a magnet and transfer the supernatant to a fresh tube. Keep on ice.
- After retrieving the fractions from the gradient, pool together up to two fractions constituting the peak of intact EXOSC10-containing exosomes (up to ~450 μl total volume).
- RNA PAGE and gel imaging
- Cast a 20% urea-polyacrylamide 10-well gel accordingly to the manufacturer’s instructions.
- Pre-run the gel in 1x TBE (see Recipe 5) at 15 W for 15-20 min.
Note: The pre-running step clears excess free ions from the gel which affect the electrical current and heat generation. The cassette will warm during the pre-run and run; typical operating temperatures are between 45-60 °C. The heat produced may help maintain the RNA sample in a denatured, single stranded form; this is not expected to be a major variable for small oligo substrates lacking strong secondary structure. - Heat the samples at 80 °C for 30 sec and then place on ice. Spin down briefly before loading.
Note: Heating the sample denatures the RNA and dissociates RNA-protein assemblies. - Wash the urea out from the wells using a syringe with a bent needle.
- Load on the gel the entire volume of each sample (20 µl) per well.
- Run at 15 W until the bromophenol blue tracking dye reaches the first line from the bottom of the plastic gel cassette.
- Open the cassette and image the gel directly using an appropriately configured imager (SYBR green on Fuji LAS series).
Note: The gel does not need to be fixed, washed, or dried prior to imaging. Depending on the level of activity detected, some adjustment of the image brightness/contrast in software, after data collection, may improve visualization.
- Cast a 20% urea-polyacrylamide 10-well gel accordingly to the manufacturer’s instructions.
Data analysis
The distributive, 3’-5’ exoribonucleolytic activity of EXOSC10 can be observed as ‘laddering’ of the substrate degradation intermediates, shifting increasingly over time toward lower molecular mass species; and thus, increasingly toward the bottom of the gel. Such a pattern is depicted in Figure 1. The last step of the ladder (i.e., latest time point) may appear compressed relative to earlier time points because the enzyme may no longer exhibit appreciable activity on a terminally shortened pool of substrates and the rest of the pool converges on this size. A 3’-PO4 blocked substrate will prevent EXOSC10-derived 3’-5’ activity; full length substrate and an absence of breakdown products should be observed across all time points in this case. Note, however, that this expectation presumes the absence of DIS3 or that any DIS3 present in the purified exosome has been inactivated (Domanski et al., 2016; Zinder et al., 2016).
Figure 1. Representative result: RNA degradation assay using affinity-purified human RNA exosome complexes. Schematic diagram of a urea-polyacrylamide gel separation of the reaction products of an exosome RNase assay as described in this protocol, consistent with a distributive activity pattern and the original data. B. The original gel, reproduced from (Domanski et al., 2016): an ExoI preparation was incubated either with a generic or blocked substrate and the indicated time points were collected.
Recipes
- Recapture solution
100 mM NaCl
20 mM HEPES, pH 7.4
1% Triton X-100 - Wash solution
100 mM NaCl
20 mM Tris-HCl, pH 8.0
0.01% Triton X-100 - 2x reaction solution
20 mM Tris-HCl, pH 8.0
20 mM DTT
0.5 mM MgCl2
5% (v/v) RNasin® ribonuclease inhibitors
0.4 pmol/µl RNA substrate - 2x RNA loading solution
95% formamide
1% DNA loading dye
20 mM EDTA - 1x TBE (5x or 10x stock can be prepared)
89 mM Tris base
89 mM boric acid
2 mM EDTA
Acknowledgments
We thank Professors Michael P. Rout and Torben Heick Jensen for their invaluable support of our research. We also thank Mr. Artem Serganov and Dr. Zhanna Hakhverdyan for proof reading. This work was supported in part by the National Institutes of Health grants P41GM109824 and P50GM107632, the Lundbeck Foundation, and the Danish National Research Foundation.
References
- Domanski, M. and LaCava, J. (2017). Affinity purification of RNA exosomes from HEK-293 cells. Bio-Protoc 7(08): e2238.
- Domanski, M., Molloy, K., Jiang, H., Chait, B. T., Rout, M. P., Jensen, T. H. and LaCava, J. (2012). Improved methodology for the affinity isolation of human protein complexes expressed at near endogenous levels. Biotechniques 0(0): 1-6.
- Domanski, M., Upla, P., Rice, W. J., Molloy, K. R., Ketaren, N. E., Stokes, D. L., Jensen, T. H., Rout, M. P. and LaCava, J. (2016). Purification and analysis of endogenous human RNA exosome complexes. RNA 22(9): 1467-1475.
- Farrell, R. E. (2010). RNA Methodologies. 4th edition.
- Kilchert, C., Wittmann, S. and Vasiljeva, L. (2016). The regulation and functions of the nuclear RNA exosome complex. Nat Rev Mol Cell Biol 17(4): 227-239.
- LaCava, J., Jiang, H. and Rout, M. P. (2016). Protein complex affinity capture from cryomilled mammalian cells. J Vis Exp e54518.
- Sambrook, J. and Russell, D. W. (2006). The Condensed Protocols. 1st edition. Cold Spring Harbor Laboratory Press.
- Wasmuth, E. V. and Lima, C. D. (2012). Structure and activities of the eukaryotic RNA exosome. Enzymes 31: 53-75.
- Zinder, J. C., Wasmuth, E. V. and Lima, C. D. (2016). Nuclear RNA exosome at 3.1 A reveals substrate specificities, RNA paths, and allosteric inhibition of Rrp44/Dis3. Mol Cell 64(4): 734-745.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Domanski, M. and LaCava, J. (2017). RNA Degradation Assay Using RNA Exosome Complexes, Affinity-purified from HEK-293 Cells. Bio-protocol 7(8): e2239. DOI: 10.21769/BioProtoc.2239.
Category
Molecular Biology > RNA > RNA degradation
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











