- Protocols
- Articles and Issues
- About
- Become a Reviewer
Articles In Press
Articles In Press are peer reviewed and have been accepted for publication. Please note that these versions may be subject to further edits before their final online publication. Nevertheless, Articles In Press are citable using the DOI. Upon the formal online publication, the article will no longer be listed here, but existing links will automatically redirect to the final version in the corresponding issue.
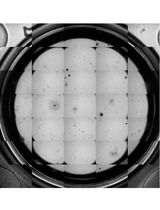
A Reliable Method for Thawing Primary AML and CMML Mononuclear Cells to Preserve Viability and Function
Human mononuclear cells derived from peripheral blood and bone marrow are valuable resources for the study of hematological malignancies, including acute myeloid leukemia (AML) and chronic myelomonocytic leukemia (CMML). Cryopreservation enables long-term storage of patient samples for downstream assays; while thawing protocols have been described, subsequent recovery of viable cells after thawing can be challenging, particularly for fragile blast and monocyte populations. Here, we describe a reliable protocol for thawing cryopreserved AML and CMML mononuclear cells designed to preserve post-thaw viability, recovery, and functional integrity. The method incorporates controlled dilution of cells out of cryoprotectant with anticoagulant-supplemented thaw buffer, DNase I treatment, and gentle resuspension steps. Using this approach, post-thaw viability consistently exceeded 80% with a mean recovery of 55.6% across samples. Recovered cells retained functional capacity, as demonstrated by colony-forming assays, and maintained immunophenotypic characteristics by flow cytometry. This protocol provides a robust and reproducible method for the recovery of cryopreserved AML and CMML mononuclear cells and may be broadly applicable to other fragile or monocyte-rich patient-derived hematopoietic samples.
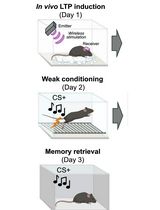
Optogenetic LTP Manipulation and Mathematical Modeling to Investigate Value Plasticity of the Instructive Signal in Mice
Adaptive behaviors shaped by prior experience are essential for increasing animal survival. Aversive experiences play a pivotal role in memory formation and in updating subsequent learning rules. While the negative value of aversive signals, which are both necessary and sufficient to drive a conditioned response, is considered to be innately specified, it can also be subject to experience-dependent scaling. Previous reports demonstrated synaptic potentiation in nociceptive pathways following robust aversive learning. However, the neuronal basis of experience-dependent value updating remains largely unknown. Recently, we demonstrated that long-term potentiation (LTP) in the parabrachial-central amygdala (PB-CeA) pathway, an important circuit involved in pain processing and aversive learning, enhances the negative value and thereby updates future learning rules. Here, we present a protocol that combines behavioral analysis using pathway-specific optogenetic induction of in vivo LTP with mathematical modeling to examine value modification using Bayesian inference of the unconditioned stimulus value using the Rescorla–Wagner model. This protocol enables investigation of the mechanisms underlying experience-dependent value modulation and learning-rule changes in mice. Potentially, this protocol may provide a framework for understanding learning rules across a wide range of species and for the development of treatments for stress-related disorders.
Construction and Functional Evaluation of Cyclic Peptide-Based CAR T Cells in Tumor Models
Cyclic peptides are emerging as a promising class of recognition modules for chimeric antigen receptor (CAR) engineering. Compared with single-chain variable fragment (scFv)-based CARs, disulfide-directed multicyclic peptides (DDMPs) represent a novel alternative, offering a markedly smaller molecular size (<5 kDa), enhanced structural stability through disulfide-directed cyclization, and broad tolerance to sequence diversification that supports systematic affinity and specificity optimization. DDMP-based CAR T cells leverage these properties to mediate antigen-dependent cytotoxicity while exhibiting an attenuated cytokine secretion profile, supporting the development of potentially safer immunotherapies for solid tumors. Here, we present a comprehensive workflow spanning CAR construct design and generation through in vitro and in vivo functional evaluation. While DDMPs are used as the exemplar recognition module, sections A and C–L of the protocol are directly applicable to any CAR format, including scFv- and nanobody-based designs with minimal modifications, making the workflow accessible to the broader CAR T-cell research community. The protocol includes the generation of Jurkat NFAT reporter cell lines and luciferase-expressing tumor target lines, which are widely used in different assays. Together, these standardized readouts enable rigorous, objective comparison of CAR T-cell efficacy and safety across tumor models.
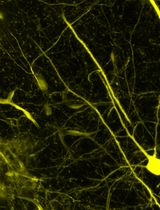
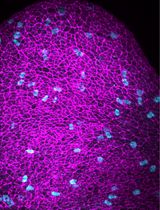
Detection of Target Molecules Within One-Millimeter-Thick Mouse Brain Slices by Using Peroxidase-Fused Nanobodies and Fluorochromized Tyramide-Glucose Oxidase Reaction
Three-dimensional immunohistochemistry (3D-IHC) shows the organization of molecular assemblies in the context of tissue architecture. Deep and rapid antibody penetration into 3D tissues and highly sensitive detection are crucial for high-throughput analysis of 3D-IHC imaging. Here, we provide a detailed protocol for a nanobody (nAb)-based 3D-IHC technique, namely POD-nAb/FT-GO 3D-IHC, for high-speed and high-sensitivity detection of targets within 1-mm-thick mouse brain tissues. Peroxidase-fused nAb (POD-nAb) is a genetically encoded recombinant antibody, which consists of a camelid nAb and a variant of horseradish peroxidase, and fluorochromized tyramide-glucose oxidase (FT-GO) is a fluorescent tyramide signal amplification (TSA) system. POD-nAb/FT-GO 3D-IHC incorporates three main components: 1) tissue permeabilization, 2) POD-nAb binding, and 3) 3D-TSA reaction with FT-GO. POD-nAbs enhance signal penetration depth and allow for highly sensitive detection when combined with FT-GO signal amplification. By using the 3D-IHC protocol provided herein, we can visualize target molecules in mouse brain tissues of 1-mm thickness with drastic signal enhancement within three days. This protocol for POD-nAb/FT-GO 3D-IHC could facilitate structural and molecular interrogation of 3D tissues.
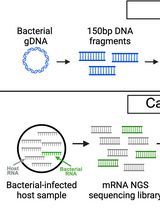
Enriching Bacteria-Specific RNA From Host Samples Before NGS With Transcript-Capture
Pathogen gene expression from host samples is often challenging to study due to low signal and high host RNA background. PCR probes have been recently used to hybridize and extract bacterial sequences from next-generation sequencing (NGS) libraries generated from in vitro and animal models of infection; however, these strategies require purchasing commercially synthesized probes that often do not capture the entire transcriptome. Transcript-capture sequencing is a novel capture approach for extracting RNA of a target bacterial species from samples in which there is substantial contamination by the host or other microbes. Biotinylated 150-base-pair DNA probes are generated in-house from bacterial DNA spanning the entire bacterial genome. Probes are hybridized to the cDNA of NGS sequencing libraries prepared from host samples to capture and enrich for bacterial-specific RNA reads before sequencing. This method results in a >200-fold increase in bacterial RNA reads from infected host samples (including in vitro, animal, and human samples) and generates complete bacterial transcriptomes with high gene coverage (>80%). Use of this protocol on infected host samples reveals a snapshot of bacterial activity during disease that may improve understanding of the physiological state of pathogens within their hosts.
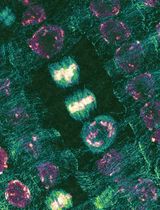
Quantitative Analysis of Cell and Tissue Shape During Mouse Cranial Neural Tube Closure
Neural tube closure is a critical process that transforms the neural plate, an open epithelial tissue, into the closed tube that serves as the structural basis of the central nervous system. Defects in this process are among the most common and severe developmental diseases in the human population, with failures in cranial closure accounting for approximately one-third of total defects. However, the cell and tissue mechanisms that drive cranial closure remain opaque relative to the better studied process of spinal closure, in large part due to the unique challenges in characterizing cranial tissues. Here, we present protocols for quantifying cell dynamics and tissue-level remodeling events that enable highly spatiotemporally resolved investigations of the causes of cranial closure defects in mouse embryos. These include brightfield morphometric approaches, fluorescent staining and confocal imaging, and quantitative pipelines to analyze these image-based datasets. At the conclusion of these approaches, users will be able to quantify several parameters of overall tissue shape in the cranial neural tissues and produce rich quantitative datasets about cell-level parameters, particularly apical cell area. These can be used to identify correlative and causative differences between mutants and control embryos. Given their flexibility, many of these approaches can be generalized to other tissue morphogenetic contexts.
Customizable High-Throughput Chemical Phenotyping of Root Bacteria
Chemical phenotyping is a fundamental technique to study the metabolic properties or chemical sensitivities of bacteria. Traditional methods such as dilution methods, discs, or gradient diffusion assays are labor-intensive, often have high material requirements, and are limited in scalability. High-throughput cultivation approaches based on 96-well plates scale efficiently to large numbers of samples. A stacker, when coupled with a plate reader system (often already available in most laboratories), greatly enhances assay scalability and robustness. Here, we describe a customized high-throughput, flexible, scalable, robust, and affordable method for the chemical phenotyping of bacteria. This liquid culture–based growth system allows screening many bacteria in parallel and in a replicated manner for their tolerance to various chemicals, including specialized metabolites of plants, antibiotics, or pesticides. Compared to commercial solutions, our approach offers high flexibility in experimental conditions while keeping costs for consumables low.
Versatile Dual Mounting Enables Larval Zebrafish Imaging Across Microscope Configurations
Larval zebrafish are often mounted laterally to ensure consistent anatomical positioning and to standardize imaging of body axes across early development. However, this conventional approach often tethers sample orientation to a single microscope configuration and limits optical accessibility. We present a mounting protocol for larval zebrafish that enables optical access from both dorsal and ventral orientations while preserving lateral sample position. This approach uses common laboratory consumables to establish a mounting platform that eliminates any need to remount samples between the use of upright and inverted microscopes. By establishing a hydrophobic seal, mounted embryos can be inverted with ease to access the sample from either orientation. A seamless transition here facilitates reliable identification and longitudinal tracking of the same biological region of interest across microscope configurations. This protocol is broadly applicable to live imaging experiments requiring flexibility in imaging geometry, minimal sample handling, and high reproducibility.
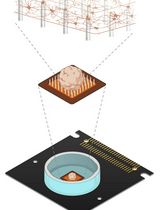
Measuring Electrophysiological Activity in Acute Brain Slices, Spheroids, and Organoids Using 3D High-Density Multielectrode Arrays
Animal and human stem cell–derived three-dimensional models to study physio-pathological brain functioning are becoming a gold standard for in vitro electrophysiology, as they enable the recapitulation of complex network properties by accounting for spatial architectural features that better reflect in vivo conditions than simpler 2D models. Standard planar multielectrode arrays (MEAs), typically providing tens of recording electrodes, are commonly used to record activity from 2D neuronal cultures. However, when adapted for use with 3D models, planar 2D MEAs showed limited effectiveness. The main issues are limited specimen adhesion to the chip, a low number of sensing elements, inability to retrieve signals from within the tissue, and reduced perfusion and vitality of the tissue in contact with sensors. To overcome these limitations, a new generation of microchip-based 3D high-density MEAs (3D HD-MEA) has been developed and validated in recent years. This technological advancement has improved the sensing capabilities and the vitality of 3D models, providing a tool tailored to maximize their potential. Here, we present an optimized protocol for neural network activity recordings in 3D models (including acute slices, brain spheroids, and organoids) from various brain regions using 3D HD-MEAs. First, we summarize the critical steps for 1) obtaining viable acute slices from the mouse cerebellum, cortico-hippocampal circuit, and prefrontal cortex, 2) establishing efficient coupling of the slices with the chip, and 3) performing recordings and analyses. We then describe the main procedures required to obtain human and animal brain spheroids and neural organoids, as well as standardized routines to perform effective recordings and analyses. For each section, we highlight the crucial steps, identify tips for specific applications, and propose troubleshooting procedures. For example, the same type of preparation (e.g., acute slices) requires different adjustments when working with different brain areas. The specific information provided here is intended to assist researchers in their daily efforts to obtain efficient and reproducible functional recordings from 3D models by using the cutting-edge technique of 3D HD-MEA.
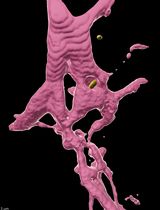
ROOT-ExM: Super-Resolution Imaging of Proteins in Arabidopsis Roots by Expansion Microscopy
Conventional light microscopy is limited in resolution by the diffraction limit of light, restricting the visualization of the nanoscale organization of biomolecules. Expansion microscopy (ExM) has emerged as a powerful technique to overcome this barrier by physically expanding the specimen embedded in a swellable hydrogel without requiring specialized or high-cost imaging hardware. ExM is widely used in animal models, whereas its application to plant tissues has been challenging due to their multicellularity, in which each cell is encompassed by the rigid cell wall, which resists the expansion forces and prevents isotropic swelling. Here, we describe a robust and optimized ExM protocol specifically designed for Arabidopsis thaliana root tissues. This protocol details critical steps, including immunostaining, anchoring, gelation, denaturation, cell wall digestion, and expansion. Our method achieves an expansion factor of approximately 4.3×, enabling an effective lateral resolution of ~60 nm using a standard confocal microscope. We demonstrate the visualization of microtubules with preserved ultrastructure. This accessible protocol allows plant researchers to perform super-resolution imaging without specialized optical equipment, facilitating detailed structural analysis of plant cells.
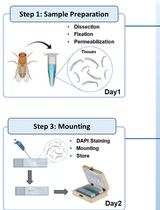
Oligo(dT) Fluorescence In Situ Hybridization to Visualize the Poly(A) mRNAs in the Internal Tissues of Drosophila
Fluorescence in situ hybridization (FISH) is a cytological method used to visualize specific oligonucleotide sequences within the cell. This method relies on the specific binding of a fluorescence-tagged probe, a short stretch of single-stranded polynucleotide, to its complementary sequence in the DNA or RNA, forming stable double-stranded hybrids. Fluorochromes, such as fluorescein, Alexa Fluor, cyanine dyes, or rhodamine, are attached to these probes to help in detecting their presence within the cell. Based on sequence complementarity, FISH allows for the visualization of the DNA or RNA with which they have hybridized. The distribution of these fluorochrome-tagged probes can be observed under a fluorescence or confocal microscope. The oligo(dT) FISH technique specifically utilizes a fluorochrome-tagged stretch of 40–50 thymidine (T) oligonucleotides that binds to the poly(A) tails of mature mRNAs within the cell. Newly transcribing pre-mRNAs and certain non-coding RNAs may not have poly(A) tails and therefore cannot be detected by this method. This step-by-step protocol outlines the oligo(dT) FISH technique for visualizing the cellular distribution of polyadenylated mRNAs in the tissues of Drosophila and other related model organisms.
Comprehensive Protocol for Handling Human Small Airway Epithelial Cells (HSAECs) to Establish Air–Liquid Interface (ALI) Cultures With TEER-Based Barrier Integrity Assessment
Understanding epithelial barrier function is essential for studying both its normal physiology and its role in disease, yet choosing an appropriate experimental model remains challenging. Animal models are commonly used but often suffer from interspecies differences that limit translational relevance. Human-derived cell lines offer a more suitable alternative, although establishing them often requires immortalisation strategies that involve overexpression of oncogenes, which can introduce phenotypic and functional changes. In contrast, primary cells, such as human small airway epithelial cells (HSAECs), provide a more physiologically accurate model. A critical aspect of replicating the native respiratory environment is maintaining continuous air exposure, which can be achieved through air–liquid interface (ALI) culture. This protocol provides a unified, step-by-step workflow for cultivating primary HSAECs under ALI conditions, covering the entire process from initial recovery after cryopreservation to the formation of a barrier-like layer. The protocol incorporates non-invasive methods such as transepithelial electrical resistance (TEER) measurements to monitor its integrity. While individual elements of this workflow have been described separately in different studies, a consolidated version encompassing the full workflow has not been widely available. This resource is intended for researchers with limited experience in airway epithelial culture and offers practical, clear guidance through each step of the process.
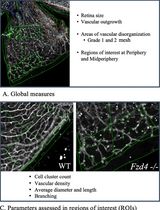
Computational Quantification of Mouse Retinal Vasculature Using ImageJ
Postnatal mouse retinal vascular development is a widely used model for studying retinal vascular diseases and evaluating candidate therapies. This is particularly relevant for inherited disorders such as familial exudative vitreoretinopathy (FEVR), in which impaired vascular growth and organization are central to disease pathogenesis. Numerous approaches have been used to assess retinal vasculature in mouse flat mounts, ranging from qualitative descriptions to limited quantitative measurements of vascular growth. However, phenotypic variability across genetic models, including different models of FEVR, complicates comparisons and underscores the need for standardized, comprehensive multi-parameter analyses that are suitable for rapid and cost-effective screening studies. We describe a standardized morphometric protocol using ImageJ software to quantitatively analyze mouse retinal vasculature in a reproducible manner. The protocol begins with measurement of areas of vascular disorganization (meshes) as well as total vascular and retinal area. Two defined regions in the peripheral and midperipheral retina are then selected to quantify cell clusters, followed by image processing, binarization, and skeletonization. From these processed images, vascular density, branch number, branch length and thickness, junction number, triple points, and box-counting fractal dimension and lacunarity are quantified. Overall, this protocol provides a rapid, cost-effective, and standardized framework for quantifying retinal vascular phenotypes across diverse mouse models. By capturing multiple structural features and accommodating phenotypic variability, it is well-suited for comparative studies and therapeutic screening in retinal vascular disease.
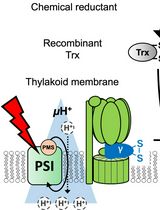
Evaluating Thioredoxin-Mediated CFoCF1 Reduction Using an In Vitro Thylakoid Assay
The activity of chloroplast ATP synthase (CFoCF1) is precisely regulated through a thioredoxin (Trx)-mediated dithiol/disulfide reaction in response to varying light conditions. This regulatory mechanism is further controlled by ΔpH formation across the thylakoid membrane. To better understand this complicating regulatory function of CFoCF1, a method is required to evaluate the extent of CFoCF1 reduction by Trx under controlled ΔpH conditions and to directly evaluate the redox state of CFoCF1. In this study, we present a simple in vitro procedure to assess the CFoCF1 reduction system using spinach thylakoids. The method consists of three key steps: (A) simple preparation of intact thylakoids from spinach leaves; (B) reduction of CFoCF1 on the thylakoid membrane using recombinant Trx under light irradiation; and (C) in situ determination of the redox state of CFoCF1 by labeling thiol groups with a maleimide reagent followed by protein detection using western blotting. The redox state of CFoCF1 was determined by mobility shifts on non-reducing SDS-PAGE. This protocol provides a refined strategy for elucidating the regulatory mechanism controlling energy conversion by CFoCF1 under fluctuating photosynthetic conditions.
One-Step Affinity Purification of MarathonRT Reverse Transcriptase for RNA Sequencing Applications
Transfer RNAs (tRNAs) are important regulators of translation and cellular function. Several high-throughput sequencing methods have been developed to quantitatively analyze tRNA isoacceptors in cells. However, the strong secondary structures and extensive post-transcriptional modification of most tRNA molecules present significant challenges for many reverse transcriptases, negatively impacting sequencing library preparation and causing quantification biases. Currently, the field utilizes processive next-generation reverse transcriptases (ngRTs), such as Induro (New England Biolabs) and UltraMarathonRT (RNAConnect), to address these issues. Despite being used in multiple protocols, these commercial products face little competition and remain costly. However, non-commercial alternatives, such as the original MarathonRT (MRT), are available from gene repositories. MRT is a next-generation reverse transcriptase derived from the Eubacterium rectale group II intron maturase, which can read through RNA secondary structures and chemical modifications. Here, we present a simplified expression and purification protocol for producing highly active MRT that is stable over 1 year. This cost-effective protocol yields a heterogeneous protein preparation with no discernible competing enzymatic activities; it mitigates previously reported precipitation issues, saving one day of laboratory work and eliminating two chromatography-based purification steps. Moreover, the use of the resulting protein preparation has been verified in the mim-tRNAseq pipeline, where it was shown to perform equally to the commercial alternatives Induro and UltraMarathonRT. In addition, we have developed a simple and cost-effective assay for measuring the enzymatic activity of MRT, allowing for batch comparison.
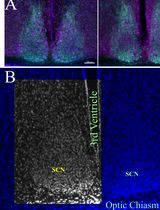
Histomorphometrical Analyses of the Mouse Suprachiasmatic Nucleus
The mammalian central circadian clock resides in the suprachiasmatic nucleus (SCN) of the hypothalamus in the brain and is responsible for coordinating daily rhythms of biological processes spanning from gene expression to behavior. Light, the primary environmental zeitgeber, entrains the SCN via melanopsin-expressing intrinsically photosensitive retinal ganglion cells that project through the retino-hypothalamic tract. Altered circadian rhythms are common in individuals diagnosed with neurodevelopmental and neurodegenerative disorders, and often, associated with structural alterations of the SCN and impaired retinal input; importantly, these anomalies can be recapitulated in animal models. Here, we describe step-by-step protocols for quantitative histomorphometrical analysis of the SCN and the assessment of retinal–SCN connectivity, previously used in mouse models of neurodevelopmental and neurodegenerative disorders. These include measurement of the SCN area, perimeter, height and width using Nissl- or DAPI-stained coronal sections, as well as densitometric and plot profile analyses of cholera toxin β-subunit–labeled retinal projections using Axiovision or Fiji/ImageJ. The protocols incorporate standardized region-of-interest, measurements by masked observers, and consistent scaling procedures to enhance reproducibility. These methods provide a rigorous framework for detecting structural anomalies and connectivity defects in the circadian system and can be broadly applied to other experimental models of circadian dysfunction.
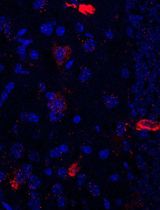
Simultaneous Immunofluorescence-Based In Situ mRNA Expression and Protein Detection in Bone Marrow Biopsy Samples
Fluorescence in situ hybridization (FISH) can be employed to study the expression and subcellular localization of nucleic acids by using labeled antisense strands that hybridize with the target RNA or DNA molecules. Likewise, immunofluorescence antibody staining (IF) takes advantage of the specific interaction between a fluorophore-labeled antibody and its corresponding antigen. This protocol reports the combination of RNA-FISH and IF antibody staining for simultaneous detection of both RNA transcripts and proteins of interest in routine formalin-fixed paraffin-embedded (FFPE) bone marrow biopsy samples. Herein, we provide a detailed description of the methodology that we have developed and optimized to study the spatial expression of two transcripts—TGFB1 and PDGFA1—in human hematopoietic (CD45+) and non-hematopoietic (CD271+) cells in the bone marrow of patients with acute lymphoblastic leukemia (ALL).
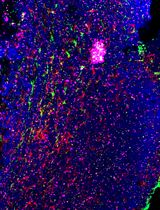
Using combined fluorescent in situ hybridization with Immunohistochemistry to co-localize mRNA in diverse neuronal cell types
Understanding gene expression within defined neuronal populations is essential for dissecting the cellular and molecular diversity of the brain. mRNA assays provide a direct readout of gene expression, capturing transcriptional changes that may precede or occur independently of protein abundance, whereas protein assays reflect the cumulative effects of translation, modification, and degradation. Moreover, in histological analysis, immunohistochemical protein detection results in visually diffuse labeling, which makes it difficult to quantitatively assess levels and locations of expression at high resolution. Here, we present a protocol that allows for mRNA detection in single neuronal cell types with a high degree of sensitivity and anatomical resolution. This protocol combines fluorescent in situ hybridization (FISH) with immunohistochemistry (IHC) on the same tissue section. Briefly, FISH is carried out by ACDBio RNAscope® fluorescent in situ hybridization technology, which involves processing the tissue sections, followed by signal amplification. This involves target retrieval, probe hybridization, and signal enhancement. Then, the tissue section is processed for IHC, which involves blocking nonspecific sites and incubation with primary antibodies, followed by development of a fluorescent signal with secondary antibodies. Typically, visual mRNA detection with FISH can be seen as individual puncta, whereas targeting the protein with an antibody results in filled cells or processes. The variation in staining pattern allows for the quantification of distinct mRNA transcripts within different neuronal populations, which renders co-localization analyses easy and efficient.
Electrophoretic Mobility Shift Assay (EMSA) for Assessing RNA–Protein Binding and Complex Formation Using Recombinant RNA-Binding Proteins and In Vitro–Transcribed RNA
Evaluating RNA–protein interactions is key to understanding post-transcriptional gene regulation. Electrophoretic mobility shift assays (EMSAs) remain a widely used technique to study these interactions, revealing information about binding affinities and binding modalities, including cooperativity and complex formation. Here, we detail, in a step-by-step protocol, how to perform EMSAs. We describe how to generate, purify, and quantitate 32P-radiolabeled RNA by in vitro transcription, as well as the expression and purification of recombinant RNA-binding proteins in E. coli using ELAV as an example. We then describe how to set up binding reactions using serial dilutions in a microtiter plate format of recombinant ELAV and in vitro–transcribed RNA and how to perform EMSAs using native low-crosslinked acrylamide gels, with detailed graphically supported instructions and troubleshooting guides.
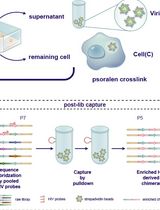
High-resolution mapping of RNA-RNA interactions across the HIV-1 genome with HicapR
The genomes of RNA viruses can fold into dynamic structures that regulate their own infection and immune evasion processes. Proximity ligation methods (e.g., SPLASH) enable genome-wide interaction mapping but lack specificity when dealing with low-abundance targets in complex samples. Here, we describe HiCapR, a protocol integrating in vivo psoralen crosslinking, RNA fragmentation, proximity ligation, and hybridization capture to specifically enrich viral RNA–RNA interactions. Captured libraries are sequenced, and chimeric reads are analyzed via a customized computational pipeline to generate constrained secondary structures. HiCapR generates high-resolution RNA interaction maps for viral genomes. We applied it to resolve the in vivo structure of the complete HIV-1 RNA genome, identifying functional domains, homodimers, and long-range interactions. The protocol's robustness has been previously validated on the SARS-CoV-2 genome. HiCapR combines proximity ligation with targeted enrichment, providing an efficient and specific tool for studying RNA architecture in viruses, with broad applications in virology and antiviral development.
Enhanced RNA-Seq Expression Profiling and Functional Enrichment in Non-model Organisms Using Custom Annotations
Functional enrichment analysis is essential for understanding the biological significance of differentially expressed genes. Commonly used tools such as g:Profiler, DAVID, and GOrilla are effective when applied to well-annotated model organisms. However, for non-model organisms, particularly for bacteria and other microorganisms, curated functional annotations are often scarce. In such cases, researchers often rely on homology-based approaches, using tools like BLAST to transfer annotations from closely related species. Although this strategy can yield some insights, it often introduces annotation errors and overlooks unique species-specific functions. To address this limitation, we present a user-friendly and adaptable method for creating custom annotation R packages using genomic data retrieved from NCBI. These packages can be directly imported as libraries into the R environment and are compatible with the clusterProfiler package, enabling effective gene ontology and pathway enrichment analysis. We demonstrate this approach by constructing an R annotation package for Mycobacterium tuberculosis H37Rv, as an example. The annotation package is then utilized to analyze differentially expressed genes from a subset of RNA-seq dataset (GSE292409), which investigates the transcriptional response of M. tuberculosis H37Rv to rifampicin treatment. The chosen dataset includes six samples, with three serving as untreated controls and three exposed to rifampicin for 1 h. Further, enrichment analysis was performed on genes to demonstrate changes in response to the treatment. This workflow provides a reliable and scalable solution for functional enrichment analysis in organisms with limited annotation resources. It also enhances the accuracy and biological relevance of gene expression interpretation in microbial genomics research.
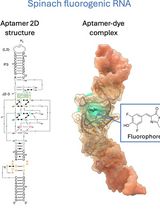
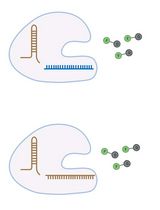
Visualizing diverse RNA functions in living cells with Spinach™ family of fluorogenic aptamers
RNA is now recognized as a highly diverse and dynamic class of molecules whose localization, processing, and turnover are central to cell function and disease. Live-cell RNA imaging is therefore essential for linking RNA behavior to mechanism. Existing approaches include quenched hybridization probes that directly target endogenous transcripts but face delivery and sequestration issues, protein-recruitment tags such as MS2/PP7 that add large payloads and can perturb localization or decay, and CRISPR–dCas13 imaging that requires substantial protein cargo and careful control of background and off-target effects. Here, we present a protocol for live-cell RNA imaging using the SpinachTM family of fluorogenic RNA aptamers. The method details the design and cloning of SpinachTM-tagged RNA constructs, selection and handling of cognate small-molecule fluorophores, expression in mammalian cell lines, dye loading, and image acquisition on standard fluorescence microscopes, followed by quantitative analysis of localization and dynamics. We include controls to verify aptamer expression and signal specificity, guidance for multiplexing with related variants (e.g., Broccoli, Corn, Squash, Beetroot), and troubleshooting for dye permeability and signal optimization. Application examples illustrate use in tracking cellular delivery of mRNA therapeutics, monitoring transcription and decay in response to perturbations, and the forming of toxic RNA aggregates. Compared with prior methods, SpinachTM tags are compact, genetically encodable, and fluorogenic, providing high-contrast imaging in both the nucleus and cytoplasm with single-vector simplicity and multiplexing capability. The protocol standardizes key steps to improve robustness and reproducibility across cell types and laboratories.
Amplification-Free Detection of Highly Structured RNA Molecules Using SCas12aV2
The CRISPR/Cas12a system has revolutionized molecular diagnostics; however, conventional Cas12a-based methods for RNA detection typically require transcription and pre-amplification steps. Our group has recently developed a diagnostic technique known as the SCas12a assay, which combines Cas12a with a split crRNA, achieving amplification-free detection of miRNA. However, this method still encounters challenges in accurately quantifying long RNA molecules with complex secondary structures. Here, we report an enhanced version termed SCas12aV2 (split-crRNA Cas12a version 2 system), which enables direct detection of RNA molecules without sequence limitation while demonstrating high specificity in single-nucleotide polymorphism (SNP) applications. We describe the general procedure for preparing the SCas12a system and its application in detecting RNA targets from clinical samples.
Enhancement of RNA Imaging Platforms by the Use of Peptide Nucleic Acid-Based Linkers
RNA imaging techniques enable researchers to monitor RNA localization, dynamics, and regulation in live or fixed cells. While the MS2-MCP system—comprising the MS2 RNA hairpin and its binding partner, the MS2 coat protein (MCP)—remains the most widely used approach, it relies on a tag containing multiple fluorescent proteins and has several limitations, including the potential to perturb RNA function due to the tag’s large mass. Alternative methods using small-molecule binding aptamers have been developed to address these challenges. This protocol describes the synthesis and characterization of RNA-targeting probes incorporating a peptide nucleic acid (PNA)-based linker within the cobalamin (Cbl)-based probe of the Riboglow platform. Characterization in vitro involves a fluorescence turn-on assay to determine binding affinity (KD) and selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE) footprinting analysis to assess RNA-probe interactions at a single nucleotide resolution. To show the advancement of PNA probes in live cells, we present a detailed approach to perform both stress granule (SG) and U-body assays. By combining sequence-specific hybridization with structure-based recognition, our approach enhances probe affinity and specificity while minimizing disruption to native RNA behavior, offering a robust alternative to protein-based RNA imaging systems.
Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics