- Protocols
- Articles and Issues
- About
- Become a Reviewer
Past Issue in 2024
Volume: 14, Issue: 16
Biological Engineering
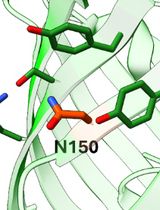
Tetrazine Amino Acid Encoding for Rapid and Complete Protein Bioconjugation
Cell Biology
Calibrating Fluorescence Microscopy With 3D-Speckler (3D Fluorescence Speckle Analyzer)
Developmental Biology
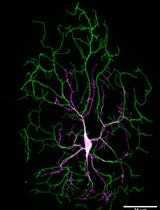
Protocol for Imaging the Same Class IV Neurons at Different Stages of Development
Microbiology
Extraction of Bacterial Membrane Vesicle and Phage Complex by Density Gradient Ultracentrifugation
Molecular Biology
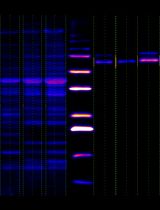
Simple Analysis of Gel Images With IOCBIO Gel Software
Neuroscience
Using Localization Microscopy to Quantify Calcium Channels at Presynaptic Boutons
Plant Science
In Vitro Hyphal Branching Assay Using Rhizophagus irregularis
Systems Biology
Chloroform/Methanol Protein Extraction and In-solution Trypsin Digestion Protocol for Bottom-up Proteomics Analysis
Correction