- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
EMSA Analysis of DNA Binding By Rgg Proteins
Published: Vol 3, Iss 15, Aug 5, 2013 DOI: 10.21769/BioProtoc.838 Views: 15253
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
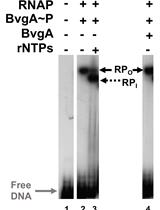
Combining Gel Retardation and Footprinting to Determine Protein-DNA Interactions of Specific and/or Less Stable Complexes
Meng-Lun Hsieh [...] Deborah M. Hinton
Dec 5, 2020 3593 Views
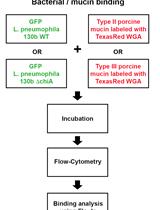
Assay for Assessing Mucin Binding to Bacteria and Bacterial Proteins
Lubov S. Grigoryeva [...] Nicholas P. Cianciotto
Mar 5, 2021 4775 Views
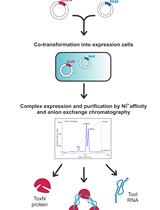
Large-scale Purification of Type III Toxin-antitoxin Ribonucleoprotein Complex and its Components from Escherichia coli for Biophysical Studies
Parthasarathy Manikandan [...] Mahavir Singh
Jul 5, 2023 2175 Views
Abstract
In bacteria, interaction of various proteins with DNA is essential for the regulation of specific target gene expression. Electrophoretic mobility shift assay (EMSA) is an in vitro approach allowing for the visualization of these protein-DNA interactions. Rgg proteins comprise a family of transcriptional regulators widespread among the Firmicutes. Some of these proteins function independently to regulate target gene expression, while others have now been demonstrated to function as effectors of cell-to-cell communication, having regulatory activitiesthat that are modulated via direct interaction with small signaling peptides. EMSA analysis can be used to assess DNA binding of either type of Rgg protein. EMSA analysis of Rgg protein activity has facilitated in vitro confirmation of regulatory targets, identification of precise DNA binding sites via DNA probe mutagenesis, and characterization of the mechanism by which some cognate signaling peptides modulate Rgg protein function (e.g. interruption of DNA-binding in some cases).
Materials and Reagents
- General chemicals
- Non-specific DNA
e.g. Poly-dIdC (Sigma-Aldrich, catalog number: P4929 )
Sheared salmon sperm DNA (Amresco, catalog number: E213 ) - BSA (10 mg/ml)
- Polyacrylamide-bis (40%) for PAGE (e.g. Bio-Rad Laboratories, catalog number: 161-0146 )
- Fluorescently-labeled and unlabeled primers for generation of DNA probe (e.g. IDT)
- Protein of interest
- 4x binding buffer (see Recipes)
- 20x running buffer (see Recipes)
- 5% Polyacrylamide Gel (see Recipes)
Equipment
- Mini-PROTEAN electrophoresis apparatus (Bio-Rad Laboratories)
- Fluorescence imaging device (e.g. GE Life Sciences Typhoon PhophorImager )
- Mini-PROTEAN casting system (Bio-Rad Laboratories)
Procedure
- Procedure for generating fluorescently-tagged DNA probes
- Obtain synthetic DNA oligo primers that will be used for PCR amplification of the target DNA of interest (we have had success using primers from IDT). Primers should be designed with the same strategy as general PCR primers (namely a length of 18-30 nt, melting temperatures of both primers in a pair being near 60 °C and within 3 °C of each other, and G + C content between 40%-60%, when possible). At least one of the primers must include a 5'-fluorescent tag (e.g. 6-carboxyfluorescein (6FAM)), but visualization sensitivity can be doubled if both primers are labeled. Generate the fluorescent DNA probe by a standard PCR protocol with any general polymerase enzyme. Probe sizes amenable to EMSA analysis can range from very small (25 bp) to very large (1,000 bp); however, probes between 100-500 bp work well in providing optimal resolution of DNA-protein complexes without requiring extensively long gel running times. If smaller probes are required, the gel percentage may need to be increased or the running time and/or voltage decreased.
- Excess oligos and free nucleotides should be separated from the DNA probe using a standard PCR reaction clean-up kit or by gel purifying the product of interest. The DNA probe should be suspended in dH2O or preferred buffer.
- The concentration of the probe should be determined using a spectrophotometer. A recommended working concentration is 200 nM (final concentration in EMSA reaction, 10 nM).
- Store probes at -20 °C in the dark. Probes can be stored in the manner successfully for several months, although fluorescence intensity may begin to decrease after the first month.
- Obtain synthetic DNA oligo primers that will be used for PCR amplification of the target DNA of interest (we have had success using primers from IDT). Primers should be designed with the same strategy as general PCR primers (namely a length of 18-30 nt, melting temperatures of both primers in a pair being near 60 °C and within 3 °C of each other, and G + C content between 40%-60%, when possible). At least one of the primers must include a 5'-fluorescent tag (e.g. 6-carboxyfluorescein (6FAM)), but visualization sensitivity can be doubled if both primers are labeled. Generate the fluorescent DNA probe by a standard PCR protocol with any general polymerase enzyme. Probe sizes amenable to EMSA analysis can range from very small (25 bp) to very large (1,000 bp); however, probes between 100-500 bp work well in providing optimal resolution of DNA-protein complexes without requiring extensively long gel running times. If smaller probes are required, the gel percentage may need to be increased or the running time and/or voltage decreased.
- Procedure for purifying Rgg protein of interest
We have been unable to identify a generic purification scheme compatible with various types of Rgg proteins. Solubility of expressed Rgg proteins varies greatly between homologs, necessitating individual optimization of buffer composition and affinity tag utilization. Examples of successful purification schemes can be found in the methods sections of the references provided below. Regardless of the method of purification, it should be mentioned that it is preferable to generate affinity-tagged Rgg's in such a way as to allow for removal the affinity tag from Rgg prior to use in EMSA assays, thereby helping to ensure that the tag is not interfering with the DNA-binding activity of the protein. - Procedure for EMSA
- Cast a 5% non-denaturing polyacrylamide gel following the suggested recipe below. We have successfully used the Bio-Rad mini-PROTEAN casting system for these purposes.
- Dilute the Rgg protein to the desired concentration(s) in a solution ensuring its solubility (e.g. storage or binding buffer). The optimal concentration range will depend on the Rgg protein, however serial dilutions resulting in final concentrations of 25-200 nM in the binding reactions should provide a good starting point for such optimization.
- Prepare the reaction mixture containing the following components (can prepare master mix of as many components as are shared between all reactions). Dye should not be included in the reaction in order to avoid interference with binding. It should be also noted that control reactions (i.e. probe alone and control probe of DNA not expected to be recognized by the protein) will need to be included as additional lanes on the gel:
5 μl 4x Binding Buffer + DTT
X μl non-specific DNA*
3 μl 80% glycerol
2 μl 10x BSA*
1 μl 10 mM CaCl2*
Y μl dH2O
18 μl total volume
Note: The concentrations of the starred components will need to be determined empirically and may be altered or eliminated depending on the activity and DNA-binding specificity of the Rgg protein being used, but the quantities outlined above are good starting points (as is 0.001 U/ml poly-dIdC and 50 μg/ml salmon sperm DNA for non-specific DNA). - To above binding reaction, add 1 μl Rgg protein (or buffer as control).
- To each reaction, add 1 μl fluorescently-labeled DNA probe.
- Incubate reactions at room temperature (25 °C) for 30 min.
- While the reactions are incubating, pre-run gel (no samples) at 100 V, 10 min, 4 °C in 1x running buffer (50 mM potassium phosphate pH 7.5) as the running buffer.
- Load 5-10 μl of each reaction on the gel and run at 100 V at 4 °C. The run time should be optimized depending on the size of the DNA probe (for probes between 150-300 bp 60 min is sufficient). Additionally, dye can be run in an empty lane to help visualize migration through the gel (bromophenol blue and xylene cyanol run at approximately 65 nt and 260 nt, respectfully, on a 5% gel).
- Transfer the gel to a fluorescence imaging device. We found that sandwiching gels between clear polypropylene sheet protectors was convenient for handling.
Note: The above binding reaction can be amended to include competitor DNA and/or signaling peptide by reducing the amount of dH2O in the reaction such that the total reaction volume remains 20 μl. Competitor DNA is often included at a concentration of 10-100 fold molar excess relative to the labeled probe, but will need to be optimized for each protein. If competitor DNA is added, add the DNA prior to or simultaneously with the probe. If signaling peptide is included, add the peptide (or empty vehicle as a control) 20 min after the incubation is started (10 min before loading the sample on the gel).
- Cast a 5% non-denaturing polyacrylamide gel following the suggested recipe below. We have successfully used the Bio-Rad mini-PROTEAN casting system for these purposes.
Recipes
- 4x binding buffer
80 mM HEPES (pH 7.9)
80 mM KCl
20 mM MgCl2
0.8 mM EDTA
2 mM dithiothreitol (added immediately before use) - 20x running buffer
1 M potassium phosphate (pH 7.5)
Note: The easiest way to make this is to prepare separate 1 M stocks of both K2HPO4 and KH2PO4, then mix them until the desired pH is reached. Separately, it should be noted that we customarily use potassium phosphate buffer in our EMSAs and have successfully visualized DNA-binding by three different Rgg proteins using this system. However, should it be preferred, it is possible that Tris-based buffer systems would also yield successful results (if the running buffer is changed, the gel buffer will also need to be changed accordingly). - 5% Polyacrylamide Gel (enough for 2 gels; can adjust the gel percentage as needed)
3.5 ml Acrylamide-bis (40%)
1 ml 1 M potassium phosphate (pH 7.5)
16.3 ml dH2O
200 μl 10% ammonium persulfate (make fresh)
20 μl TEMED catalyst
Note: Extra gels can be wrapped in saran wrap or damp paper towels and stored at 4 °C.
References
- Chang, J. C., LaSarre, B., Jimenez, J. C., Aggarwal, C. and Federle, M. J. (2011). Two group A streptococcal peptide pheromones act through opposing Rgg regulators to control biofilm development. PLoS Pathog 7(8): e1002190.
- LaSarre, B., Aggarwal, C. and Federle, M. J. (2013). Antagonistic Rgg regulators mediate quorum sensing via competitive DNA binding in Streptococcus pyogenes. MBio 3(6): e00333-12.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
LaSarre, B. and Federle, M. J. (2013). EMSA Analysis of DNA Binding By Rgg Proteins. Bio-protocol 3(15): e838. DOI: 10.21769/BioProtoc.838.
Category
Microbiology > Microbial biochemistry > Protein > Interaction
Molecular Biology > DNA > DNA-protein interaction
Biochemistry > Protein > Interaction > EMSA
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










