- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
High-throughput β-galactosidase and β-glucuronidase Assays Using Fluorogenic Substrates
Published: Vol 3, Iss 14, Jul 20, 2013 DOI: 10.21769/BioProtoc.827 Views: 16157
Reviewed by: Fanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
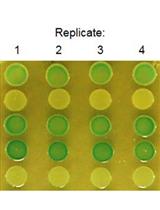

In vivo Analysis of Cyclic di-GMP Cyclase and Phosphodiesterase Activity in Escherichia coli Using a Vc2 Riboswitch-based Assay
Ying Liu [...] Ute Römling
Mar 5, 2018 7924 Views
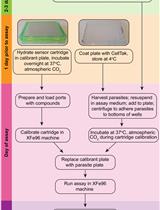
Real-Time Analysis of Mitochondrial Electron Transport Chain Function in Toxoplasma gondii Parasites Using a Seahorse XFe96 Extracellular Flux Analyzer
Jenni A. Hayward [...] Giel G. van Dooren
Jan 5, 2022 4175 Views
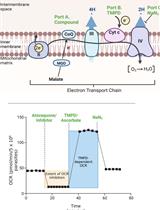
Analysis of Plasmodium falciparum Mitochondrial Electron Transport Chain Activity Using Seahorse XFe96 Extracellular Flux Assays
SaiShyam Ramesh [...] Alexander G. Maier
Nov 5, 2023 2447 Views
Abstract
β-galactosidase and β-glucuronidase enzymes are commonly used as reporters for gene expression from gene promoter-lacZ or uidA fusions (respectively). The protocol described here is a high-throughput alternative to the commonly used Miller assay (Miller, 1972) that utilises a fluorogenic substrate (Fiksdal et al., 1994) and 96-well plate format. The fluorogenic substrates 4-Methylumbelliferyl β-D-galactoside (for β-galactosidase assays) (Ramsay et al., 2013) or 4-Methylumbelliferyl β-D-glucuronide (for β-glucuronidase assays) (Ramsay et al., 2011) are cleaved to produce the fluorescent product 4-methylumbelliferone. Cells are permeabilized by freeze-thawing and lysozyme, and the production of 4-methylumbelliferone is monitored continuously by a fluorescence microplate reader as a kinetic assay. The rate of increase in fluorescence is then calculated, from which relative gene-expression levels are extrapolated. Due to the high sensitivity fluorescence-based detection of 4-methylumbelliferone and the high density of time points collected, this assay may offer increased accuracy in the quantification of low-level gene expression. The assay requires small sample volumes and minimal preparation time. The permeabilisation conditions outlined in this protocol have been optimised for Gram-negative bacteria (specifically Escherichia coli and Serratia), but is likely suitable for other organisms with minimal optimisation.
Keywords: LacZMaterials and Reagents
- Bacteria cell culture
- 4-Methylumbelliferyl β-D-galactoside (Life Technologies, catalog number: M-1489MP ) (for β-galactosidase assays)
- 4-Methylumbelliferyl β-D-glucuronide (Life Technologies, catalog number: M-1490 ) (for β-glucuronidase assays)
- Phosphate-Buffered Saline Tablets (Life Technologies, InvitrogenTM, catalog number: 00-3002 )
- Lysozyme from chicken egg white (Sigma-Aldrich, catalog number: L7651 )
- 200x stock solution for 4-Methylumbelliferyl β-D-galactoside or 4-Methylumbelliferyl β-D-glucuronide (see Recipes)
- Final working reagent of 4-Methylumbelliferyl β-D-galactoside or 4-Methylumbelliferyl β-D-glucuronide (see Recipes)
Equipment
- Fluorescence microplate reader, for example: Gemini XPS Fluorescence Microplate Reader (Molecular Devices) or Infinite 200 PRO (Tecan Group Ltd.)
- 96-Well microplates (Thermo Fisher Scientific, catalog number: 269787 )
- Flat-bottomed clear 96-well microplates (low autofluorescence and/or absorbance at 365 and 445 nm is preferable)
- Multi-channel pipette(s) (12 or 8 channel) capable of dispensing 10 μl and 100 μl
- Ultra-low temperature freezer (-70 °C)
Procedure
- Record the OD600 of the samples to be assayed. It is recommended that the OD600 of the samples to be analysed are in the 0.1-1 OD600 range. Dilute the samples in growth media to give an OD600 in the 0.1-1 range. Depending on the range of gene expression levels observed, the dilution factor may need to be optimised by analysing multiple dilutions.
- Collect 100 μl of the diluted samples (0.1-1 OD600 range) to be assayed and freeze at -70 °C in a 96-well microplate (the "master plate"). Once all samples are collected, return plate to the -70 °C and leave to freeze overnight.
For example: For a time-course experiment in E. coli, 100 μl samples could be collected every hour for 12 h in a single microplate, returning the microplate to the -70 °C after collecting each sample. This microplate now acts as the "master plate", from which replicate assays can be carried out at a later date if desired. Negative media-only controls can also be added to the master plate in free wells. - After all samples are collected (make sure all samples have been fully frozen at -70 °C), defrost the master plate in the 37 °C incubator with the lid off (to avoid cross-contamination via condensation).
- When the master plate has fully defrosted, aliquot 10 μl from each well into a new microplate (the "assay plate") using a multichannel pipette and place the assay plate in the -70 °C for at least 15 min.
- Pre-warm the microplate reader to 37 °C prior to carrying out the assay and initialise the microplate reader software so it is ready to go (see Notes about microplate reader settings for example settings).
- Prepare the final working reagent.
Note: Prepare immediately prior to use and avoid prolonged exposure to light (i.e. Use the same day). Make enough for 100 μl per reaction. - Defrost the assay plate in the 37 °C incubator with the lid off (to avoid cross-contamination via condensation) for at least 10 min and/or until any visible ice crystals have dissolved in all wells.
- Place yourself within close distance of the microplate reader. Dispense 100 μl of working reagent into each well of the 96-well microplate (or just wells containing samples) using a multichannel pipette. Try to minimize the time between dispensing reagent to each well. Immediately place the microplate in the plate reader with the lid off and start the program. Samples containing large amounts of β-galactosidase and β-glucuronidase can saturate the assay within the first 10 min, therefore it is important to capture reads quickly after the addition of substrate.
Note: For highly expressing samples that saturate the detector early, data is still recoverable. However as there are likely to be fewer timepoints with a linear increase in fluorescence the rate estimation may be less accurate. Sample dilution will improve accurate measurement of these samples. - Extract the rate of increase in fluorescence from the reads using the microplate software or export to excel (see Figure 1):
- Plot the relative fluorescence intensity (the machine carries out internal fluorescence normalisation, hence it is "relative" fluorescence intensity) over time (per second is usually convenient). Choose a linear portion of the graph with the steepest slope (Vmax) from which to extrapolate the rate (in excel, the equation = SLOPE() can be used). This will provide relative fluorescence units per second (RFU/sec). Some platereader software will calculate Vmax automatically and in real time during the assay. If the graph is curved or very noisy, you may have problems with cell permeabilization or the fluorescence detector gain settings, respectively (see Troubleshooting below).
- Optional: Subtract the average RFU/sec of negative control wells (media-only with final working reagent) from all samples (in practice this value is usually zero or very small and so this step is usually not necessary).
- Normalise to optical cell density (OD600) recorded in step 1. This will give you units of RFU/sec/OD600.
- Values can be normalized to account for sample volume used (i.e. Divided by 0.01 ml to give values as RFU/sec/ml/OD600), however this normalization is somewhat arbitrary if the same volumes (10 μl) are always used. Alternatively, standard curves with known concentrations of purified β-galactosidase or β-glucuronidase can be used to estimate actual units of enzyme concentration.

Figure 1. Plot of raw relative fluorescence units over time. Biological triplicate samples were analysed by the β-galactosidase assay over a 30-min period, as described above. Mean slope and standard deviation for the three replicates are indicated next to each group. The highly-expressed samples saturate the detector after 600 sec. Statistically significant low-level expression is also detected from the lowly-expressed sample compared to the negative media-only (with final reagent mix) control. Final values for publication/presentation are normalised to OD600.
- Plot the relative fluorescence intensity (the machine carries out internal fluorescence normalisation, hence it is "relative" fluorescence intensity) over time (per second is usually convenient). Choose a linear portion of the graph with the steepest slope (Vmax) from which to extrapolate the rate (in excel, the equation = SLOPE() can be used). This will provide relative fluorescence units per second (RFU/sec). Some platereader software will calculate Vmax automatically and in real time during the assay. If the graph is curved or very noisy, you may have problems with cell permeabilization or the fluorescence detector gain settings, respectively (see Troubleshooting below).
Notes about microplate reader settings
- Universal settings:
Additional settings may be optional on some machines, however shaking between each read is recommended if avaliable.Temperature 37 °C Excitation wavelength 360-365 nm Emmission wavelength 445-460 nm Total assay time 10-30 min depending on expression level - Specific machine settings:
Gemini XPS Fluorescence Microplate Reader
Select full plate readTemperature: 37 °C Autoshake: on Read type: kinetic read of 30 min, read intervals 1 min Excitation: 360 nm Emission: 450 nm cut-off: 435 nm Reads: 8 reads/well PMT: High Calibrate: On Infinite 200 PRO
Nuclon flat-bottom transparentlid mode off Temperature 37 °C Number of cycles 30 Kinetic time interval 1 min Shaking duration 5 sec Shaking amplitude 1 mm Shaking mode Linear Excitation 360 nm Emission 460 nm Plate read mode Top Gain Manual Gain mode Gain 85 Lagtime 0 Integration time 20 Multiple reads per well No
Troubleshooting
- Excitation/emission
4-methylumbelliferone has an excitation peak at 365 nm and emission peak at 448 nm. However the peak absorption and emission spectra observed on your particular machine and the settings available to you may vary slightly. Additionally, autofluorescence and/or absorbance from the media and/or microplate may require you to select wavelengths outside the peaks. It is best to optimize for your setup by carrying out an emission/excitation wavelength scan with a positive control, such as a sample with known β-galactosidase or β-glucuronidase activity. In our experience, LB, TY and minimal medium and Nunc MicroWell Flat-bottomed 96-Well Microplates do not generate any problems with autofluorescence. It is possible that a particular organism may generate products that fluoresce in this assay. Excitation/emission wavelength scans of samples containing appropriate negative control strains (lacking lacZ or uidA) may reveal if this is the case. - Cell permeabilization
In our hands this assay is very sensitive to the degree of cell permeabilization. In this protocol, optimized for Gram-negative bacteria, efficient cell permeabilization is achieved by freeze-thawing the samples twice between -70 and 37 °C and through the addition of high concentrations of lysozyme in the assay buffer. If cells are not completely permeabilized or become increasingly permeabilized throughout the assay, the RFU/sec plot will appear curved rather than linear. You may need to optimise the protocol to allow efficient permeabilization of your samples. - Microplate reader gain
Some microplate readers automatically adjust the detector gain over the entire plate or for each individual well. This can increase the dynamic range of the reader, as ideally both very lowly and very highly fluorescent samples can be analysed in the same plate. However for some microplate readers, the gain may need to be set manually. If fluorescence is not detected at all then the gain is likely too low. If the gain is too high, samples will saturate the detector almost immediately after the assay begins. To determine the optimal gain for your assay and machine, take samples from your most highly expressed samples and your most lowly expressed samples and carry out trial assays, adjusting the gain with each iteration. Choose a level at which lowly expressed samples have a detectible increase and the highest expressed sample doesn't saturate the detector before 10 min.
Recipes
- 200x stock solution for 4-Methylumbelliferyl β-D-galactoside or
4-Methylumbelliferyl β-D-glucuronide Dissolve 4-Methylumbelliferyl β-D-galactoside or 4-Methylumbelliferyl β-D-glucuronide in DMSO at 50 mg ml-1.
Store in small aliquots away from light at -70 °C. Avoid repeated freeze-thawing. - Final working reagent of 4-Methylumbelliferyl β-D-galactoside or 4-Methylumbelliferyl β-D-glucuronide
Dilute 200x stock to 1x in PBS buffer containing 2 mg ml-1 lysozyme
References
- Fiksdal, L., Pommepuy, M., Caprais, M.-P. and Midttun, I. (1994). Monitoring of fecal pollution in coastal waters by use of rapid enzymatic techniques. Appl Environ Microbiol 60(5): 1581-1584.
- Miller, J. H. (1972). Experiments in molecular genetics, Cold Spring Harbor Laboratory Cold Spring Harbor, New York. p. 352-355.
- Ramsay, J. P., Major, A. S., Komarovsky, V. M., Sullivan, J. T., Dy, R. L., Hynes, M. F., Salmond, G. P. and Ronson, C. W. (2013). A widely conserved molecular switch controls quorum sensing and symbiosis island transfer in Mesorhizobium loti through expression of a novel antiactivator. Mol Microbiol 87(1): 1-13.
Article Information
Copyright
© 2013 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ramsay, J. P. (2013). High-throughput β-galactosidase and β-glucuronidase Assays Using Fluorogenic Substrates. Bio-protocol 3(14): e827. DOI: 10.21769/BioProtoc.827.
Category
Microbiology > Microbial cell biology > Cell-based analysis > Reporter assay
Molecular Biology > DNA > Gene expression
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











