- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vitro Homeostatic Proliferation of Human CD8 T Cells
Published: Vol 7, Iss 22, Nov 20, 2017 DOI: 10.21769/BioProtoc.2619 Views: 12973
Reviewed by: Ivan ZanoniLokesh KalekarAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Isolation and Culture of Ferret Airway Stem Cells
Ziying Yan [...] Feng Yuan
Jul 20, 2025 2413 Views
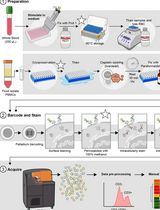
Dual Phospho-CyTOF Workflows for Comparative JAK/STAT Signaling Analysis in Human Cryopreserved PBMCs and Whole Blood
Ilyssa E. Ramos [...] James M. Cherry
Nov 20, 2025 2351 Views
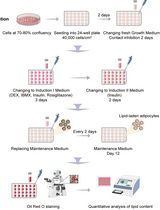
Optimization of Adipogenic Differentiation Protocol for Murine and Human Cell Culture Models
Junwan Fan [...] Wenyan He
Jan 20, 2026 232 Views
Abstract
Long-lived T-cell–mediated immunity requires persistence of memory T cells in an antigen-free environment while also maintaining a heightened capacity to recall effector functions. Such antigen-independent homeostatic proliferation is mediated in part by the common gamma-chain cytokines IL-7 and IL-15. To further explore the mechanisms governing maintenance of effector functions in long-lived memory T cells during antigen-independent proliferation, human naïve and memory CD8 T cells can be sorted from peripheral blood mononuclear cells (PBMCs), labeled with the proliferation-tracking dye carboxyfluorescein succinimidyl ester (CFSE), and then purified based on their levels of cell division. This allows investigators to assess differences in the desired molecular target in cells that have undergone cytokine-driven proliferation. We provide here a protocol for assessing epigenetic programs in divided and undivided human naïve and memory CD8 T cells following 7 days in culture with IL-7 and IL-15 to illustrate how this approach can shed light on the mechanism(s) that governs the preservation of effector functions during homeostasis of long-lived memory CD8 T cells.
Keywords: Homeostatic proliferationBackground
A cardinal feature of adaptive immunity is the development of immunological memory against previously encountered pathogens (Plotkin et al., 2013). Memory CD8 T cells play a major role in providing life-long protection against pathogens previously encountered by the host, but in order to provide long-lived protection, T cells must have acquired the ability to persist and maintain effector functions in an antigen-free environment. During this time, memory T cells undergo antigen-independent proliferation in response to IL-7/15 cytokines, albeit to different degrees.
The total pool of memory CD8 T cells is a heterogeneous combination of several different cell subsets that respond differently to homeostatic cytokines. For instance, the highly proliferative central memory (Tcm) subset of CD8 T cells expresses the chemokine receptor CCR7, which enables those cells to access lymphoid tissue. Conversely, the low proliferative effector memory (Tem) subset of CD8 T cells lacks CCR7 expression and has limited access to lymphoid tissue (Sallusto et al., 1999; Masopust et al., 2001). Another recently defined subset of memory T cells, stem-cell memory (Tscm), which were named based on their heightened ability to self-renew and give rise to other memory subsets, exhibits the greatest level of cytokine-driven proliferation, compared to the other subsets (Gattinoni et al., 2011). Although phenotypically and functionally distinct, all of these memory T-cell subsets retain their ability to rapidly recall effector function upon antigen re-exposure. Thus, a common defining feature for the development of the various memory CD8 T-cell subsets is the maintenance of effector functions in the absence of antigen. However, the cell-intrinsic mechanisms that maintain memory-associated effector functions remain poorly understood. Below we describe an in vitro homeostatic proliferation assay that enables us to analyze mechanisms that regulate fate commitment during human memory T-cell homeostasis. We specifically describe loci-specific bisulfite-sequencing DNA-methylation analysis to examine the stability of effector DNA-methylation programs in memory CD8 T-cell subsets over several rounds of cell division (Abdelsamed et al., 2017).
Materials and Reagents
- ART tips 1,000 μl (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 2079G )
- ART tips 200 μl (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 2069G )
- ART tips 10 μl (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 2139-RT )
- 5 ml STRIPETTE (Corning, Costar®, catalog number: 4487 )
- 10 ml STRIPETTE (Corning, Costar®, catalog number: 4488 )
- 25 ml STRIPETTE (Corning, Costar®, catalog number: 4489 )
- 18 G x 1-inch disposable needles (BD, catalog number: 305195 )
- 30 ml BD Luer-Lok syringe (BD, catalog number: 302832 )
- Falcon conical tubes, 50-ml (Corning, Falcon®, catalog number: 352098 )
- Falcon conical tubes, 15-ml (Corning, Falcon®, catalog number: 352057 )
- 96-well cell culture plates, round bottom (Corning, Costar®, catalog number: 3799 )
- Sterile polystyrene Petri dishes 100 x 15 mm (Fisher Scientific, catalog number: FB0875712 )
- Apheresis blood unit
- XL10-Gold Ultracompetent cells (Agilent Technologies, catalog number: 200314 )
- pGEM-T Vector System I (Promega, catalog number: A3600 )
- 1x calcium- and magnesium-free PBS (Lonza, catalog number: 17-516Q )
- Ficoll-Paque PLUS (GE Healthcare, catalog number: 17-1440-02 )
- Trypan blue solution, 0.4% (Sigma-Aldrich, catalog number: T8154 )
- EasySep Human CD8+ T Cell Enrichment Kit (STEMCELL Technologies, catalog number: 19053 )
- Antibodies
CD8 (BioLegend, catalog number: 301033 )
CD45RO (BioLegend, catalog number: 304210 )
CD45RA (BioLegend, catalog number: 304134 )
CCR7 (BD, BD Biosciences, catalog number: 561271 )
CD95 (BioLegend, catalog number: 305622 )
Live dead stain (Tonbo Biosciences, catalog number: 13-0870 ) - CpGenome Direct Prep Bisulfite Modification Kit (Merck, catalog number: 17-10451 )
- Zymoclean Gel DNA Recovery Kit (ZYMO RESEARCH, catalog number: D4002 )
- S.O.C. medium (Thermo Fisher Scientific, InvitrogenTM, catalog number: 15544034 )
- DirectPrep 96 MiniPrep Kit (QIAGEN, catalog number: 27361 )
- Primers for methylation-specific PCR (Purchased from Integrated DNA Technologies) (see Notes for design)
- Heat-inactivated fetal bovine serum (FBS) (GE Healthcare, HycloneTM, catalog number: SH30910.03 )
- RPMI 1640 with L-glutamine (Mediatech, catalog number: 10-040-CM )
- 0.5 M EDTA (Invitrogen, catalog number: 15575-038 )
- Sodium azide (Sigma-Aldrich, catalog number: S2002-25G )
- Carboxyfluorescein succinimidyl ester (CFSE) (Life Technologies, catalog number: C1165 )
- Gentamicin (50 mg/ml) (Thermo Fisher Scientific, GibcoTM, catalog number: 15750060 )
- Penicillin-streptomycin (10,000 U/ml) (Thermo Fisher Scientific, GibcoTM, catalog number: 15140122 )
- Human recombinant IL-7 stock, 250 µg/ml (PeproTech, catalog number: AF-200-07 )
- Human recombinant IL-15 stock, 250 µg/ml (PeproTech, catalog number: AF-200-15 )
- 10x Tris/Boric Acid/EDTA (TBE), nucleic acid electrophoresis buffer (Bio-Rad Laboratories, catalog number: 1610733 )
- Agarose (Hoefer, catalog number: GR140-500 )
- Ethidium bromide 10 mg/ml (Thermo Fisher Scientific, InvitrogenTM, catalog number: 15585011 )
- LB agar miller (Sigma-Aldrich, catalog number: L3027-1KG )
- 2x LB broth (Teknova, catalog number: L8080 )
- Ampicillin (Sigma-Aldrich, catalog number: A9518-5G )
- IPTG (Sigma-Aldrich, catalog number: I6758-10G )
- X-gal (Lambda Biotech)
- JumpStart Taq ReadyMix 2x (Sigma-Aldrich, catalog number: P2893 )
- Heat-inactivated FBS (see Recipes)
- RPMI 1640 with glutamine and 4% FBS (see Recipes)
- Enrichment buffer (see Recipes)
- RPMI 1640 with glutamine, 4% FBS, and 0.02% sodium azide (see Recipes)
- FACS buffer (see Recipes)
- 4 µM CFSE working solution (see Recipes)
- RPMI 1640 with glutamine and 10% FBS (see Recipes)
- Culture medium (see Recipes)
- IL-7 and IL-15 working solutions (see Recipes)
- 1x TBE (see Recipes)
- 2% agarose gel (see Recipes)
- LB agar plates (see Recipes)
Equipment
- Neubauer hemacytometer (Hausser Scientific, catalog number: 3200 )
- P1000 (Gilson, catalog number: F123602 )
- P200 (Gilson, catalog number: F123601 )
- P10 (Gilson, catalog number: F144802 )
- Water bath
- 37 °C shaker
- 37 °C incubator
- Sorval Legend XTR centrifuge (Thermo Fisher Scientific, Thermo ScientificTM, model: SorvalTM LegendTM XTR , catalog number: 75004520)
- EasySep STEMCELL Technologies 15-ml and 50-ml magnets
- Tissue culture incubator
- Biosafety cabinet
- Horizontal gel electrophoresis unit (Hoefer, model: SUBHT )
- Vacuum manifold
- Nikon Eclipse TS100 microscope (Nikon Instruments, model: Eclipse TS100 )
- Sony Biotechnology SY3200 sorter (Synergy) (Sony Biotechnology, model: SY3200 )
- Autoclave (Getinge, model: 133LS )
Software
- FlowJo Software (Version 9.7.6)
- QUMA Software (http://quma.cdb.riken.jp/)
Procedure
- Isolation of human peripheral blood mononuclear cells (PBMCs)
- Collect the blood from an apheresis blood unit by using a 30 ml syringe with an 18-G, 1-inch needle.
Note: Dispense all blood products, tubes, and solutions in a biohazard bag. - Slowly dispense 10 ml blood on the wall of a 50 ml tube containing 25 ml 1x calcium- and magnesium-free PBS.
- With steady flow, overlay the diluted blood on the wall of a 50 ml tube containing 15 ml Ficoll.
- Spin at 400 x g for 15 min without brakes.
- Collect the interphase, which contains the peripheral blood mononuclear cells, and transfer it to a 50 ml tube.
- Wash with RPMI 1640 with glutamine and 4% FBS (see Recipes) at 10x the volume and then spin at 400 x g for 5 min.
- Collect the blood from an apheresis blood unit by using a 30 ml syringe with an 18-G, 1-inch needle.
- Enrichment of human CD8 T cells
- Decant the supernatant from step A6, and resuspend the cell pellet in enrichment buffer for counting (see Recipes).
- Take 10 µl of the cell suspension, add to 90 µl trypan blue (1:10 dilution), and then use a hemacytometer for counting (expected cell number is 108-109).
- After counting, prepare cell suspension at a concentration 5 x 107 cells/ml (if > 4.25 x 108 total cells, then use a 50 ml tube; if < 4.25 x 108 total cells, then use a 15-ml tube).
- Add the enrichment cocktail (included in the EasySep kit) at 50 µl/ml cells, mix well by pipetting up and down, and incubate at room temperature for 10 min.
- During the 10 min incubation, vortex the EasySep D magnetic particles for 30 sec or until you observe a uniform suspension.
- Add the particles at 150 µl/ml cells, mix well, and incubate at room temperature for 5 min.
- After a 5 min incubation, use enrichment buffer (see Recipes) to bring the cell suspension to a total volume of 10 ml if cell number < 4.25 x 108 total cells or 20 ml if > 4.25 x 108 total cells.
- Place the tube on the EasySep magnet.
- After 5 min, use a pipette to collect the negative fraction from the 50 ml tube. If a 15 ml tube was used, pour off the desired fraction, in one continuous motion, into a new 50-ml tube for counting (similar to step B2). For more details about the protocol and this step, please visit https://www.stemcell.com/easysep-human-cd8-t-cell-enrichment-kit.html.
- Decant the supernatant from step A6, and resuspend the cell pellet in enrichment buffer for counting (see Recipes).
- Sorting human naïve and memory CD8 T cell subsets
- After counting (expected cell number is 10-15% of the total PBMCs count), in a 50 ml tube wash the cells with RPMI and 4% FBS (till the 50 ml mark) and spin at 400 x g for 5 min.
- Decant, resuspend in FACS buffer, and distribute the cells as 50 x 106 cells per FACS tube in a 100 µl final volume.
- Stain for CCR7 antigen and then incubate in a water bath at 37 °C for 20 min (see Notes and Materials and Reagents’ above for fluorochromes, clones, manufacturer, catalog numbers and dilutions-Table 1).
Table 1. List of human antibody conjugated fluorochromes used for cell sorting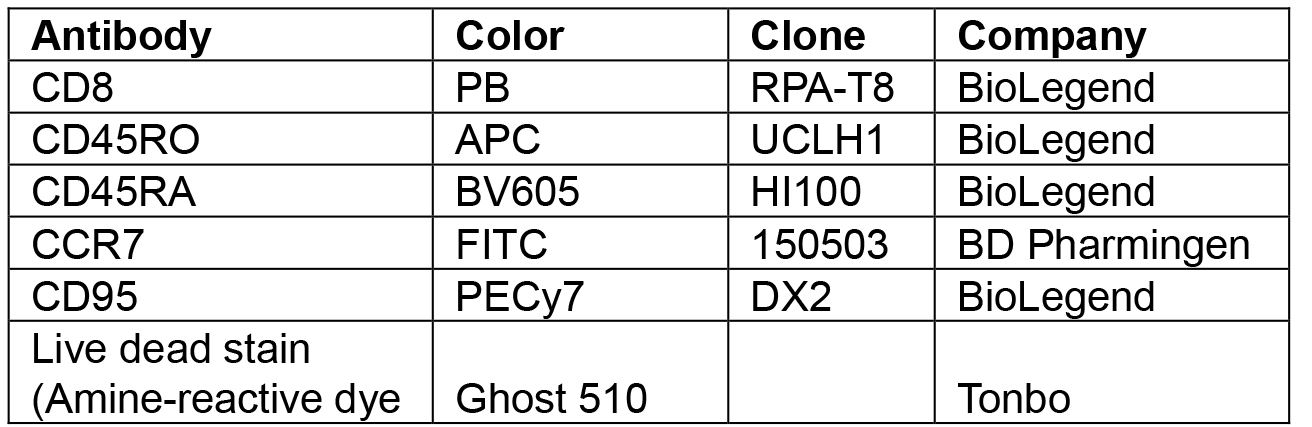
Note: All antibody dilutions are 1:100 except for anti-CD95 PECy7 antibody that is 1:20. Final volume is 100 µl. Live dead staining is performed prior to surface stains in serum-free pbs. - Wash the cells with RPMI 4% FBS, and 0.02% sodium azide (see Recipes and Notes) and then spin at 400 x g for 5 min.
- Decant and stain for CD8, CD45RO, CD45RA, CD95, and living/dead stain and incubate at room temperature in the dark for 30 min.
- Wash the cells with FACS buffer and then spin at 400 x g for 5 min.
- Decant and pool all of the stained cells into a single tube for sorting.
- Sort live naïve and memory CD8 T cell subsets based on the following surface markers:
- Naïve CD8 T cells: CCR7+, CD45RO-, CD45RA+, CD95-
- Tem CD8 T cells: CCR7-, CD45RO+
- Tcm CD8 T cells: CCR7+, CD45RO+
- Tscm CD8 T cells: CCR7+, CD45RO-, CD95+
- Naïve CD8 T cells: CCR7+, CD45RO-, CD45RA+, CD95-
- After counting (expected cell number is 10-15% of the total PBMCs count), in a 50 ml tube wash the cells with RPMI and 4% FBS (till the 50 ml mark) and spin at 400 x g for 5 min.
- Labeling sorted T cells with the proliferation-tracking dye CFSE
- Wash the sorted cells with prewarmed FACS buffer (see Recipes) and then spin at 400 x g for 5 min.
- Decant and resuspend 500,000 cells in 1 ml prewarmed FACS buffer.
- Prepare 4 µM CFSE working solution (see Recipes), then add 1 ml working solution to 1 ml cell suspension, and incubate at 37 °C for 5 min.
- Wash with 50 ml cold RPMI and 20% FBS (see Recipes) and then place on ice for 5 min.
- Spin at 400 x g for 5 min at 4 °C.
- Repeat steps D4 and D5.
- Decant completely, resuspend in 150 µl culture medium (see Recipes), and transfer the cell suspension into a 96-well, round-bottom plate.
- Prepare IL-7 and IL-15 working solutions at 50 ng/ml each (see Recipes), and then add 150 µl working solution to the 150 µl cell suspension in the 96-well plate.
- Incubate at 37 °C in 5% CO2.
- After 7 days of culture, sort undivided and divided cell populations (third division or more) for DNA-methylation analysis as shown in Figure 1A (Abdelsmed et al., 2017).
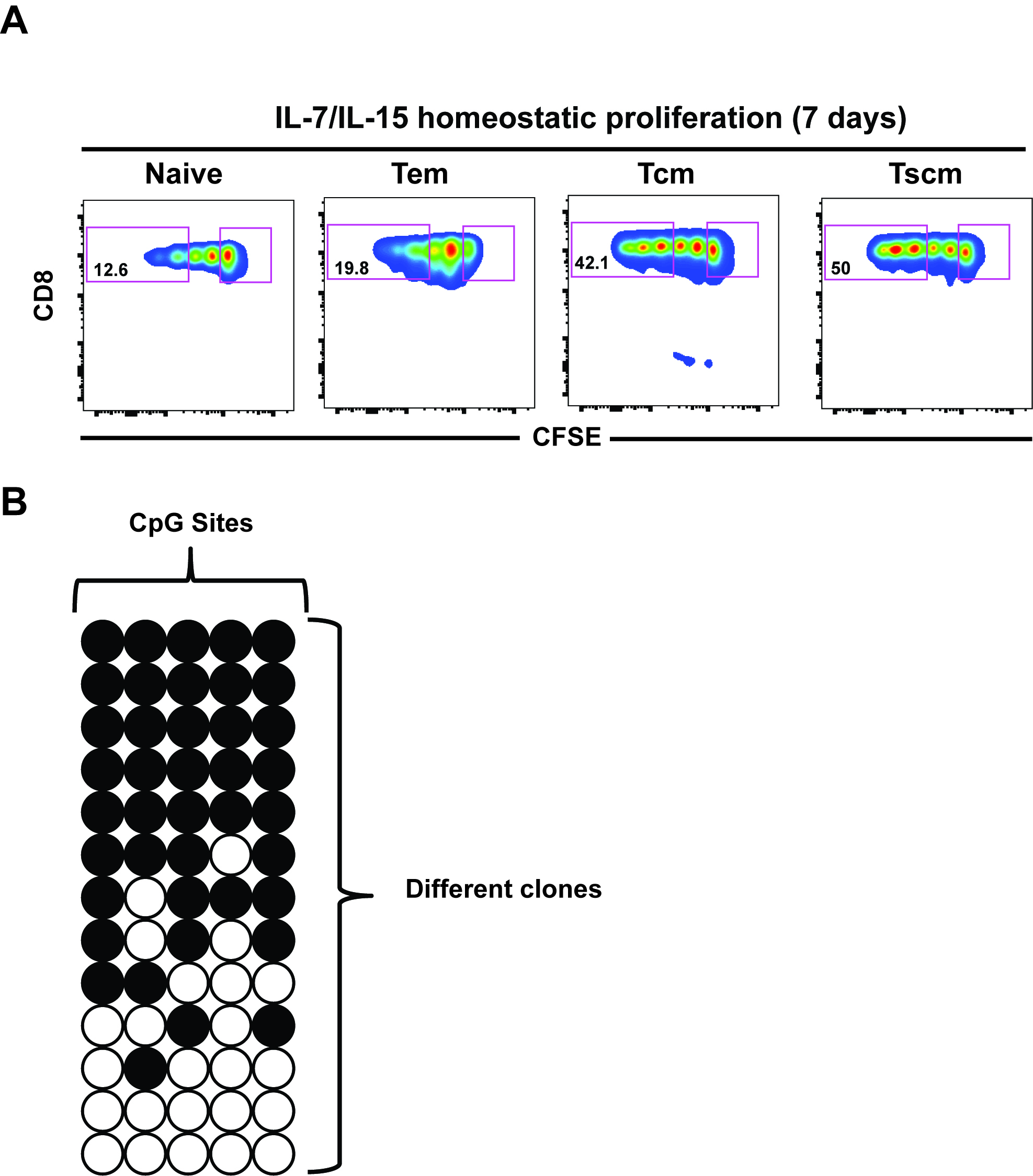
Figure 1. Representative data analysis. A. FACS plots: CFSE dilution from one representative donor showing proliferation of sorted CD8 T cell subsets after IL-7/IL-15 in vitro culture for 7 days. Percentage in each gate represents cells dividing more than 3 divisions. B. Bisulfite sequencing analysis: Each horizontal line represents a clone while vertical lines are different CpG sites in the differentially methylated region. Open circles depict unmethylated CpG while filled circles depict methylated CpG. These data and analyses are representative of published data (Abdelsamed et al., 2017; Ghoneim et al., 2017).
- Wash the sorted cells with prewarmed FACS buffer (see Recipes) and then spin at 400 x g for 5 min.
- Loci-specific bisulfite-sequencing analysis (DNA-methylation analysis)
- Sorted cells are lysed, and DNA undergoes bisulfite treatment using the CpGenome Direct Prep Bisulfite Modification Kit.
- Bisulfite-treated DNA is used as a template to amplify a specific target sequence using JumpStart ReadyMix 2x. The Methylation-specific PCR program is as follows:
- 95 °C for 5 min
- 95 °C for 30 sec
- 55 °C 1:30 min
- Go to step b 49 times
- 4 °C indefinitely
- 95 °C for 5 min
- Prepare 2% agarose gel containing ethidium bromide (see Recipes).
- Pour the gel into the gel tray and then wait until it solidifies.
- Load the PCR product into the gel and then run the gel using 1x TBE (see Recipes).
- Cut the appropriate band size, digest the gel, and extract the DNA using the Zymoclean Gel DNA Recovery Kit.
- Use the entire PCR product for the ligation reaction using pGEM-T Vector System I kit.
Note: Overnight incubation at 4 °C is recommended. - Mix the ligation product with 50 µl XL-10–competent bacteria and place on ice for 20 min.
Note: It is recommended to use one third of the ligation product for transformation and save the rest in -20 °C. - Incubate the tubes containing bacteria/DNA mix at 42 °C for 45 sec.
- Place the tubes immediately on ice for 2 min.
- Add 400 µl S.O.C. medium to each tube and then incubate the tubes in a shaker (150 rpm) at 37 °C for 90 min.
- Spread 150 µl of the bacterial suspension on LB agar plates (see Recipes).
- Incubate at 37 °C for 18 h.
- In a deep 96-well plate, inoculate one white bacterial colony in each well containing 900 µl LB medium with 0.1 mg/ml ampicillin (see Notes).
- Incubate the cell cultures at 37 °C for 18-22 h with shaking (220 rpm).
- Spin the 96-deep well plates at 1,600 x g for 15 min.
- Decant the supernatant and then extract the plasmid from the bacterial pellet by following the instructions in DirectPrep 96 Miniprep Kit.
- Elute the DNA with 60 µl elution buffer, dry the 96-well plate overnight, and then use 300-600 ng DNA for Sanger sequencing using pUC/M13 reverse primers.
- Upload the Ab1 sequencing files as zip files to the QUMA website (QUantification tool for Methylation Analysis, http://quma.cdb.riken.jp/).
- After analyzing the data by using the QUMA website, its readout is depicted as shown in Figure 1B.
- Sorted cells are lysed, and DNA undergoes bisulfite treatment using the CpGenome Direct Prep Bisulfite Modification Kit.
Data analysis
FlowJo Software (Version 9.7.6) was used to analyze flow cytometry data (Figure 1A), while DNA methylation data was analyzed using QUMA Software (http://quma.cdb.riken.jp/). Each sequencing AB1 file represents a clone. Briefly, all AB1 files for each clone were compressed and uploaded to the software then the target sequence of the gene of interest was aligned against the sequenced clones. The software readout is shown in Figure 1B.
Notes
- The methylated cytosines are protected from a deamination reaction, while the nonmethylated cytosines are converted to uracils. Using Sanger sequencing, this single-nucleotide–resolution method detects methylated cytosines as cytosines and nonmethylated cytosines as thymines. To learn more about bisulfite conversion, visit the Zymo Research website (www.zymoresearch.com/epigentics).
- Primers used in the methylation-specific PCR program should have the following conditions:
- Do not contain any CpG sites
- Annealing temperature: 56-60 °C
- Length: 26-30 bp
- All cytosines should be converted to thymines. In case of the forward primer, all cytosines are converted to thymines while in case of reverse primer; all cytosines are converted to thymines followed by reverse complement.
- Do not contain any CpG sites
- Sodium azide is recommended to be added to the staining buffer since anti-CD95 antibody can induce cell apoptosis.
- CCR7 is a chemokine receptor that recycles through the cell membrane. Incubation at 37 °C improves its detection.
- To obtain representative data, you need to inoculate 16-24 bacterial colonies per agar plate, which represents one given condition.
Recipes
- Heat-inactivated FBS
Incubate a bottle of FBS in a water bath at 56 °C for 30 min and then prepare 50 ml aliquots and store at -20 °C - RPMI 1640 with glutamine and 4% FBS
Add 40 ml heat-inactivated FBS to 1 L RPMI with glutamine - Enrichment buffer
Add 10 ml heat-inactivated FBS to 500 ml 1x PBS without calcium and magnesium + 1 ml EDTA (0.5 M EDTA) - RPMI 1640 with glutamine, 4% FBS, and 0.02% sodium azide
Add 40 ml heat-inactivated FBS to 1 L RPMI with glutamine + 0.2 g sodium azide - FACS buffer
Add 20 ml heat-inactivated FBS to 500 ml 1x PBS without calcium and magnesium - 4 µM CFSE working solution
Add 1.5 µl CFSE from stock (44.8 mM or 25 mg/ml) to 10 ml warm PBS - RPMI 1640 with glutamine and 20% FBS
Add 100 ml heat-inactivated FBS to 1 L RPMI with glutamine - Culture medium
In 1 L RPMI with glutamine, add the following:
100 ml heat-inactivated FBS
1 ml gentamycin (1:1,000 dilution)
10 ml penicillin-streptomycin (1:100 dilution) - IL-7 and IL-15 working solutions
- Add 1 µl of 250 µg/ml IL-7 and 1 µl of 250 µg/ml IL-15 into 2.5 ml of culture medium
- Dilute into culture medium at 1:1 ratio, then use directly 150 µl and add to 150 µl of CFSE labeled cells seeded in a 96-well plate
- Add 1 µl of 250 µg/ml IL-7 and 1 µl of 250 µg/ml IL-15 into 2.5 ml of culture medium
- 1x TBE
Dilute 10x TBE to 1x TBE using deionized water - 2% agarose gel
- Add 2 g agarose LE to 100 ml of 1x TBE and then heat in a microwave until the gel dissolves completely
- Add 5 µl ethidium bromide (10 mg/ml) to achieve the final concentration of 5 ng/ml
- Mix well and then pour into the gel tray
- Add 2 g agarose LE to 100 ml of 1x TBE and then heat in a microwave until the gel dissolves completely
- LB agar plates
- Dissolve 40 g LB agar into 1 L of double-distilled water and then autoclave
- Bring the temperature down to 45 °C and then add 100 mg ampicillin, 230 mg IPTG, and 100 µl X-gal
- Mix well and pour 15 ml into a sterile Petri dish and leave open under a biosafety cabinet
- Wait until the agar plates are solidified then store at 4 °C for up to one month
- Dissolve 40 g LB agar into 1 L of double-distilled water and then autoclave
Acknowledgments
We thank Dr. Angela McArthur for scientific editing. This work was supported by the National Institutes of Health grant 1R01AI114442 and ALSAC (to B.Y.). The authors declare no competing financial interests. This protocol was adapted and modified from Lugli et al., 2013-Identification, isolation and in vitro expansion of human and nonhuman primate T stem cell memory cells. Nat Protoc 8(1): 33-42.
References
- Abdelsamed, H. A., Moustaki, A., Fan, Y., Dogra, P., Ghoneim, H. E., Zebley, C. C., Triplett, B. M., Sekaly, R. P. and Youngblood, B. (2017). Human memory CD8 T cell effector potential is epigenetically preserved during in vivo homeostasis. J Exp Med 214(6): 1593-1606.
- Gattinoni, L., Lugli, E., Ji, Y., Pos, Z., Paulos, C. M., Quigley, M. F., Almeida, J. R., Gostick, E., Yu, Z., Carpenito, C., Wang, E., Douek, D. C., Price, D. A., June, C. H., Marincola, F. M., Roederer, M. and Restifo, N. P. (2011). A human memory T cell subset with stem cell-like properties. Nat Med 17(10): 1290-1297.
- Ghoneim, H. E., Fan, Y., Moustaki, A., Abdelsamed, H. A., Dash, P., Dogra, P., Carter, R., Awad, W., Neale, G., Thomas, P. G. and Youngblood, B. (2017). De novo epigenetic programs inhibit PD-1 blockade-mediated T cell rejuvenation. Cell 170(1): 142-157 e119.
- Lugli, E., Gattinoni, L., Roberto, A., Mavilio, D., Price, D. A., Restifo, N. P. and Roederer, M. (2013). Identification, isolation and in vitro expansion of human and nonhuman primate T stem cell memory cells. Nat Protoc 8(1): 33-42.
- Masopust, D., Vezys, V., Marzo, A. L. and Lefrancois, L. (2001). Preferential localization of effector memory cells in nonlymphoid tissue. Science 291(5512): 2413-2417.
- Plotkin, S. A., Orenstein, W. A. and Offit, P. A. (2013). Vaccines. (6th edition). 1550.
- Sallusto, F., Lenig, D., Forster, R., Lipp, M. and Lanzavecchia, A. (1999). Two subsets of memory T lymphocytes with distinct homing potentials and effector functions. Nature 401(6754): 708-712.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Abdelsamed, H. A., Zebley, C. C. and Youngblood, B. (2017). In vitro Homeostatic Proliferation of Human CD8 T Cells. Bio-protocol 7(22): e2619. DOI: 10.21769/BioProtoc.2619.
Category
Immunology > Immune cell differentiation > T cell
Immunology > Immune cell function > Cytokine
Cell Biology > Cell isolation and culture > Cell differentiation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link