- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Minimum Inhibitory Concentration (MIC) Assay for Antifungal Drugs
Published: Vol 2, Iss 20, Oct 20, 2012 DOI: 10.21769/BioProtoc.252 Views: 40131

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Intracellular cAMP Measurements in Candida albicans Biofilms
Liuliu Jiang [...] Xin Wei
Dec 5, 2019 4686 Views

Competition Assays to Quantify the Effect of Biocontrol Yeasts against Plant Pathogenic Fungi on Fruits
Electine Magoye [...] Florian M. Freimoser
Feb 5, 2020 5426 Views
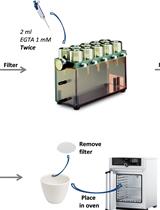
Determination of the Cellular Ion Concentration in Saccharomyces cerevisiae Using ICP-AES
Louise Thines [...] Pierre Morsomme
Aug 20, 2020 3450 Views
Abstract
The Minimum Inhibitory Concentration (MIC) Assay is widely used to measure the susceptibility of yeasts to antifungal agents. In serial two-fold dilutions, the lowest concentration of antifungal drug that is sufficient to inhibit fungal growth is the MIC. Typically, 50% inhibitory (MIC50) or 80% inhibitory (MIC80) values are reported. To facilitate visualization of antifungal susceptibility data, heat maps are generated whereby optical density values are represented quantitatively with colour.
Materials and Reagents
- Strain to be analyzed
- Overnight cultures of the strains to be analyzed and parental controls
- Glucose
- MOPS
- Yeast extract
- Bactopeptone
- Yeast nitrogen base
- Media
- RPMI medium 1640 (Life Technologies, Gibco®, catalog number: 31800-089 ) (see Recipes)
Note: It is the standard medium recommended by the CLSI standard protocol.
- YPD (see Recipes)
Note: It is a rich medium commonly used to assess susceptibility of diverse mutants.
- Synthetic defined medium (see Recipes)
Note: Supplemented with required amino acids to enable growth of auxotrophic strains is commonly used for experiments requiring plasmid selection.
- RPMI medium 1640 (Life Technologies, Gibco®, catalog number: 31800-089 ) (see Recipes)
Equipment
- 96-well tissue culture plate (flat bottom with lid) (SARSTEDT AG, catalog number: 83.1835 )
- Microplate reader
- Multichannel pipette [optional: Multichannel electronic pipette (Rainin, model: EDP3-Plus )]
Software
- Soft Max Pro or equivalent spectrophotometer software (Microplate Data Processing)
- Microsoft Excel (Data Analysis)
- JavaTree View (Heat Map Generation, http://jtreeview.sourceforge.net/)
Procedure
- Grow an overnight culture of the strains to be assayed, including relevant controls, in 3 ml of the appropriate medium being tested.
- Prepare the MIC plate with the drug gradient to be tested.
- Always set up the plates in duplicate.
- Add 100 µl of the medium to all 96 wells except for column 12 using a multi-channel pipette.
- To column 12, add 200 µl of the medium that contains drug at 2x the final concentration that the titration will start at.
- Perform 2-fold dilutions by transferring 100 µl of the drug-containing medium from column 12 to column 11 (mix well and repeat across the row). Stop at column 2 and discard the excess 100 µl so that column 1 is the drug-free control.
- Measure the OD600 of 100 µl of the overnight cultures in duplicates in a 96 well plate.
- Subtract the OD600 reading from medium only wells to account for background.
- Calculate the average OD600 for each strain.
- To prepare the inoculum in 5 ml of medium, add 1/OD600 x 1.5 µl of the overnight culture.
- Add 100 µl of diluted inoculum (~103 cells) to every well from columns 1 to 12, one strain per row.
- Incubate the plates statically at the appropriate temperature (35 °C is the standard temperature approved by CLSI standard, 37 °C is a clinically relevant temperature, and 30 °C is a commonly used laboratory temperature) for the desired amount of time, often for 24, 48, or 72 h.
- To prepare plates for OD600 reading, mix wells with a multi-channel pipette.
- Process the data in Excel.
- Copy OD600 values for each drug replicate into a new worksheet.
- Above the first row of values, label the drug gradient with the appropriate concentration for each column.
- Label the strains in the left-most column.
- Calculate the average of medium-only values and subtract it from all values on the plate. Convert any negative numbers to “0”.
- Calculate the average of values from each duplicate.
- Optional: To normalize growth in the drug gradient relative to its drug-free control, divide the OD600 values of each well by the OD600 value of the well in the column 1 of the corresponding row.
- Copy the referenced values to a new worksheet using “paste special”, select “values only” and arrange in the following format.
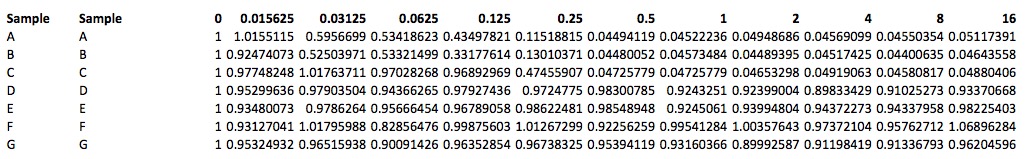
Note: This data was updated by authors on Oct. 15, 2015.
- Save as a text (tab delimited) file.
- Plot the data as heat map in JavaTree View.
- Open the text (tab delimited) file in JavaTree View.å.
- Under setting, make the following changes in pixel.
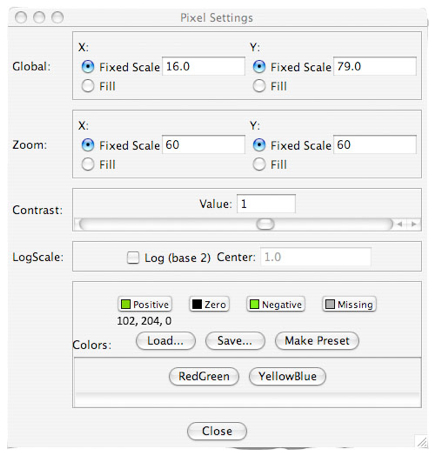
Zoom X: Fixed Scale = 60
Zoom Y: Fixed Scale = 60
Contrast Value: 1
Positive: 102, 204, 0 (Green)
- Export the file
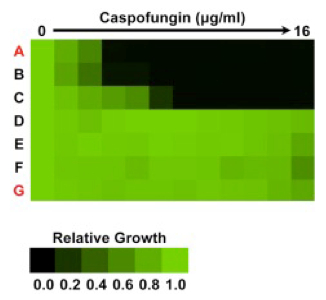
The above figure is adapted from Reference 2, and plotted based on the data shown in the table associated with point 8 g above. Growth measured by optical density is quantitatively displayed with color as indicated in the color bar, such that bright green represents full growth and black represents no growth. As indicated above, growth of each strain can be plotted as values normalized to the drug-free control for the same strain, or growth can be directly plotted without normalization.
Recipes
- RPMI medium 1640 (pH 7.0)
3.5% MOPS
2% glucose
- YPD
1% yeast extract
2% bactopeptone
2% glucose
- Synthetic defined medium
0.67% yeast nitrogen base
2% glucose
Acknowledgments
The method reported in this protocol is based on that described in Singh-Babak et al. (2012). JLX was supported by an Ontario Graduate Scholarship, a University of Toronto Open Fellowship, and a Canadian Institutes of Health Research (CIHR) Frederick Banting & Charles Best CGS Doctoral Award, SDSB by a CIHR Frederick Banting & Charles Best CGS Doctoral Award, and LEC by a Career Award in the Biomedical Sciences from the Burroughs Wellcome Fund, by a Canada Research Chair in Microbial Genomics and Infectious Disease, by a Ministry of Research and Innovation Early Researcher Award, and by NSERC Discovery Grant 355965-2009.
References
- Clinical and Laboratory Standards Institute (CLSI). Reference Method for Broth Dilution Antifungal Susceptibility Testing of Yeasts; Approved Standard-Third Edition. CLSI document M27-A3 (ISBN 1-56238-666-2). Clinical and Laboratory Standards Institute, 940 West Valley Road, Suite 1400, Wayne, Pennsylvania 19087-1898 USA, 2008.
- Singh-Babak, S. D., Babak, T., Diezmann, S., Hill, J. A., Xie, J. L., Chen, Y. L., Poutanen, S. M., Rennie, R. P., Heitman, J. and Cowen, L. E. (2012). Global analysis of the evolution and mechanism of echinocandin resistance in Candida glabrata. PLoS Pathog 8(5): e1002718.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Xie, L., Singh-Babak, S. D. and Cowen, L. E. (2012). Minimum Inhibitory Concentration (MIC) Assay for Antifungal Drugs. Bio-protocol 2(20): e252. DOI: 10.21769/BioProtoc.252.
- Singh-Babak, S. D., Babak, T., Diezmann, S., Hill, J. A., Xie, J. L., Chen, Y. L., Poutanen, S. M., Rennie, R. P., Heitman, J. and Cowen, L. E. (2012). Global analysis of the evolution and mechanism of echinocandin resistance in Candida glabrata. PLoS Pathog 8(5): e1002718.
Category
Microbiology > Microbial biochemistry > Other compound
Microbiology > Antimicrobial assay > Antifungal assay
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









