- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Establishment of a Human Cell Line Persistently Infected with Sendai Virus
Published: Vol 7, Iss 16, Aug 20, 2017 DOI: 10.21769/BioProtoc.2512 Views: 9112
Reviewed by: Yannick DebingAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
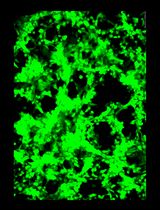
General Maintenance and Reactivation of iSLK Cell Lines
Ariana C. Calderón-Zavala [...] Ekaterina E. Heldwein
Jun 5, 2025 1896 Views
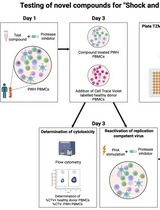
Inducible HIV-1 Reservoir Reduction Assay (HIVRRA), a Fast and Sensitive Assay to Test Cytotoxicity and Potency of Cure Strategies to Reduce the Replication-Competent HIV-1 Reservoir in Ex Vivo PBMCs
Jade Jansen [...] Neeltje A. Kootstra
Jul 20, 2025 2460 Views
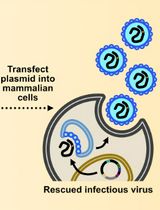
Assembly and Mutagenesis of Human Coronavirus OC43 Genomes in Yeast via Transformation-Associated Recombination
Brett A. Duguay and Craig McCormick
Aug 20, 2025 3038 Views
Abstract
Interferon regulatory transcription factor 3 (IRF3) is a transcription factor that upon activation by virus infection promotes the synthesis of antiviral genes, such as the interferons (Hiscott, 2007). In addition to inducing genes, IRF3 triggers antiviral apoptosis by RIG-I-like receptor-induced IRF3 mediated pathway of apoptosis (RIPA), which is independent of its transcriptional activity. RIPA protects against lethal virus infection in cells and mice (Chattopadhyay et al., 2016). In the absence of RIPA, caused by genetic ablation, chemical mutagenesis or inhibition of the pattern recognition receptor (PRR) retinoic acid-inducible gene I (RIG-I), Sendai virus (SeV) infection does not trigger cellular apoptosis and become persistently infected (Peters et al., 2008; Chattopadhyay et al., 2013). IRF3-expressing wild type (WT) cells (U4C) undergo SeV-induced apoptosis; however, the P2.1 cells, which are deficient in IRF3 expression are not capable of triggering viral apoptosis (Figure 1). Ectopic expression of human IRF3 restores the apoptotic activity in P2.1 cells (P2.1/IRF3, Figure 1). SeV is used as a model for studying pathogenic human viruses, which are difficult to work with or require BSL3 facility. We have previously reported that both human and mouse cells can establish SeV persistence in the absence of IRF3’s apoptotic activity (Chattopadhyay et al., 2013). Here, we outline a detailed procedure for the development of a persistently SeV-infected human cell line (Figure 2), which continuously expresses viral protein and produces low levels of infectious viral particles.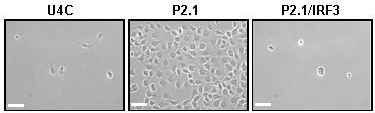
Figure 1. SeV-induced apoptosis is IRF3-dependent. HT1080-derived cell lines (U4C, P2.1 and P2.1/IRF3) were infected with Sendai virus and three days post infection culture fields were photographed, scale bar represents 50 µm.
Background
IRF3 is essential for initiating antiviral defense mechanisms in host cells by way of promoting transcription of antiviral genes (Hiscott, 2007; Chattopadhyay and Sen, 2017). Upon recognition of viral dsRNA by PRRs in the cell, IRF3 becomes phosphorylated, dimerizes, and translocates to the nucleus, where it binds to the interferon-sensitive response element (ISRE), and promotes transcription of type-1 interferons, e.g., IFN-β. IRF3 is also critical for triggering apoptosis via a distinct pathway, which does not require its transcriptional activity. In a series of previous studies, we have discovered the pathway, which we named RIPA that triggers apoptosis in virus-infected cells. In RIPA, IRF3 interacts with BCL-2-Associated X protein (BAX), a pro-apoptotic factor (Chattopadhyay et al., 2010). Upon binding to BAX, IRF3 translocates to the mitochondria, and initiates a signaling cascade that ultimately promotes apoptosis (Chattopadhyay et al., 2010). In the absence of IRF3 or other components of RIPA, the cells establish viral persistence when infected with Sendai virus (SeV) (Peters et al., 2008; Chattopadhyay et al., 2013). These persistent cell lines are useful for studying the full anti-viral mechanisms of cells because the cells do not undergo apoptotic cell death. In the current protocol, we provide a detailed method to create a SeV persistent human cell line, which are defective in IRF3 expression. Viral persistence is common for many viruses, which efficiently antagonize the cell death pathways of the infected cells. An in vitro approach to study persistently infected cells will reveal ways to avoid the establishment of viral persistence. It will also be evaluated in future whether the absence of RIPA can be used as a tool to generate persistently infected cells using viruses of different lifestyles.
Materials and Reagents
- Materials
- Pipette tips (USA Scientific)
- 6 cm tissue culture plates (USA Scientific, Cyto-One, catalog number: CC7672-3359 )
- Cryovials (USA Scientific, catalog number: 1412-9100 )
- 1.5 ml Eppendorf tubes (USA Scientific)
- PVDF membrane (Bio-Rad Laboratories, catalog number: 1620177 )
- Autoradiography film (Denville Scientific, catalog number: E3012 )
- 6-well plate
- Pipette tips (USA Scientific)
- Cells
- U4C cells: these cells were generated from HT1080 cells and are deficient in IFN signaling
Note: These cells are maintained in DMEM containing 10% FBS, 100 international units of penicillin, 100 µg/ml streptomycin (complete DMEM). - P2.1 cells: these cells were generated from U4C cells and are deficient in IRF3 expression
Note: These cells are maintained in DMEM containing 10% FBS, 100 µg/ml penicillin, 100 µg/ml streptomycin (complete DMEM). - P2.1/IRF3 cells: these cells were generated by stably expressing human IRF3 in P2.1 cells and were selected under puromycin (1 µg/ml)
Note: These cells are maintained in DMEM containing 10% FBS, 100 µg/ml penicillin, 100 µg/ml streptomycin and puromycin (1 µg/ml). - LLC-MK2 cells (ATCC, catalog number: CCL-7 ):
Note: These cells are maintained in Medium 199 containing 10% FBS, 100 µg/ml penicillin, 100 µg/ml streptomycin.
Note: U4C, P2.1 and P2.1/IRF3 cells were generated in the authors’ laboratory and were described previously. See Chattopadhyay et al., 2010 and 2016. These cell lines are available from the authors upon request. - U4C cells: these cells were generated from HT1080 cells and are deficient in IFN signaling
- Viruses
Sendai virus (SeV) Cantell strain (Charles River laboratories)–this strain was originally obtained from ATCC (ATCC, catalog number: VR-907 ) - Reagents
- Dulbecco’s modified Eagle’s medium (DMEM) (Lerner Research Institute Central Cell Services, catalog number: 11-500p )
- Fetal bovine serum (FBS) (Atlanta Biologicals, catalog number: S11550 )
- Complete EDTA-free protease inhibitor (Roche Diagnostics, catalog number: 11873580001 )
- Cryoprotective medium (Lonza, catalog number: 12-132A )
- Phosphate-buffered saline (PBS) (Lerner Research Institute Central Cell Services, catalog number: 123-1000p )
- SDS-PAGE loading buffer (Laemelli) (Bio-Rad Laboratories, catalog number: 1610737 )
- 10x SDS-PAGE running buffer (AMRESCO, catalog number: 0783 )
- Protein assay dye (Bio-Rad Laboratories, catalog number: 5000006 )
- 10x transfer buffer (AMRESCO, catalog number: 0307 )
- Nonfat dry milk (Bio-Rad Laboratories, catalog number: 1706404XTU )
- Tris buffered saline with Tween-20 (TBST) (AMRESCO, catalog number: K873 )
- Antibodies
- Sendai Virus C antibody (generated in author’s laboratory) (Chattopadhyay, 2016)
Note: Another anti-SeV antibody from Abcam, catalog number: ab33988 may be used to detect the presence of SeV. - Goat-anti-rabbit secondary antibody conjugated with horseradish peroxidase was obtained from Rockland Lab (Rockland, catalog number: 611-103-122 )
- Sendai Virus C antibody (generated in author’s laboratory) (Chattopadhyay, 2016)
- ECL plus solution (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 80196 )
- Medium 199 (Thermo Fisher Scientific, GibcoTM, catalog number: 11150059 )
- Agar (BD, DifcoTM, catalog number: 214050 )
- Guinea pig red blood cells (Colorado Serum Company, catalog number: 30100 )
- Tris buffer (pH 7.4) (Affymetrix, catalog number: 22639 )
- 5 M sodium chloride solution (NaCl) (Affymetrix, USB, catalog number: 75888 )
- Triton X-100 (Sigma-Aldrich, catalog number: T9284 )
- Sodium orthovanadate (Sigma-Aldrich, catalog number: S6508-10G )
- Sodium fluoride (Sigma-Aldrich, catalog number: S6521 )
Note: This product has been discontinued. - β-Glycerophosphate disodium salt hydrate (Sigma-Aldrich, catalog number: G9422 )
- Sodium pyrophosphate (Sigma-Aldrich, catalog number: S6422 )
- Cell lysis buffer (see Recipes)
- Dulbecco’s modified Eagle’s medium (DMEM) (Lerner Research Institute Central Cell Services, catalog number: 11-500p )
Equipment
- Micropipettes (10 μl, 200 μl, 1,000 μl) (Eppendorf)
- SDS-PAGE and transfer apparatus (Bio-Rad Laboratories, model: Mini PROTEAN-II )
Note: This equipment is no longer available at manufacturer (also use the same machine for agarose gel). - Vortex (Thermo Fisher Scientific)
- Tissue culture incubator (at 37 °C with 5% CO2) (Thermo Fisher Scientific)
- Biosafety cabinet (Baker SterilGARD)
- Table top centrifuge (Eppendorf)
- Heating block (at 95 °C) (Benchmark)
- Rocker (Benchmark)
- Rotator (Labnet)
- Refrigerator (4 °C) and freezers (-20 °C and -80 °C)
- Autoradiography film processor (Kodak)
- Spectrophotometer (Benchmark)
- Liquid Nitrogen Tank Cryosafe CM2 (D.A.I. Scientific Equipment)
Procedure
- Infection of P2.1 cells with SeV
- Seed 500,000 P2.1 cells into 6 cm tissue culture dishes (Figure 2).
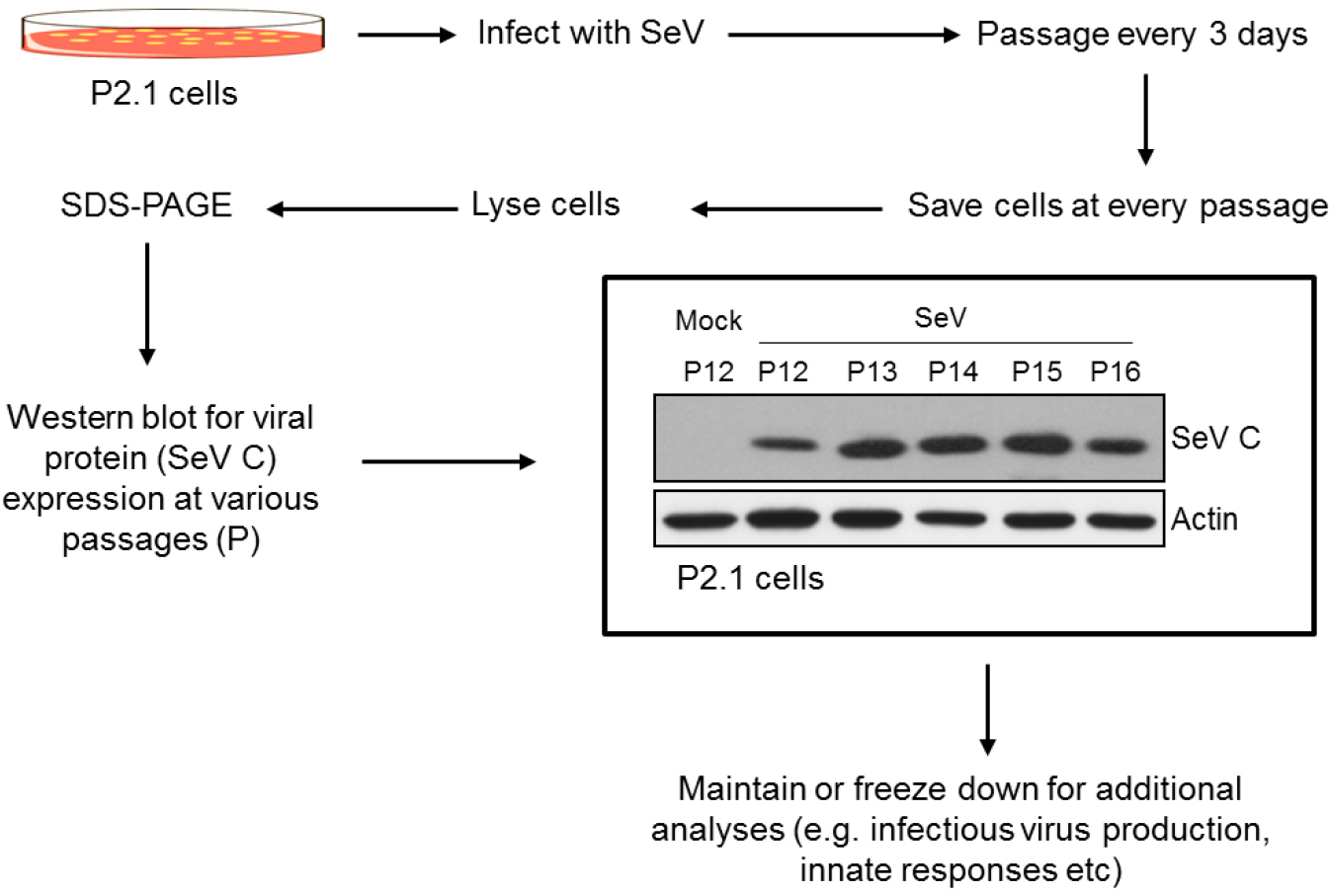
Figure 2. Development of SeV-persistent cell lines using P2.1 parental cells. A flow chart of generation and maintenance of SeV-persistently infected cells using P2.1 cell line. The Western blot shows the consistent levels of viral protein (SeV C) expression at various passage numbers (indicated by P), and actin was used as a loading control. - After cells have adhered (about 16-18 h post seeding), check for 80% confluence, wash cells two times with DMEM containing 2% FBS (virus infection medium), then add a minimum amount (1 ml) of virus infection medium.
- Add SeV at a concentration of 80 hemagglutinating units/ml. Incubate the cells with virus for 1 h with gentle physical agitation every 10 min.
- Remove the virus-containing medium, and wash cells two times with DMEM containing 10% FBS.
- Continue culturing cells in DMEM containing 10% FBS, passaging cells every 3 days, setting aside 10% for detection of SeV and 50% for freezing.
- After each passage, reserve some of the cells for future analysis.
- Combine 0.5 ml cell suspension in DMEM containing 10% FBS with 0.5 ml cryoprotective medium in a labeled cryovial.
- Immediately place in a freezer kept at -80 °C.
- For long-term storage, store cells in liquid nitrogen tank.
- Combine 0.5 ml cell suspension in DMEM containing 10% FBS with 0.5 ml cryoprotective medium in a labeled cryovial.
- To determine the presence of virus, harvested cells are analyzed by Western blot for viral protein detection, as described below.
- Seed 500,000 P2.1 cells into 6 cm tissue culture dishes (Figure 2).
- SDS-PAGE and Western blot to confirm SeV infection
- Seed P2.1 cells and SeV infected P2.1 cells into separate 6 cm tissue culture dishes.
- After 24 h or when the cells are confluent, harvest cell lysates.
- Remove media, and wash the plates once with cold PBS.
- Lyse cells in 50 µl of cold lysis buffer (see Recipes). Gently vortex and keep cell lysates on ice for 30 min.
- Centrifuge cell lysates at 16,200 x g (Rcf) for 20 min in a cold (4 °C) centrifuge.
- Remove media, and wash the plates once with cold PBS.
- Load equal amounts of protein on a 20% SDS-PAGE gel.
Note: Prior to loading, combine cell lysates with equal volumes of 2x SDS-PAGE loading buffer. - Electrophorese the proteins at 100 V for 2 h, or until adequate separation of proteins in the 10-100 kDa region is achieved.
- Transfer proteins onto a PVDF membrane using a Bio-Rad semi-dry transfer apparatus at 0.06 mA for 1.5 h.
- Block the membrane in TBST containing 5% nonfat dry milk on a rocker at room temperature for 30 min.
- Add the anti-Sendai Virus C antibody at 1:5,000 dilution in blocking buffer. Incubate on a rocker at room temperature for 1 h or 12-16 h at 4 °C.
- Wash the membrane three times in TBST, and add the secondary (HRP-conjugated) antibody, diluted to 1:5,000 dilution in blocking buffer, incubating for 1 h at room temperature.
- Wash the membrane three times in TBST and incubate in ECL solution for 1-3 min.
- Develop on autoradiograph film using an autoradiograph film processor.
- Seed P2.1 cells and SeV infected P2.1 cells into separate 6 cm tissue culture dishes.
- Quantification of infectious virus particles in persistently infected cells
The infectious SeV titers in the culture supernatants were determined by previously described procedure (Peters et al., 2008) and was represented as pfu/ml. Briefly, the following procedure is performed:- Seed LLC-MK2 cells in 6-well plate at 100,000 cells per well in Medium 199 and allow the cells to become confluent (in about three days).
- The culture supernatants from the persistently infected cell lines are serial diluted and then used to infect confluent LLC-MK2 monolayer, using the virus infection protocol (as described in Procedure A).
- After infection, the cells are washed and overlaid with Medium 199 containing 0.5% agar.
- After 3 days, the agar layer is removed and the cells are washed with PBS.
- Virus colonies are visualized by incubating the monolayer with a 0.1% suspension of guinea pig red blood cells for 20-30 min. The monolayer is then washed with PBS and the hemabsorbed plaques are counted.
- Seed LLC-MK2 cells in 6-well plate at 100,000 cells per well in Medium 199 and allow the cells to become confluent (in about three days).
Data analysis
To confirm that viral persistence has been achieved, a few relatively simple assays may be performed. Sendai virus expresses a non-structural protein (C), also known as SeV C, which can be detected by Western blotting of the persistently infected P2.1 cells, the procedure for which is outlined in the previous section. To confirm the presence, or rather, the absence of clearance of SeV in P2.1 cells, an anti-SeV C antibody can be applied to the Western blot, which, if a band of approximately 25 kDa is elucidated, can confirm persistence (Figure 2).
Notes
Western blot for detection of SeV C protein should be carefully evaluated because the antibody also cross reacts with other viral and cellular proteins which are of similar sizes. Careful separation of these proteins on a 20% SDS gels will facilitate the analyses (Figure 3).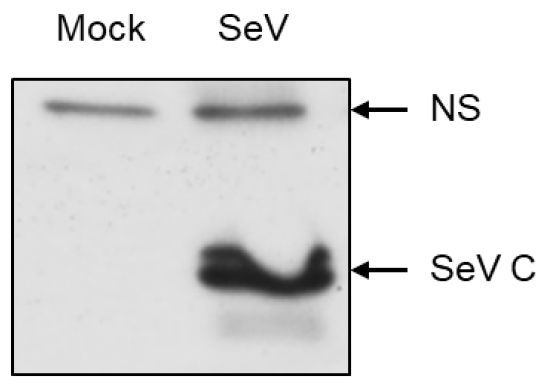
Figure 3. A representative Western blot for SeV C protein expression. Human cells either mock infected or infected with SeV for 16 h, when cell lysates were analyzed by Western blot. NS, non specific.
Recipes
- Cell lysis buffer
50 mM Tris buffer, pH 7.4
150 mM NaCl
0.1% Triton X-100
1 mM sodium orthovanadate
10 mM sodium fluoride
10 mM β-glycerophosphate
5 mM sodium pyrophosphate
Acknowledgments
Our work was supported by the American Heart Association Scientist Development grants 15SDG25090212 (SC) and The University of Toledo College of Medicine and Life Sciences startup funds (SC). The protocol and some representative results were adapted from our previous work (Peters et al., 2008).
References
- Chattopadhyay, S., Fensterl, V., Zhang, Y., Veleeparambil, M., Yamashita, M. and Sen, G. C. (2013). Role of interferon regulatory factor 3-mediated apoptosis in the establishment and maintenance of persistent infection by Sendai virus. J Virol 87(1): 16-24.
- Chattopadhyay, S., Kuzmanovic, T., Zhang, Y., Wetzel, J. L. and Sen, G. C. (2016). Ubiquitination of the transcription factor IRF-3 activates RIPA, the apoptotic pathway that protects mice from viral pathogenesis. Immunity 44(5): 1151-1161.
- Chattopadhyay, S., Marques, J. T., Yamashita, M., Peters, K. L., Smith, K., Desai, A., Williams, B. R. and Sen, G. C. (2010). Viral apoptosis is induced by IRF-3-mediated activation of Bax. EMBO J 29(10): 1762-1773.
- Chattopadhyay, S. and Sen, G. C. (2017). RIG-I-like receptor-induced IRF3 mediated pathway of apoptosis (RIPA): a new antiviral pathway. Protein Cell 8(3): 165-168.
- Hiscott, J. (2007). Triggering the innate antiviral response through IRF-3 activation. J Biol Chem 282(21): 15325-15329.
- Peters, K., Chattopadhyay, S. and Sen, G. C. (2008). IRF-3 activation by Sendai virus infection is required for cellular apoptosis and avoidance of persistence. J Virol 82(7): 3500-3508.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Coakley, C., Peter, C., Fabry, S. and Chattopadhyay, S. (2017). Establishment of a Human Cell Line Persistently Infected with Sendai Virus. Bio-protocol 7(16): e2512. DOI: 10.21769/BioProtoc.2512.
Category
Immunology > Host defense > Human
Microbiology > Microbe-host interactions > Virus
Molecular Biology > Protein > Anti-microbial analysis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









