- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Tandem Affinity Purification in Drosophila Heads and Ovaries
Published: Vol 2, Iss 15, Aug 5, 2012 DOI: 10.21769/BioProtoc.245 Views: 14271

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
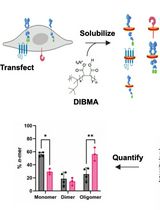
SiMPull-POP: Quantification of Membrane Protein Assembly via Single Molecule Photobleaching
Ryan J. Schuck [...] Rajan Lamichhane
Jan 5, 2026 303 Views
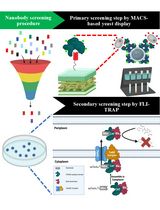
Isolation of Antigen-Specific Nanobodies From Synthetic Libraries Using a Protein Selection Strategy That Combines MACS-Based Screening of YSD and FLI-TRAP
Apisitt Thaiprayoon [...] Dujduan Waraho-Zhmayev
Jan 20, 2026 473 Views
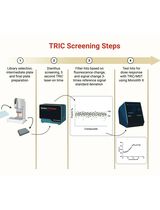
Orthogonal Temperature-Related Intensity Change and Time-Resolved Förster Resonance Energy Transfer High-Throughput Screening Platform for the Discovery of SLIT2 Binders
Moustafa T. Gabr [...] Somaya A. Abdel-Rahman
Feb 20, 2026 97 Views
Abstract
Tandem affinity purification (TAP) (Pugi et al.,2001; Rigaut et al., 1999) is a method that uses a tagging approach of a target protein of interest for a two-step purification scheme in order to pull down protein complexes under native conditions and expression levels. The TAP tag consists of three components: a calmodulin-binding peptide, a Tobacco etch virus (TEV) protease cleavage site and Protein A which is an immunoglobulin G (IgG)-binding domain. This protocol was modified from the original methodology used in yeast cells(Pugi et al.,2001; Rigaut et al., 1999) for isolation of protein complexes from Drosophila heads and ovaries expressing a TAP tagged protein of interest. To determine in vivo binding partners of the Drosophila fragile X protein (dFMR1), we developed a transgenic strain of flies expressing a recombinant form of dFMR1 with a carboxy-terminal TAP tag (Tsai and Carstens, 2006). To ensure that the construct was expressed at wild-type levels, we engineered this form of the tagged protein in the context of a genomic rescue construct that rescued a mutant sterility phenotype. The purification process was performed using mild conditions to maintain native protein interactions. For TAP methods in Drosophila S2 cell culture, we have successfully used a protocol previously published by Tsai and Carstens (Tsai and Carstens, 2006; Bhogal et al., 2011).
Materials and Reagents
- 1x Phosphate buffered saline (PBS)
- Fly sieves: 106, 355, 600 and 850 mm
- IgG Sepharose Fast Flow Beads (GE Healthcare Dharmacon, catalog number: 17-0969-01 )
- AcTEV Protease (Life Technologies, Invitrogen™, catalog number: 12575-015 )
- Calmodulin Sepharose 4B (GE Healthcare Dharmacon, catalog number: 17-0529-01 )
- Micro Bio-Spin column (Bio-Rad Laboratories, catalog number: 732-6204 )
- NuPAGE Novex 4-12% Bis-Tris gel (Life Technologies, catalog number: NP0321 )
- Silverquest Silver Staining Kit (Life Technologies, catalog number: LC6070 ), Colloidal Blue Staining Kit (Life Technologies, catalog number: LC6025 ) or Coomassie Blue.
- Beta-mercaptoethanol
- 1 M CaCl2
- Hepes
- MgCl2
- KCl
- NP-40
- DTT
- Na3VO4
- EDTA
- EGTA
- NaF
- Glycerol
- Mg-acetate
- Imidazole
- Bouwmeester's buffer (see Recipes)
- Buffer A (see Recipes)
- Buffer B (see Recipes)
- Buffer C (see Recipes)
- TEV Cleavage buffer (see Recipes)
- Calmodulin binding buffer (see Recipes)
- Calmodulin blution buffer (see Recipes)
Equipment
- Benchtop centrifuge (that fits 1.5 ml microcentrifuge tubes)
- 1.5 ml microcentrifuge tubes
- Nutator
- Vortexer
- Ceramic mortar and pestle
- Dounce homogenizer
- Disposable pestles for 1.5 ml tubes
- Beckman table top centrifuge (that fits 15 ml and 50 ml conical tubes and can be cooled to 4 °C)
- Sorvall with a Beckman JA-20 rotor that is capable to cool to 4 °C
- Mass Spectrometry Facility
Procedure
- Isolation of protein complexes
- For protein complex isolation from drosophila ovaries:
- Feed (1-3 day old) flies expressing your TAP-tagged protein for three days with yeast paste.
- Dissect ovaries in 1x PBS, collect in a 1.5 ml microcentrifuge tube, and freeze at -80 °C. Collect more than 200 pairs of ovaries. Ovaries can be collected following several independent rounds of dissection and frozen to amass an appropriate amount.
- Homogenize ovaries in 1 ml Bouwmeester's buffer using disposable pestles that fit 1.5 ml microcentrifuge tube.
- Incubate on ice for 20 min and transfer to Beckman centrifuge tubes (1/2 x 2 inch).
- Spin tubes at 4 °C, 20,000 x g for 15 min.
- Remove and save supernatant on ice. This is your cytoplasmic extract.
- Spin tubes at 4 °C, 100,000 x g for 45 min to remove remaining extract following additional spin.
- Remove and save supernatant on ice. This is your nuclear extract (can combine the extracts from step A6 and A8, if you want total extracts, or keep separate).
- Take out 20 μl from each of your isolated extracts as a lysate control (control A). Freeze in liquid nitrogen and store at -80 °C. Go to step #a in Section 2: Pulldown of protein complexes.
- For protein complex isolation from drosophila heads:
- Freeze flies expressing TAP-tagged protein of interest at –80 °C. You will need at least 50 ml of frozen flies to begin. Weigh starting materials on a standard scale and be sure to have at least 1 gram of heads.
- Chill sieves in liquid nitrogen in preparation for head popping procedure.
- Pop heads off using liquid nitrogen and the proper sized meshes (http://cmgm.stanford.edu/~vesicle/protocols/droshead.html). Keep purified head samples on dry ice or at -80 °C until you are ready for the next step.
- Pre-freeze mortar and pestle using liquid nitrogen. Add heads and liquid nitrogen to chilled mortar and pestle and grind sample on dry ice, continuing to add liquid nitrogen until you have a fine powder.
- When you have fine powder, add more liquid nitrogen to pour out powder into a 50 ml falcon tube.
- Let liquid nitrogen de-gas in uncovered tube for 5-10 min.
- Add 2 ml bouwmeester buffer to powder, transfer to a dounce homogenizer and dounce homogenize 10 times. Transfer to several 1.5 ml microcentrifuge tubes. Spin 15 min, 15,000 x g at 4 °C, remove supernatant and repeat spin to remove as much as supernatant as possible.
- Take out 20 μl as a lysate control from each sample (control A). Snap freeze in liquid nitrogen and store at -80 °C.
- For protein complex isolation from drosophila ovaries:
- Pulldown of protein complexes
- Prepare 200 μl IgG Sepharose Fast Flow beads in a 15 ml falcon tube and wash twice with 500 μl buffer B (add 100 μl PIC and 5 μl DTT fresh to 10 ml buffer B). After each wash with buffer B, spin at 4 °C, 100 x g for 2 min in a Beckman table top centrifuge and remove supernatant.
- Add lysate and rotate overnight at 4 °C on a nutator.
- After overnight incubation, spin at 4 °C, 100 x g for 5 min in a Beckman table top centrifuge and remove supernatant. Save 20 μl of supernatant for post-incubation control (control B).
- Wash beads 3 times in 1 ml Buffer B. Spin 4 °C, 100 x g for 2 min after each wash in a Beckman table top centrifuge.
- Wash beads 4 times in 1 ml Buffer C. Spin 4°C, 100 x g for 2 min after each wash in a Beckman table top centrifuge.
- Wash twice in 1 ml TEV cleavage buffer. Spin 4 °C, 100 x g for 2 min after each wash in a Beckman table top centrifuge. Remove 10 μl beads as a control (control C).
- Add 400 μl TEV cleavage buffer and 4 μl AcTEV protease to beads and transfer to a 1.5 ml microcentrifuge tube. Rotate overnight at 4 °C on a nutator.
- Spin 300 x g, 1 min at 4 °C in a bench top centrifuge.
- Take off supernatant and add 0.6 μl 1 M CaCl2 per 200 μl supernatant (usually 1.2 μl for 400 μl total supernatant).
- Prepare a 1.5 ml microcentrifuge tube with 100 μl Calmodulin sepharose resin and wash twice with 500 μl Calmodulin binding buffer. Spin 2 min, 1,500 rpm, 4 °C after each wash.
- Add 3x volume Calmodulin binding buffer to supernatant and add this to prepared calmodulin resin. Rotate 3-4 h at 4 °C.
- Spin 2 min, 1,500 rpm, 4 °C.
- Wash three times with 500 μl Calmodulin binding buffer. Spin 2 min, 1,500 rpm, 4 °C after each wash.
- Elute TAP-tagged protein with Calmodulin elution buffer using preferred strategy.
- Run eluate on a 4-12% Bis-Tris 10 well gel.
- Run controls and protein ladder on a separate gel from final eluate so there is no contamination.
- Stain with either Coomassie Blue, Colloidal Blue Staining Kit, or Silverquest Kit. Cut out bands of interest and send to the Mass Spectrometry Facility.
Recipes
Note: Make buffers without Protease Inhibitor Cocktail without EDTA (PIC w/o EDTA), Na3VO4, CaCl2 and/or DTT. These should be added fresh right before use.
- Buffer A (20 ml)
See note aboveStock and concentration volume Final concentration 1 M Hepes (pH 7.9) 200 ml 10 mM Hepes 1 M MgCl2 30 ml 1.5 mM MgCl2 2 M KCl 100 ml 10 mM KCl 1 M DTT 10 ml 0.5 mM DTT PIC w/o EDTA 20 ml 1x H2O to 20 ml - Bouwmeester's buffer (20 ml)
Filter Sterilize and store at 4 °C.
See note aboveStock and concentration volume Final concentration 1 M Tris-HCl (pH 7.5) 1 ml 50 mM Tris-HCl 5 M NaCl 2.5 ml 125 mM NaCl 100% glycerol 1 ml 5% glycerol 100% NP-40 80 ml 0.2% NP-40 1 M MgCl2 30 ml 1.5 mM MgCl2 1 M DTT 20 ml 1 mM DTT 1 M NaF 500 ml 25 mM NaF 200 mM Na3VO4 100 ml 1 mM Na3VO4 500 mM EDTA 40 ml 1 mM EDTA 2 mM EGTA 80 ml 500 mM EGTA PIC w/o EDTA 20 ml 1x H2O to 20 ml - Buffer B (20 ml)
Make fresh on the day of experiment
See note aboveStock and concentration volume Final concentration 1 M Hepes (pH 7.9) 400 ml 20 mM Hepes 87% glycerol 4.6 ml 20% glycerol 100% NP-40 200 ml 0.5% NP-40 2 M KCl 2.5 ml 200 mM KCl 1 M DTT 10 ml 0.5 mM DTT 500 mM EDTA 40 ml 1 mM EDTA 20 mM EGTA 80 ml 500 mM EGTA PIC w/o EDTA 200 ml 1x H2O to 20 ml - Buffer C (20 ml)
See note aboveStock and concentration volume Final concentration 1 M Hepes (pH 7.9) 400 ml 20 mM Hepes 87% glycerol 4.6 ml 20% glycerol 100% NP-40 200 ml 0.5% NP-40 2 M KCl 2.5 ml 200 mM KCl 1 M DTT 10 ml 0.5 mM DTT H2O to 20 ml - TEV cleavage buffer (100 ml)
Filter sterilize
See note aboveStock and
concentrationvolume Final concentration 1 M Tris-HCl (pH 8.0) 1 ml 10 mM Tris-HCl 5 M NaCl 3 ml 150 mM NaCl 20% NP-40 500 ml 0.1% NP-40 1 M DTT 50 ml 0.5 mM EDTA H2O to 100 ml - Calmodulin binding buffer (100 ml)
Stock and
concentrationVolume Final concentration 1 M Tris-HCl (pH 8.0) 1 ml 10 mM Tris-HCl Beta-mercaptoethanol 69.7 ml 10 mM
Beta-mercaptoethanol5 M NaCl 3 ml 150 mM NaCl 1 M Mg-acetate 100 ml 1 mM Mg-acetate 1 M Imidazole 100 ml 1 mM Imidazole 1 M CaCl2 200 ml 2 mM CaCl2 20% NP-40 500 ml 0.1% NP-40 H2O to 100 ml - Calmodulin elution buffer (100 ml)
Stock and
concentrationVolume Final concentration 1 M Tris-HCl (pH 8.0) 1 ml 10 mM Tris-HCl Beta-mercaptoethanol 69.7 ml 10 mM Beta-mercaptoethanol 5 M NaCl 3 ml 150 mM NaCl 1 M Mg-acetate 100 ml 1 mM Mg-acetate 1 M Imidazole 100 ml 1 mM Imidazole 20% NP-40 500 ml 0.1% NP-40 2 mm EGTA 400 ml 500 mM EGTA H2O to 100 ml
Acknowledgments
This protocol was modified from the original methodology used in yeast cells (Pugi et al., 2001; Rigaut et al., 1999).
References
- Bhogal, B., Jepson, J. E., Savva, Y. A., Pepper, A. S., Reenan, R. A. and Jongens, T. A. (2011). Modulation of dADAR-dependent RNA editing by the Drosophila fragile X mental retardation protein. Nat Neurosci 14(12): 1517-1524.
- Puig, O., Caspary, F., Rigaut, G., Rutz, B., Bouveret, E., Bragado-Nilsson, E., Wilm, M. and Seraphin, B. (2001). The tandem affinity purification (TAP) method: a general procedure of protein complex purification. Methods 24(3): 218-229.
- Rigaut, G., Shevchenko, A., Rutz, B., Wilm, M., Mann, M. and Seraphin, B. (1999). A generic protein purification method for protein complex characterization and proteome exploration. Nat Biotechnol 17(10): 1030-1032.
- Tsai, A. and Carstens, R. P. (2006). An optimized protocol for protein purification in cultured mammalian cells using a tandem affinity purification approach. Nat Protoc 1(6): 2820-2827.
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Pepper, A. S., Bhogal, B. and Jongens, T. (2012). Tandem Affinity Purification in Drosophila Heads and Ovaries. Bio-protocol 2(15): e245. DOI: 10.21769/BioProtoc.245.
Category
Developmental Biology > Cell signaling > TAP tag
Biochemistry > Protein > Interaction > Protein-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









