- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Cell Type-specific Metabolic Labeling of Proteins with Azidonorleucine in Drosophila
Published: Vol 7, Iss 14, Jul 20, 2017 DOI: 10.21769/BioProtoc.2397 Views: 8972
Reviewed by: Jyotiska ChaudhuriManish ChamoliRosario Gomez-Garcia

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Click Chemistry (CuAAC) and Detection of Tagged de novo Synthesized Proteins in Drosophila
Kathrin Marter [...] Daniela C. Dieterich
Jan 20, 2019 11684 Views
Evaluation of Translation Rate Through L-azidohomoalanine (AHA) Incorporation and Subsequent Alkyne Fluorophore–Mediated Click Chemistry in Yeast
Mainak Pratim Jha and Koyeli Mapa
Jul 20, 2025 2558 Views
Identification of the Subcompartment-Specific Mitochondrial Proteome by APEX2 Proximity Labeling in Saccharomyces cerevisiae
Lorenz Spänle and Johannes M. Herrmann
Feb 20, 2026 70 Views
Abstract
Advanced mass spectrometry technology has pushed proteomic analyses to the forefront of biological and biomedical research. Limitations of proteomic approaches now often remain with sample preparations rather than with the sensitivity of protein detection. However, deciphering proteomes and their context-dependent dynamics in subgroups of tissue-embedded cells still poses a challenge, which we meet with a detailed version of our recently established protocol for cell-selective and temporally controllable metabolic labeling of proteins in Drosophila. This method is based on targeted expression of a mutated variant of methionyl-tRNA-synthetase, MetRSL262G, which allows for charging methionine tRNAs with the non-canonical amino acid azidonorleucine (ANL) and, thus, for detectable ANL incorporation into nascent polypeptide chains.
Keywords: Metabolic labelingBackground
The protein composition of any given cell is intimately linked to its state of differentiation and functionality. Changes in a cell’s proteome may reflect its response to cell-intrinsic cues or to signals originating from elsewhere inside the respective organism or its environment. In turn they are indicative of the significance of those signaling cues. Deciphering proteomes and their dynamics in a cell type-specific fashion has thus become a main focus in current research, reaching a better understanding of molecular events underlying physiological or pathophysiological processes. Any proteomic approach in this direction, however, is challenged by the heterogeneity of cell types that are interconnected within a tissue or organ of interest. In the brain, for instance, different types of neurons and glial cells form the networks required to control animal or human behavior. Moreover, it is well established that information processing within these networks leading to long-term memory is strictly dependent on de novo protein synthesis and degradation. While this has been exemplified for a number of neuronal proteins (e.g., immediate early gene proteins), it is obvious that proteins that are up- or down-regulated in just a limited number of cells (or even are regulated oppositely in different groups of cells) may easily escape conventional modes of detection, where cellular proteomes are averaged across entire brain areas.
A number of labeling methods for cellular proteomes have been published in the last two decades, e.g., using isotope-coded affinity tags (Gygi et al., 1999) or isobaric tags for relative and absolute quantification (Ross et al., 2004), quantitative proteomic analysis using samples from cells grown in 14N or 15N media (Washburn et al., 2002; MacCoss et al., 2003), and stable isotope labeling by amino acids in cell culture (Ong et al., 2002; Andersen et al., 2005). Moreover puromycin (Schmidt et al., 2009) and non-canonical amino acids, e.g., azidohomoalanine (AHA) or homopropargylglycine, in combination with click chemistry have been used to decipher cellular proteomes (Link et al., 2003; Link and Tirrell, 2003; Beatty et al., 2006; Dieterich et al., 2006; Link et al., 2006; Dieterich et al., 2010). All of these strategies, however, fail to uncover cell-type specific proteomes within tissue or organ samples. Most recently, novel strategies to resolve this issue have been reported for C. elegans and Drosophila (Elliott et al., 2014; Erdmann et al., 2015; Yuet et al., 2015). They have in common the use of either a mutated aminoacyl-tRNA synthetase or an orthogonal aminoacyl-tRNA synthetase/tRNA for tagging of newly synthesized proteins with food-supplied non-canonical amino acids. Specifically, we could show that upon cell type-specific expression of a mutant Methionyl-tRNA synthetase (MetRSL262G) as achieved by employing the well-established Gal4/UAS-system, the non-canonical amino acid ANL can be incorporated into proteins of selectable cell types in living Drosophila larvae and adult flies. An accompanying study by Niehues et al. (2015) used this method to show the causal involvement of mutated glycyl-tRNA synthetase in a model for the neurodegenerative Charcot Marie Tooth disease.
ANL-containing proteins can either be analyzed in protein extracts by using biochemistry and mass spectrometry or can be visualized in situ by fluorescence microscopy (Erdmann et al., 2015; Niehues et al., 2015). For more information see ‘Click Chemistry (CuAAC) and detection of tagged de novo synthesized proteins’. The following protocol details the metabolic labeling of proteins in larvae and adult flies with ANL.
Materials and Reagents
- Fly vials (e.g., VWR, catalog number: 734-2254 )
- Fly vial plugs (e.g., Carl Roth, catalog number: PK13.1 )
- Gal4 activator strains of choice (e.g., C57-Gal4 for muscle-specific expression [from Ulrich Thomas, Magdeburg, Germany], elavC155-Gal4 for pan-neuronal expression [from Bloomington stock center, Bloomington, Indiana, USA], repo-Gal4 for glial expression [from Christian Klämbt, Münster, Germany])
- UAS-dMetRSL262G effector strains [available at request from Daniela C. Dieterich & Ulrich Thomas]. As described in Erdmann et al. (2015) various lines expressing dMetRSL262G either tagged with 3xmyc or EGFP are available
Note: We traditionally use ONM. The standard corn meal medium has also been successfully used in Niehues et al. (2015) for ANL labeling, thus, we anticipate that other media can be used as well without any limitations. - Otto-normal-medium (ONM, see Recipes)
- Agar-Agar (Carl Roth, catalog number: 5210 )
- Semolina (local food store)
- Mashed raisins (local food store)
- Baker’s yeast (local food store)
- Sugar beet sirup (local food store)
- Honey (local food store)
- Tap water
- 20% (w/v) Nipagin (see Recipes)
- 200 mM ANL stock solution (for the synthesis of ANL see [Link et al., 2007; Ngo et al., 2009; Erdmann et al., 2015]) (see Recipes)
Equipment
- Beaker (kitchen/household grade)
- Immersion blender (kitchen/household grade)
- Paintbrush (art supplies)
- Fly incubator (e.g., SANYO, model: MIR-553 )
- Hotplate (kitchen/household grade)
- Pot (kitchen/household grade)
- Tablespoon (kitchen/household grade)
Procedure
- Preparation of ANL-containing fly food medium (Figure 1)
- Thaw baker’s yeast.
- Add semolina and Agar-Agar to 0.33 L water. Stir from time to time until swelling is completed.
- Heat mashed raisins, yeast, sugar beet syrup and honey in 0.66 L water. Boil the mixture for 5 min while stirring constantly.
- Add the semolina-Agar-Agar-mixture and boil once again. Don’t forget to stir constantly, as the mixture might braise at the bottom of the pot.
- Cool down the ONM to 50 °C. Stir every 15-20 min for 1 min.
- Add the Nipagin and stir for at least 1 min until the Nipagin is homogenously distributed in the food.
- Add 2 ml of ANL stock solution to 100 ml ONM in a beaker for a final concentration of 4 mM ANL. Mix for 1 min using an immersion blender.
- Aliquot ANL-containing ONM (2-4 ml) into fly vials and let cool down completely at room temperature.
- Plug the vials.
- ANL-containing ONM can be stored for approximately two weeks at 4 °C. Discard vials once the ONM detaches from the vial wall.
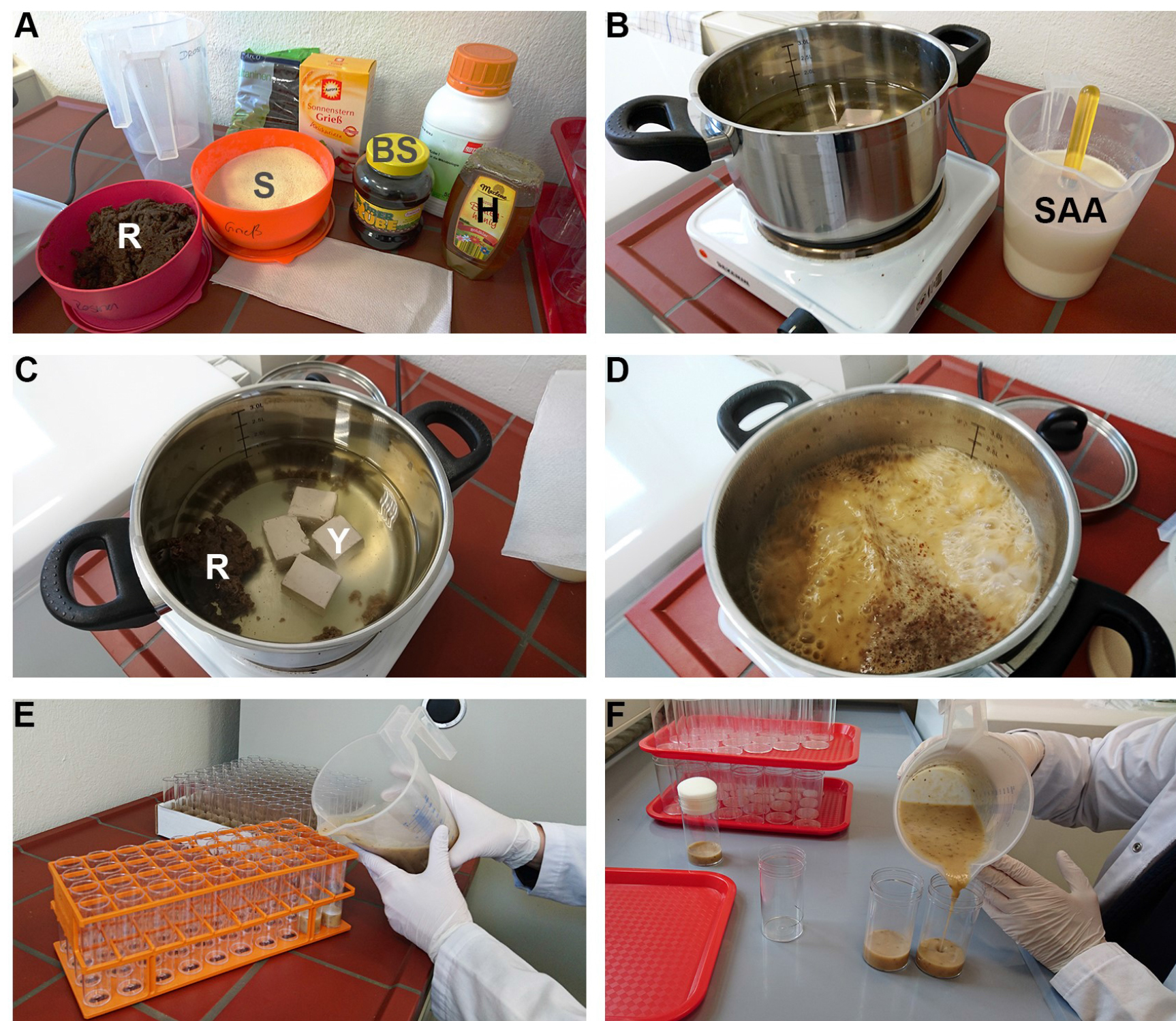

Figure 1. Preparation of ANL-containing fly food medium. A. Ingredients for ONM are shown: R (mashed raisins); S (semolina); BS (sugar beet syrup); H (honey). B. Semolina and Agar-Agar are added to water and allowed to swell, SAA. C. Raisins and Baker’s yeast (Y), sugar beet syrup (BS), and honey (H), are added into water and boiled for 5 min. D. Afterwards, the SAA mixture is added and boiled once more. After cooling down to 50 °C with stirring from time to time, Nigapin and ANL are added and mixed thoroughly. E and F. Media is then aliquoted and allowed to cool down completely before storage at 4 °C.
- Thaw baker’s yeast.
- Cell type-specific expression of dMetRSL262G-variants in Drosophila larvae and flies
- Collect virgin female flies of the respective activator strain.
Note: The number of flies and hence the number of offspring depends on your experimental design including the type of analysis (e.g., MS/MS) and the effectiveness and degree of cell selectivity of the Gal4-activator in use. - Cross virgin female flies of the activator strain to male flies of a UAS-dMetRSL262G-effector strain (Erdmann et al., 2015).
Note: Crosses can as well be set up reciprocally, that is, you may use virgin female flies of a UAS-dMetRSL262G-effector strain and cross them to males of your activator strain.
- Collect virgin female flies of the respective activator strain.
- Cell type-specific labeling of proteins with ANL–examples
- Long-term labeling of proteins in larval body wall muscles:
- Raise appropriate crosses on ANL-containing ONM such that ANL is present during all developmental stages of the progeny.
- Mid- to late 3rd larval stage is reached after approximately 5 days when crosses are raised at 25 °C and after approximately 10-12 days when kept at 18 °C.
- Perform dissection of larval body walls according to ‘Click Chemistry (CuAAC) and detection of tagged de novo synthesized proteins’. See also Bellen and Budnik (2000).
Note: Biochemical approaches on whole larval extracts have proven difficult to perform time and again, perhaps due to lytic activities. We therefore recommend separating body walls (mainly comprising muscles, epithelia, cuticle, trachea and sensory neurons) from all other tissues.
- Raise appropriate crosses on ANL-containing ONM such that ANL is present during all developmental stages of the progeny.
- Long-term labeling of proteins in fly heads (e.g., if dMetRSL262G is expressed in the CNS, compound eyes and/or antenna):
- Raise crosses on ANL-containing ONM so that ANL is present throughout development of the progeny.
- Remove parental flies before eclosure of the offspring.
- Prepare fly heads from adult progeny according to ‘Click Chemistry (CuAAC) and detection of tagged de novo synthesized proteins’.
- Raise crosses on ANL-containing ONM so that ANL is present throughout development of the progeny.
- Short-term labeling of proteins in larval body wall muscles or brains:
- Raise crosses on ANL-free ONM for 1-2 days at 25 °C.
- Transfer parental flies onto fresh ANL-free ONM for 4-6 h. Remove parental flies.
- Keep vials for 72 ± 2 h at 25 °C.
- Wash early 3rd instar larvae out of the food with warm tap water and rinse them into a mesh basket.
- Transfer 3rd instar larvae onto ANL-containing ONM using a paintbrush.
- Keep larvae at 25 °C for 24 h.
- Prepare the larval body walls or larval brains according to ‘Click Chemistry (CuAAC) and detection of tagged de novo synthesized proteins’.
- Raise crosses on ANL-free ONM for 1-2 days at 25 °C.
- Short-term labeling of proteins in fly heads:
- Raise appropriate crosses on ANL-free ONM either at 25 °C or 18 °C depending on your experimental design.
- Discard parental flies before progeny ecloses.
- Transfer progeny flies on ANL-containing ONM for a period of time according to your experimental design. For certain analyses it may also be considered to place flies back onto ANL-free food.
- Prepare fly heads according to ‘Click Chemistry (CuAAC) and detection of tagged de novo synthesized proteins’.
- Long-term labeling of proteins in larval body wall muscles:
Data analysis
ANL incorporation into fly or larval proteins can be analyzed after performing copper-catalyzed azide-alkyne cycloaddition (CuAAC, ‘click chemistry’) as described in Erdmann et al. (2015) and the accompanying bio-protocol. General fly and larvae viabilities upon ANL incorporation can be analyzed by assessing e.g., locomotor behavior and/or hatching rates as described in Erdmann et al. (2015).
Notes
- Reproducibility and variability: Uptake of ANL and thus subsequent labeling efficiency may vary when heterogeneous animal numbers are raised in the vials, as the food will be differently mashed through depending on the number of larvae. Also of note, larvae will show in general stronger ANL incorporation compared to adult flies.
- ANL-containing ONM is used best within two weeks after preparation to yield reproducible results.
Recipes
- 20% (w/v) Nipagin
150 g methyl-4-hydroxybenzoate
50 g propyl-4-hydroxybenzoate
Dissolve in 100% (abs) ethanol - Otto-normal-medium (ONM)
8.3 g Agar-Agar
50 g semolina
40 g mashed raisins
2 cubes of yeast ( 42 g each)
1 tablespoon sugar beet syrup
1 tablespoon honey
6.6 ml 20% (w/v) Nipagin - 200 mM ANL stock solution
34.4 mg per 1 ml ddH2O
Acknowledgments
This work was supported by the German Research Foundation (DFG) with an SFB 779 grant to D.C.D., SFB 854 grants to U.T. and D.C.D. and a Leibniz Society PAKT grant (LGS SynaptoGenetics) to D.C.D. and U.T. This protocol was adapted from Erdmann et al., 2015.
References
- Andersen, J. S., Lam, Y. W., Leung, A. K., Ong, S. E., Lyon, C. E., Lamond, A. I. and Mann, M. (2005). Nucleolar proteome dynamics. Nature 433(7021): 77-83.
- Beatty, K. E., Liu, J. C., Xie, F., Dieterich, D. C., Schuman, E. M., Wang, Q. and Tirrell, D. A. (2006). Fluorescence visualization of newly synthesized proteins in mammalian cells. Angew Chem Int Ed Engl 45(44): 7364-7367.
- Bellen, H. J. and Budnik, V. (2000). Drosophila, a laboratory manual. In: Ashburner, M., Hawley, S. and Sullivan. B (Eds). Cold Spring Harbor Laboratory. chap. 11.
- Dieterich, D. C., Hodas, J. J., Gouzer, G., Shadrin, I. Y., Ngo, J. T., Triller, A., Tirrell, D. A. and Schuman, E. M. (2010). In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons. Nat Neurosci 13(7): 897-905.
- Dieterich, D. C., Link, A. J., Graumann, J., Tirrell, D. A. and Schuman, E. M. (2006). Selective identification of newly synthesized proteins in mammalian cells using bioorthogonal noncanonical amino acid tagging (BONCAT). Proc Natl Acad Sci U S A 103(25): 9482-9487.
- Elliott, T. S., Townsley, F. M., Bianco, A., Ernst, R. J., Sachdeva, A., Elsasser, S. J., Davis, L., Lang, K., Pisa, R., Greiss, S., Lilley, K. S. and Chin, J. W. (2014). Proteome labeling and protein identification in specific tissues and at specific developmental stages in an animal. Nat Biotechnol 32(5): 465-472.
- Erdmann, I., Marter, K., Kobler, O., Niehues, S., Abele, J., Muller, A., Bussmann, J., Storkebaum, E., Ziv, T., Thomas, U. and Dieterich, D. C. (2015). Cell-selective labelling of proteomes in Drosophila melanogaster. Nat Commun 6: 7521.
- Gygi, S. P., Rist, B., Gerber, S. A., Turecek, F., Gelb, M. H. and Aebersold, R. (1999). Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat Biotechnol 17(10): 994-999.
- Link, A. J., Mock, M. L. and Tirrell, D. A. (2003). Non-canonical amino acids in protein engineering. Curr Opin Biotechnol 14(6): 603-609.
- Link, A. J. and Tirrell, D. A. (2003). Cell surface labeling of Escherichia coli via copper(I)-catalyzed [3+2] cycloaddition. J Am Chem Soc 125(37): 11164-11165.
- Link, A. J., Vink, M. K., Agard, N. J., Prescher, J. A., Bertozzi, C. R. and Tirrell, D. A. (2006). Discovery of aminoacyl-tRNA synthetase activity through cell-surface display of noncanonical amino acids. Proc Natl Acad Sci U S A 103(27): 10180-10185.
- Link, A. J., Vink, M. K. and Tirrell, D. A. (2007). Synthesis of the functionalizable methionine surrogate azidohomoalanine using Boc-homoserine as precursor. Nat Protoc 2(8): 1884-1887.
- MacCoss, M. J., Wu, C. C., Liu, H., Sadygov, R. and Yates, J. R., 3rd (2003). A correlation algorithm for the automated quantitative analysis of shotgun proteomics data. Anal Chem 75(24): 6912-6921.
- Ngo, J. T., Champion, J. A., Mahdavi, A., Tanrikulu, I. C., Beatty, K. E., Connor, R. E., Yoo, T. H., Dieterich, D. C., Schuman, E. M. and Tirrell, D. A. (2009). Cell-selective metabolic labeling of proteins. Nat Chem Biol 5(10): 715-717.
- Niehues, S., Bussmann, J., Steffes, G., Erdmann, I., Kohrer, C., Sun, L., Wagner, M., Schafer, K., Wang, G., Koerdt, S. N., Stum, M., Jaiswal, S., RajBhandary, U. L., Thomas, U., Aberle, H., Burgess, R. W., Yang, X. L., Dieterich, D. and Storkebaum, E. (2015). Impaired protein translation in Drosophila models for Charcot-Marie-Tooth neuropathy caused by mutant tRNA synthetases. Nat Commun 6: 7520.
- Ong, S. E., Blagoev, B., Kratchmarova, I., Kristensen, D. B., Steen, H., Pandey, A. and Mann, M. (2002). Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol Cell Proteomics 1(5): 376-386.
- Ross, P. L., Huang, Y. N., Marchese, J. N., Williamson, B., Parker, K., Hattan, S., Khainovski, N., Pillai, S., Dey, S., Daniels, S., Purkayastha, S., Juhasz, P., Martin, S., Bartlet-Jones, M., He, F., Jacobson, A. and Pappin, D. J. (2004). Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol Cell Proteomics 3(12): 1154-1169.
- Schmidt, E. K., Clavarino, G., Ceppi, M. and Pierre, P. (2009). SUnSET, a nonradioactive method to monitor protein synthesis. Nat Methods 6(4): 275-277.
- Washburn, M. P., Ulaszek, R., Deciu, C., Schieltz, D. M. and Yates, J. R., 3rd (2002). Analysis of quantitative proteomic data generated via multidimensional protein identification technology. Anal Chem 74(7): 1650-1657.
- Yuet, K. P., Doma, M. K., Ngo, J. T., Sweredoski, M. J., Graham, R. L., Moradian, A., Hess, S., Schuman, E. M., Sternberg, P. W. and Tirrell, D. A. (2015). Cell-specific proteomic analysis in Caenorhabditis elegans. Proc Natl Acad Sci U S A 112(9): 2705-2710.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Erdmann, I., Marter, K., Kobler, O., Niehues, S., Bussmann, J., Müller, A., Abele, J., Storkebaum, E., Thomas, U. and Dieterich, D. C. (2017). Cell Type-specific Metabolic Labeling of Proteins with Azidonorleucine in Drosophila. Bio-protocol 7(14): e2397. DOI: 10.21769/BioProtoc.2397.
Category
Biochemistry > Protein > Labeling
Systems Biology > Proteomics > Whole organism
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










