- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Soft Agar Colony Formation Assay as a Hallmark of Carcinogenesis
Published: Vol 7, Iss 12, Jun 20, 2017 DOI: 10.21769/BioProtoc.2351 Views: 30592
Reviewed by: Antoine de MorreeVanesa Olivares-IllanaXiaoyi Zheng

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Isolation of Murine Alveolar Type II Epithelial Cells
Fan Sun [...] Zhaoxia Qu
May 20, 2017 13116 Views

A Fast and Reliable Method to Generate Pure, Single Cell-derived Clones of Mammalian Cells
Zhe Han [...] Varun Kumar
Aug 20, 2022 4105 Views
Abstract
Soft agar colony formation assay is established to estimate the anchorage-independent growth ability of cells. In this assay, a bottom layer of agar with complete media is poured and solidified first, followed by an upper layer containing a specified number of cells suspended in medium-agar mixture. After two weeks of incubation, the number of colonies will be counted, serving as an indicator of malignancy of tumor cells.
Keywords: Anchorage-independent growthBackground
Anchorage-independent growth is an ability of cells to grow independently on a solid surface, and is considered as a hallmark of carcinogenesis (de Larco and Todaro, 1978). Soft agar colony formation assay is a well-established method to evaluate cellular anchorage-independent growth for the detection of the tumorigenic potential of malignant cells (Roberts et al., 1985), which is developed from plate colony formation assay described by Puck et al. in 1956 where cells were seeded on to a culture plate to assay the ability of cells to form colonies (Puck et al., 1956). The limitation of plate colony formation assay is that it only displays cellular abilities for anchorage-dependent growth, by which normal cells can escape from anoikis (a form of programmed cell death that occurs in anchorage-dependent cells when they detach from the surrounding extracellular matrix) and survive (Taddei et al., 2012). In contrast, malignant cells are capable of proliferating and growing without attachment to a substrate. Therefore, soft agar colony formation assay is developed to characterize this ability in vitro (Hamburger and Salmon, 1977; Yuan et al., 2017). The soft agar colony formation assay has been widely adapted for researches on cell differentiation, transformation and tumorigenesis as well as the efficacy evaluation of anti-tumor treatment.
Materials and Reagents
- Cell culture disc (75-cm2) (Corning, catalog number: 430641 )
- Cell culture plate (12-well) (Corning, Costar®, catalog number: 3513 )
- Falcon 15 ml conical centrifuge tubes (Corning, catalog number: 430791 )
- Counting slides (Bio-Rad Laboratories, catalog number: 1450011 )
- 0.1-20 ml volume pipette tips (Eppendorf, catalog number: 22492012 )
- 5-200 ml volume pipette tips (Eppendorf, catalog number: 22492039 )
- 50-1,000 ml volume pipette tips (Eppendorf, catalog number: 22492055 )
- Human SGC7901 cell line (Cell Resource Center of the Chinese Academy of Sciences, catalog number: CC-Y1456 )
- Phosphate buffered saline (PBS) pH 7.4 (Thermo Fisher Scientific, GibcoTM, catalog number: C10010500BT )
- Trypsin-EDTA (0.25%) (Thermo Fisher Scientific, GibcoTM, catalog number: 25200072 )
- RPMI 1640 medium (Thermo Fisher Scientific, GibcoTM, catalog number: C11875500BT )
- Fetal bovine serum (Thermo Fisher Scientific, GibcoTM, catalog number: 10099141 )
- Penicillin-streptomycin (5,000 U/ml) (Thermo Fisher Scientific, GibcoTM, catalog number: 15070063 )
- L-glutamine (Thermo Fisher Scientific, GibcoTM, catalog number: 25030081 )
- Agar (Biowest, catalog number: 111860 )
- Complete 1640 medium (see Recipes)
- 5% agar solution (see Recipes)
Equipment
- 2-20 μl pipettes (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 4641060N )
- 20-200 μl pipettes (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 4641080N )
- 100-1,000 μl pipettes (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 4641100N )
- Clean bench (Thermo Fisher Scientific, Thermo ScientificTM, model: HeraguardTM ECO )
- Autoclave (TOMY DIGITAL BIOLOGY, model: SX-500 )
- Water-Jacketed CO2 incubators (Thermo Fisher Scientific, Thermo ScientificTM, model: FormaTM Series II 3110 , catalog number: 3131)
- Thermostat water bath (Prima Technology, model: YB12 )
- Centrifuge (Eppendorf, model: 5424 R )
- Automated cell counter (Bio-Rad Laboratories, model: TC20TM )
- Advanced microscopy group microscope (Thermo Fisher Scientific, model: EVOS )
- Gel count colony counter (Oxford optronix, model: GelCountTM )
Software
- Statistical Program for Social Sciences 17.0 software (SPSS)
Procedure
- Preparing 5% agar solution (see Recipes).
- Production of the bottom layer of agar.
- Add 9 ml complete medium (37 °C) to 1 ml 5% agar solution (50 °C) and mix thoroughly.
- Pipette 0.8 ml mixture to each well of 12-well plate and allow it to solidify for 30 min at room temperature.
- Preparation of cell suspension.
- Remove the complete medium from culture dish and wash cells with 1x PBS.
- Add 0.5 ml 0.25% trypsin (37 °C) for 3-5 min and collect detacted cells by adding complete medium.
- Spin cells at 60 x g for 5 min and resuspend cells in complete medium, then count cells and adjust the concentration of cells to 1 x 103 cells/ml.
- Production of the upper layer of agar.
- Add 9.4 ml re-suspended cells (37 °C) to 0.6 ml 5% agar solution (50 °C) and mix homogeneously.
- Pipette 0.8 ml the cell-agar mixture onto the solidified bottom layer of agar in 12-well plate and allow it to solidify for 30 min at room temperature.
- Add 800 μl complete medium on top to prevent drying of agar and then cells were maintained in a 37 °C humidified incubator with a mixture of 95% air and 5% CO2.
- Clone counting
- Monitor colony formation for 2-3 weeks before counting.
- All colonies per well were counted with a gel count colony counter, and then determine the average number of colonies of the three replicates for each group.
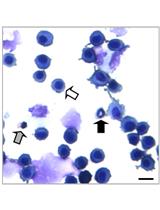
- Capture images of colonies at room temperature using an advanced microscopy group microscope (Figure 1).

Figure 1. Photographs of representative colonies from SGC7901 cell lines. SGC7901 cells were cultured in the upper layer of agar and the formation of colonies was captured at 2-3 weeks after culture. Scale bars: A, 400 μm; B, 100 μm.
Data analysis
All statistical data were analyzed with the Statistical Program for Social Sciences 17.0 software (SPSS). The experiments were conducted in triplicates. The results are presented as the mean ± SD. Differences between means were assessed using Student’s t-test or one-way analysis of variance. P < 0.05 was considered to be statistically significant.
Notes
- This assay was conducted using human SGC7901 cell line and is applicable to other cancer cell lines.
- Incubate cells typically 2-3 weeks and adjust the incubation period according to the tumorgenecity of the cell line.
- After autoclaving, agar solution should be kept sterile during the following operation.
- Pay attention to the temperature of agar solution and complete medium. It is recommended to keep agar solution and complete medium at 50 °C and 37 °C, respectively, and mix them as soon as possible to avoid in-homogenous agglomeration (Puck et al., 1956).
- Do not pour the upper layer of agar until the bottom layer of agar is coagulated completely.
- Details of the instruction of gel count colony counter can be obtained from the website: http://www.oxfordoptronix.com/product17/page501/menu2/Colony_Counting/GelCount_/GelCount_.html
Recipes
- Complete 1640 medium
10% FBS
1% penicillin-streptomycin
2 mM glutamine - 5% agar solution
- Dissolve 5 g agar powder in 100 ml saline and autoclave at 121 °C for 15 min
- Place the sterile 5% agar solution in 50 °C water bath to keep it in liquid phase
Acknowledgments
This work was funded by NSFC grants 81602641 to Dr. Xiaodi Zhao.
References
- de Larco, J. E. and Todaro, G. J. (1978). Growth factors from murine sarcoma virus-transformed cells. Proc Natl Acad Sci U S A 75(8): 4001-4005.
- Hamburger, A. W. and Salmon, S. E. (1977). Primary bioassay of human tumor stem cells. Science 197(4302): 461-463.
- Puck, T. T., Marcus, P. I. and Cieciura, S. J. (1956). Clonal growth of mammalian cells in vitro; growth characteristics of colonies from single HeLa cells with and without a feeder layer. J Exp Med 103(2): 273-283.
- Roberts, A. B., Anzano, M. A., Wakefield, L. M., Roche, N. S., Stern, D. F. and Sporn, M. B. (1985). Type β transforming growth factor: a bifunctional regulator of cellular growth. Proc Natl Acad Sci U S A 82(1): 119-123.
- Taddei, M. L., Giannoni, E., Fiaschi, T. and Chiarugi, P. (2012). Anoikis: an emerging hallmark in health and diseases. J Pathol 226(2): 380-393.
- Yuan, P., He, X. H., Rong, Y. F., Cao, J., Li, Y., Hu, Y. P., Liu, Y., Li, D., Lou, W. and Liu, M. F. (2017). KRAS/NF-κB/YY1/miR-489 signaling axis controls pancreatic cancer metastasis. Cancer Res 77(1): 100-111.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Du, F., Zhao, X. and Fan, D. (2017). Soft Agar Colony Formation Assay as a Hallmark of Carcinogenesis. Bio-protocol 7(12): e2351. DOI: 10.21769/BioProtoc.2351.
- Zhao, X. D., Lu, Y. Y., Guo, H., Xie, H. H., He, L. J., Shen, G. F., Zhou, J. F., Li, T., Hu, S. J., Zhou, L., Han, Y. N., Liang, S. L., Wang, X., Wu, K. C., Shi, Y. Q., Nie, Y. Z. and Fan, D. M. (2015). MicroRNA-7/NF-kappaB signaling regulatory feedback circuit regulates gastric carcinogenesis. J Cell Biol 210(4): 613-627.
Category
Cancer Biology > General technique > Cell biology assays > Cell isolation and culture
Cell Biology > Cell-based analysis > Colony formation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link