- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determining Genome Size from Spores of Seedless Vascular Plants
Published: Vol 7, Iss 11, Jun 5, 2017 DOI: 10.21769/BioProtoc.2322 Views: 8823
Reviewed by: Scott A M McAdamEmily CopeAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Safe DNA-extraction Protocol Suitable for Studying Tree-fungus Interactions
Susanna Keriö [...] Jared M. LeBoldus
Jun 5, 2020 8055 Views
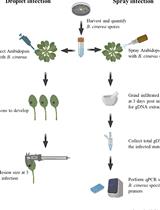
Quantification of Botrytis cinerea Growth in Arabidopsis thaliana
Patricia Scholz [...] Athanas Guzha
Aug 20, 2023 2839 Views

Optimized Protocol for DNA Extraction in Three Theobroma Species
Angie F. Riascos-España [...] Pedro A. Velasquez-Vasconez
May 5, 2025 2075 Views
Abstract
Seedless vascular plants, including ferns and lycophytes, produce spores to initiate the gametophyte stage and to complete sexual reproduction. Approximately 10% of them are apomictic through the production of genomic unreduced spores. Being able to measure the spore nuclear DNA content is therefore important to infer their reproduction mode. Here we present a protocol of spore flow cytometry that allows an efficient determination of the reproductive modes of seedless vascular plants.
Keywords: ApomixisBackground
In seedless vascular plants, sporogenesis features, such as meiotic chromosome counts, were traditionally used to infer nuclear DNA content as well as reproductive modes. However, these approaches are time-consuming, or can only provide indirect evidence. An efficient and reliable method to estimate spore nuclear DNA content of these plants had not been established until Kuo et al. (2017). Herein, we describe a protocol using flow cytometry to evaluate spore genome sizes of these plants based on the work of Kuo et al. (2017).
Materials and Reagents
- Pipette tips (10, 100, and 1,000 μl)
- 50-ml tube
- 1.7-ml tubes with caps
- 2.0-ml tubes with caps
- 2.3-mm stainless steel beads (Bio Spec Products, catalog number: 11079123ss )
- 30-μmnylon meshes (Sysmex, CellTrics®, catalog number: 04-0042-2316 )
- 20-μmnylon meshes (Sysmex, CellTrics®, catalog number: 04-0042-2315 )
- 10-μm nylon meshes (Sysmex, CellTrics®, catalog number: 04-0042-2314 )
- Glass Petri dish (Corning, PYREX®, catalog number: 423790 )
- Leaf tissue of C-value standard (e.g., Nicotiana tabacum L. ‘Xanthi’; 2C = 10.04 pg, Johnston et al., 1999)
- Spores of ferns or lycophytes (kept by dry storage and under < 4 °C)
- PVP-40 (Sigma-Aldrich, catalog number: PVP40 )
- 2-mercaptoethanol (Sigma-Aldrich, catalog number: M3148 )
- RNaseA solution (10 mg/ml in ddH2O) (Sigma-Aldrich, catalog number: R5000-100MG )
- Triton X-100
- Sodium sulfite (Na2SO3)
- Tris-HCl (pH 7.5)
- Propidium iodide
- Backmen stock buffer (see Recipes)*
*Note: LB01 buffer (Doležel et al., 2007) or GPB buffer (Loureiro et al., 2007) can be alternatively used depending on plant material properties. - PI solution (see Recipes)
Equipment
- Pipette (10, 100, and 1,000 μl)
- Vortex (Scientific Industries, model: Vortex-Genie 2 )
- Razors and razor pen
- Flow cytometer (BD, BD Biosciences, model: FACScan )*
*Note: FACScan with a 15 microW blue argon ion laser of an emission wave length of 488 nm.
Software
- BD FACSCan system (BD Biosciences, Franklin Lake, NJ, USA)
Procedure
- Prepare buffer for use
- Allocate appropriate amount of Backmen stock buffer to a 50-ml tube based on an estimation of 1-1.5 ml per sample.
- Add 0.04 g PVP-40, 5 μl 2-mercaptoethanol, 1 μl RNase per ml of buffer.
- Extract spore nuclei by bead-vortexing
- For each sample, weigh ~0.007 g spores into a 1.7-ml tube. Green spores that usually have thin spore walls, 4 times the amount of spores are recommended (Kuo et al., 2017)*.
*Note: For bead-vortexing, the detailed process can be seen in the video supplied in Kuo et al. (2017): http://onlinelibrary.wiley.com/store/10.1111/nph.14291/asset/supinfo/nph14291-sup-0002-VideoS1.mov?v=1&s=dae6f0590d33413f1444bc0add857d285fb73cd6 - Add 16 stainless steel beads into each 1.7-ml tube.
- Add 250 μl of buffer into each 1.7-ml tube.
- Vortex these tubes at a speed of 1,900 rpm for 1 min. For green spores, a speed of 3,200 rpm and a vortex duration of 0.5 min are recommended (Kuo et al., 2017).
- Filter bead-vortexed samples into 2.0-ml tubes through nylon meshes. The size of nylon mesh is selected based on spore sizes to prevent spore being filtered through the mesh.
- Add additional buffer to the samples, and ensure each of filtered spore nuclei solutions is greater than 500 μl in volume.
- Extract standard nuclei by chopping leaf tissue
- Add 500 μl of buffer to a glass Petri dish.
- Add a (~400 mm2) piece of young leaf to the Petri dish, and chop it with a razor on ice until most tissue slices are less than 1 mm in size.
- Filter the chopped sample into a 2.0-ml tube through a 30-μm nylon mesh.
- Add additional buffer to the sample, and ensure that the filtered leaf nuclei solution is greater than 500 μl in volume or more depending on need.
- Staining nuclei solutions
- Mix spore nuclei and standard leaf nuclei solutions into a 500-μl volume in 2.0-ml tubes.
- Add 10 μl PI solution into each of mixed nuclei solutions.
- Incubate in the dark at 4 °C for 1 h for staining.
Data analysis
- Set up a histogram plot of particle count vs. linear value of relative fluorescence in BD FACSCan system.
- After PI staining, measure nuclear DNA content of the samples in BD FACSCan system, and adjust the fluorescence laser voltage to visualize the nuclei peaks on the histogram plot.
- Measure > 1,300 particles for each peak, and the coefficient variation for each peak should be lower 5% as the quality criteria suggested by Greilhuber et al. (2007).
- Genome size of spore nuclei (pg or Mbp) =
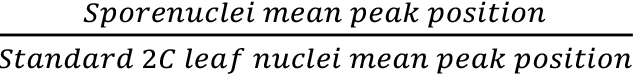 x standard 2C value (Figure 1).
x standard 2C value (Figure 1).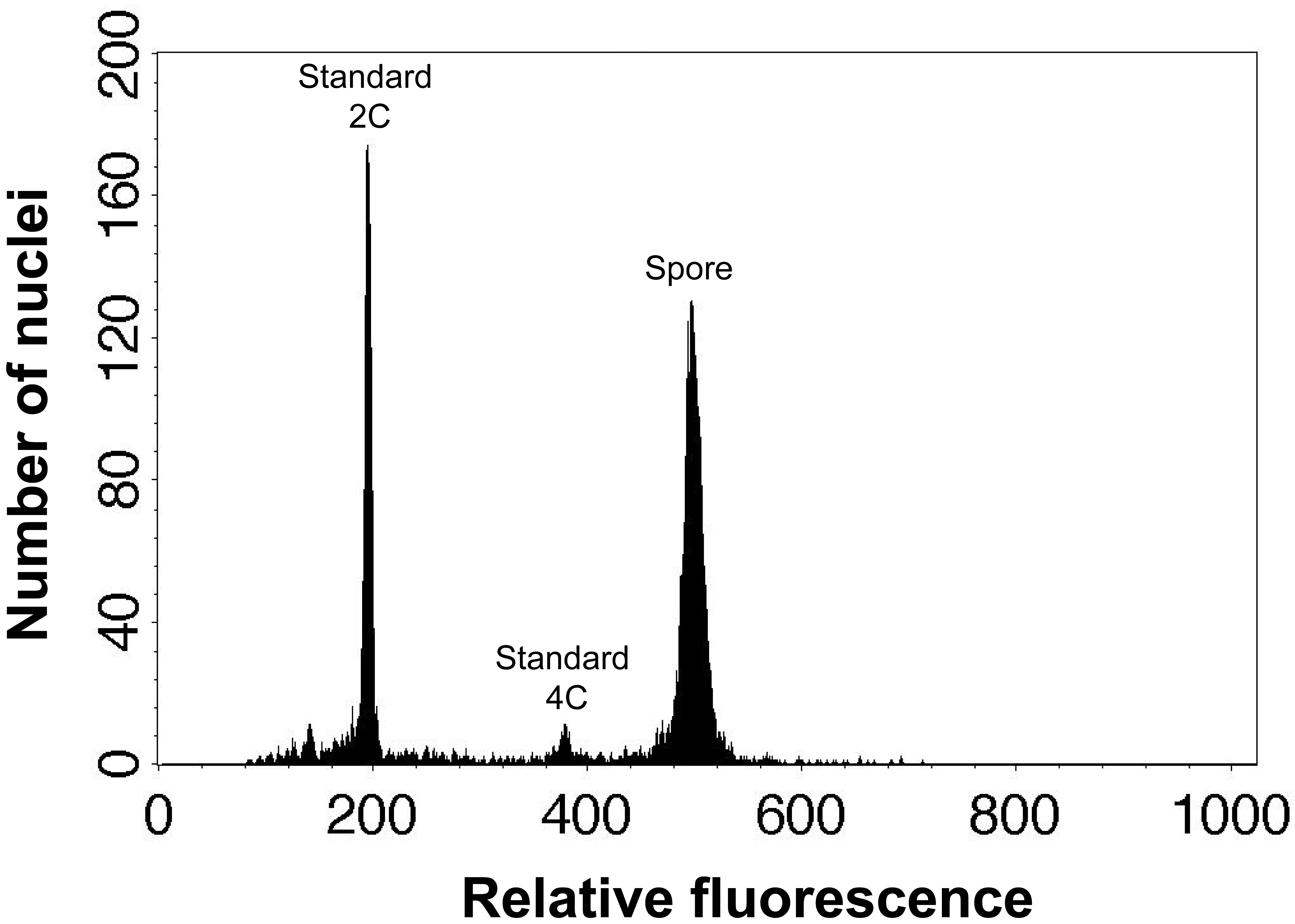
Figure 1. An example of flow cytometric result of spore and standard nuclei. These peaks are spore nuclei of Dryopteris varia and 2C and 4C leaf nuclei of Nicotiana tabacum L. ‘Xanthi’.
Notes
- To dry fertile leaves to release and collect spores, air-drying process under room temperature for 2 to 3 days is recommended. For long-term storage, spore material is better stored in tubes without any solution at a temperature lower than 4 °C.
Recipes
- Backmen stock buffer (Ebihara et al., 2005)
1.0% Triton X-100
50 mM Na2SO3
50 mM Tris-HCl (pH 7.5)
ddH2O (the solvent)
Note: Store at 4 °C up to 1 year. - PI solution
2.04 mg/ml propidium iodide
ddH2O (the solvent)
Note: Store in the dark at 4 °C for long-term storage.
Acknowledgments
The bead-vortexing condition to extract spore nuclei of seedless vascular plants is accessed and constructed by Kuo et al. (2017). We thank Fay-Wei Li and three anonymous reviewers for providing comments on the draft of this manuscript.
References
- Doležel, J., Greilhuber, J. and Suda, J. (2007). Estimation of nuclear DNA content in plants using flow cytometry. Nat Protoc 2(9): 2233-2244.
- Ebihara, A., Ishikawa, H., Matsumoto, S., Lin, S. J., Iwatsuki, K., Takamiya, M., Watano, Y. and Ito, M. (2005). Nuclear DNA, chloroplast DNA, and ploidy analysis clarified biological complexity of the Vandenboschia radicans complex (Hymenophyllaceae) in Japan and adjacent areas. Am J Bot 92(9): 1535-1547.
- Greilhuber, J., Temsch, E. M. and Loureiro, J. C. M. (2007). Nuclear DNA Content Measurement. In: Doleel, J., Greilhuber, J. and Suda, J. (Eds.). Flow cytometry with plant cells: analysis of genes, chromosomes and genomes. Wiley, pp: 67-102.
- Johnston, J., Bennett, M., Rayburn, A., Galbraith, D. and Price, H. (1999). Reference standards for determination of DNA content of plant nuclei. Am J Bot 86(5): 609-613.
- Kuo, L. Y., Huang, Y. J., Chang, J., Chiou, W. L. and Huang, Y. M. (2017). Evaluating the spore genome sizes of ferns and lycophytes: a flow cytometry approach. New Phytol 213(4):1974-1983.
- Loureiro, J., Rodriguez, E., Dolezel, J. and Santos, C. (2007). Two new nuclear isolation buffers for plant DNA flow cytometry: a test with 37 species. Ann Bot 100(4): 875-888.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kuo, L. and Huang, Y. (2017). Determining Genome Size from Spores of Seedless Vascular Plants. Bio-protocol 7(11): e2322. DOI: 10.21769/BioProtoc.2322.
Category
Plant Science > Plant cell biology > Tissue analysis
Plant Science > Plant molecular biology > DNA > DNA extraction
Cell Biology > Single cell analysis > Mass cytometry
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









