- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
RNP-IP (Modified Method)-Getting Majority of RNA from RNA Binding Protein in Cytoplasm
Published: Vol 2, Iss 13, Jul 5, 2012 DOI: 10.21769/BioProtoc.219 Views: 16790

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
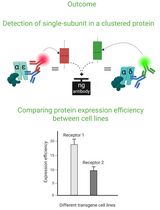
Cluster FLISA—A Method to Compare Protein Expression Efficiency Between Cell Lines and Subunit Clustering of Proteins
Sabrina Brockmöller and Lara Maria Molitor
Nov 5, 2025 1246 Views
Time-Lapse Into Immunofluorescence Imaging Using a Gridded Dish
Nick Lang [...] Andrew D. Stephens
Feb 20, 2026 246 Views
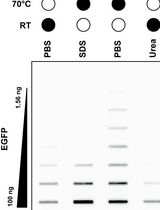
Selective Isolation of TOP3B•mRNA Covalent Intermediates Using Denaturing Oligo-dT Pulldown
Julia E. Warrick and Michael G. Kearse
Mar 5, 2026 130 Views
Abstract
Post-transcriptional regulation of gene expression is a ribonucleoprotein (RNP)-driven process, which involves RNA binding proteins (RBPs) and noncoding RNAs that regulate splicing, nuclear export, subcellular localization, mRNA stability and translation. mRNAs encoding proteins that function in a particular cell process or pathway can be found within a unique mRNP complex, which consists of mRNA and RNP. This provides valuable information regarding not only known components of a particular process or pathway, but importantly, leads to the identification of novel components representing potential therapeutic targets and biomarkers. In addition to those targets identified by pathway expansion, the specific RBPs (RNA binding proteina) regulating RNA functions may be potential therapeutic targets in their own right. RNP-IP is a technology that allows the isolation and identification of mRNAs, microRNAs and protein components of RNP complexes from cell extracts using antibodies to RBPs. Once purified, the RNAs present in the complex are analyzed to identify the target mRNAs using various molecular biology tools such as RT-PCR, gene expression analysis based on microarray technology (chip analysis), or sequencing. Using this modified method will get more RNA existing in cytoplasm. This method does not require a pre-clear step and getting the supernatant for western blot is different from the original method.
Materials and Reagents
- Normal rabbit IgG
- RIP-certified antibody (NBL, catalog number depends on what do you want to target)
- NaCl
- MgCl2
- Nonidet P40
- NaoAc
- Protein A beads (GE Healthcare Dharmacon, catalog number: 17-0780-01 ) or Protein G beads (Thermo Fisher Scientific, catalog number: 22852 )
- Ethanol (molecular biology grade)
- 2-Propanol (Molecular Biology)
- Nuclease-free PBS
- Nuclease-free water
- Isotype control IgG (if necessary)
- Digitornin
- Aprotinin (final concentration 10 μg/ml)
- Leupeptin (5 μg/ml)
- Phenylmethylsulfonyl fluoride (PMSF) (final concentration 0.5 mM)
- RNase inhibitor (Life Technologies, Invitrogen™, catalog number: 10777-019 )
- Dithiothreitol (DTT) (reducing agent)
- Lysis buffer (see Recipes)
- Wash buffer (NT2) (see Recipes)
- Precaution: Additional buffer preparation (see Recipes)
Equipment
- Microcentrifuge capable of 15,000 x g
- Microcentrifuge tube (1.5 ml or 2 ml) (nuclease-free) (recommendation; use low-adhesion tube for RIP-Assay)
- Centrifuge capable of 2,000 x g
- Centrifuge tube (15 ml or 50 ml)
- Pipette (5 ml, 10 ml, 25 ml) (nuclease-free)
- Pipette tip (10 μl, 20-100 μl , 200 μl , and 1,000 μl) (nuclease-free) (recommendation; use low-adhesion pipette tip for RIP-Assay)
- Ultra low temperature freezer (-80 °C)
- Freezer (below -20 °C)
- End-over-end rotator
- Vortex mixer
- Gloves
Procedure
- Preparation of antibody-immobilized protein A or protein G agarose beads.
Day 1- The following components in DEPEC treated water are final concentrations.
50 mM Tris (pH 7.5)
50 mM NaCl
1 mM MgCl2
0.05% Nonidet P40 - Wash beads with NT2 buffer. If you have 4 samples, you need to take 100 μl beads to each Eppendorf tube, total 4 tubes. 3,600 rpm 1 min at 4 °C, remove supernatant. Wash 3 times with 200 μl NT2 buffer.
- Add 30 μg target antibody (such as HuR 1 μg/μl from NBL) to each tube.
- Add 320 μl NT2 buffer to each tube. Overnight rotate at 4 °C.
- Centrifuge the tubes (Ab+ beads) 5,000 x g 5 min at 4 °C. Discard supernatant wash twice with freshly made NT2 buffer 1 ml. After last wash, temporarily keep supernatant and put on ice until cell lysate is ready.
- The following components in DEPEC treated water are final concentrations.
- Lysis of mammalian cells
- Prepare cell pellet:
- Detach the cells from the culture dish by pipetting or using a cell scraper, if necessary. Collect the cell suspension into centrifuge tube.
- Centrifuge the cell suspension at 300 x g for 5 min at 4 °C to pellet the cells. Carefully remove and discard the supernatant.
- Wash the cells by resuspending the cell pellet with ice-cold PBS.
- Centrifuge the cell suspension at 300 x g for 5 min at 4 °C to pellet the cells. Carefully remove and discard the supernatant.
- Wash the cells once again using steps 3-4.
- Wash the cells by resuspending the cell pellet with ice-cold nuclease-free PBS.
- Centrifuge the cell suspension at 300 x g for 5 min at 4 °C to pellet the cells. Carefully remove and discard the supernatant.
- Wash the cells by resuspending the cell pellet with ice-cold nuclease-free PBS.
- Aliquot the cell suspension to each new microcentrifuge tube.
- Centrifuge the cell suspension at 300 x g for 5 min at 4 °C to pellet the cells. Carefully remove and discard the supernatant.
- Detach the cells from the culture dish by pipetting or using a cell scraper, if necessary. Collect the cell suspension into centrifuge tube.
- Lysis cell pellet
- Make RSB buffer in DEPEC treated water with final following concentrations.
10 mM Tris (pH 7.5)
100 mM NaCl
2.5 mM MgCl2
Then add other supplements in per ml RSB buffer.
10 μl digitornin/ml buffer (Digitornin is dissolved in ethanol 4 mg/ml freshly), 3 μl RNase inhibitor/ml buffer, 40 μl proteinase inhibitor cocktail/ml. - Add 200 μl of RSB buffer to each cell pellet. On ice 2 min. Centrifuge 4,400 rpm 4 °C, 8 min.
- Make 753 μl master mix for each tube (Ab + beads). For 4 tubes, you can make 4.5 times more.
700 μl NT2 buffer
10 μl 0.1 M DTT
10 μl RNase inhibitor
33 μl 0.5 M EDTA (pH 8) - Then add each supernatant of cell lysate to tube (Ab+ beads). Rotate at 4 °C 2 h.
- Take 20 μl for western blot from the mixture. The centrifuge the tube at 5,000 x g 4 °C 2 min.
- Discard supernatant. Wash pellet with 1 ml NT2 buffer twice at 5,000 x g 4 °C 2 min.
- Further process the mixture to get mRNAs. Make DNAase master mix.
Now we have 4 tubes, so make 4.5 times master mix.
100 μl of NT2 buffer x 4.5=450 μl
5 μl of DNAase (2U/μl) x 4.5=22.5 μl
Add 105 μl of DNAase master mix to each tube and 37 °C, 10 min incubation. - Then add 1ml NT2 buffer centrifuge 5,000 x g 5 min 4 °C. Remove supernatant.
- Add proteinase K (20 mg/μl) master mix 103 μl to each tube. 55 °C, 15 min plus mixing.
Master mix: 2.5 μl proteinase K x 4.5=11.25 μl
0.5 μl 20% SDS x 4.5=2.25 μl
100 μl NT2 buffer x 4.5=450 μl - Centrifuge 5,000 x g 4 °C 5 min.
- Collect 100 μl supernatant to a new Eppendorf tube. Add 200 μl of NT2 buffer to old tube, centrifuge 5,000 x g 4 °C, 5 min again. Collect 200 μl of supernatant and pool to the new tube, which becomes 300 μl of supernatant.
- Add equal volume of acid phenol/chloroform to above supernatant.
Vertex 1 min, then centrifuge 1 min RT, 13k rpm. - Collect about 250 μl of aqueous phase (top phase)
Add: 25 μl of 3 M NaoAc (pH 5.5)
625 μl of 100% ethanol
3 μl of glycogen blue
Mix well and put it at -20 °C or below for 20 min (or for overnight, if necessary). - Centrifuge the tube at 12,000 x g for 10 min at 4 °C, then aspirate the supernatant carefully.
- Rinse with 1 ml 70% Ethanol, 13,000 x g 2 min at 4 °C. Discard supernatant. Invert the tube, air dry 5 min. The pellet is mRNAs.
- Make RSB buffer in DEPEC treated water with final following concentrations.
- Prepare cell pellet:
Recipes
- Lysis buffer
Add appropriate concentrations of protease inhibitors, RNase inhibitor, and DTT to lysis buffer just before use. Lysis buffer containing these reagents is described as lysis buffer (+) in the following protocols. The optimal concentration of each reagent for RIP-Assay is shown as follows.
Make RSB buffer in DEPEC treated water with final concentrations as follows:
10 mM Tris (pH 7.5)
100 mM NaCl
2.5 mM MgCl2
Then add other supplements in per ml RSB buffer.
10 μl digitornin/ml buffer (digitornin is dissolved in ethanol 4 mg/ml freshly), 3 μl RNase inhibitor/ml buffer, 40 μl proteinase inhibitor cocktail/ml. - Wash buffer
Add final 1.5 mM concentration of DTT to wash buffer just before use. Wash buffer containing DTT is described as wash buffer (+) in the following protocols.
Make washing buffer in DEPEC treated water with final concentrations as follows:
50 mM Tris (pH 7.5)
50 mM NaCl
1 mM MgCl2
0.05% Nonidet P40 - Precaution: Additional buffer preparation
In some cases, both the lysis buffer (+) and wash buffer (+) may require the addition of appropriate volumes of high-salt solution (in these cases, add 30 μl of high-salt solution to each ml of lysis buffer and wash buffer).
Acknowledgments
This protocol was adapted from the NBL RIP-Assay Kit (see Reference 1).
References
- Protocol from NBL RIP-Assay Kit (NBL, catalog number: RN1001).
Article Information
Copyright
© 2012 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Liu, F. (2012). RNP-IP (Modified Method)-Getting Majority of RNA from RNA Binding Protein in Cytoplasm. Bio-protocol 2(13): e219. DOI: 10.21769/BioProtoc.219.
Category
Biochemistry > RNA > RNA-protein interaction
Biochemistry > Protein > Immunodetection
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








