- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Pseudomonas syringae Flood-inoculation Method in Arabidopsis
Published: Vol 7, Iss 2, Jan 20, 2017 DOI: 10.21769/BioProtoc.2106 Views: 13630
Reviewed by: Zhaohui LiuBin TianAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
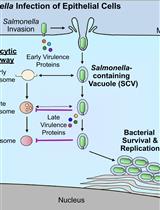
Bacterial Pathogen-mediated Suppression of Host Trafficking to Lysosomes: Fluorescence Microscopy-based DQ-Red BSA Analysis
Mădălina Mocăniță [...] Vanessa M. D'Costa
Mar 5, 2024 2929 Views
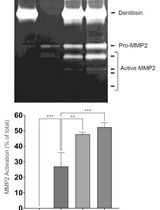
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2122 Views
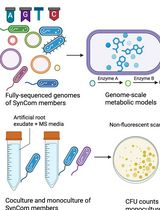
In Silico Prediction and In Vitro Validation of Bacterial Interactions in the Plant Rhizosphere Using a Synthetic Bacterial Community
Arijit Mukherjee [...] Sanjay Swarup
Nov 5, 2025 1703 Views
Abstract
Pseudomonas syringae pv. tomato strain DC3000 (Pto DC3000), which causes bacterial speck disease of tomato, has been used as a model pathogen because of its pathogenicity on Arabidopsis thaliana. Here, we demonstrate a rapid and reliable flood-inoculation method based on young Arabidopsis seedlings grown on one-half strength MS medium. We also describe a method to evaluate the bacterial growth in Arabidopsis.
Keywords: Pseudomonas syringaeBackground
The A. thaliana-Pto DC3000 model pathosystem is widely used to investigate the molecular mechanisms of microbial pathogenesis and plant innate immunity (Ishiga et al., 2012 and 2016; Ishiga and Ichinose, 2016). Although several inoculation methods have been developed to study the interactions in this model system, none of the methods reported to date are similar to those occurring in nature.
Materials and Reagents
- 1.5 ml microcentrifuge tubes (Ina-optika, catalog number: LT-0150 )
- 1 ml pipette tips (Mettler-Toledo, catalog number: 768/8 RT-1000 )
- Inoculating loops (VWR, catalog number: 12000-812 )
- 3M Micropore 2.5 cm surgical tape (3M, catalog number: 1530-1 )
- Plastic Petri plates (100 x 20 mm) (TrueLine, catalog number: TR4002 )
- Plastic Petri plates (100 x 15 mm) (VWR, catalog number: 10053-704 )
- Conical tubes (50 ml) (LabPlanet, catalog number: 3181-345-008 )
- Plant material: Arabidopsis thaliana ecotype Columbia (Col-0) was used as a wild-type plant
- Bacterial strains: P. syringae pv. tomato DC3000 (Pto DC3000) was used as pathogenic strain on A. thaliana-Pto DC3000 was kindly provided by Dr. Fumiaki Katagiri at University of Minnesota
- Ethanol (70%) (Wako Pure Chemical Industries, catalog number: 059-07895 )
- Sodium hypochlorite (Wako Pure Chemical Industries, catalog number: 197-02206 )
- Tween 20 (Wako Pure Chemical Industries, catalog number: 327-32475 )
- Sterile distilled water
- Rifampicin (50 mg/ml) (Wako Pure Chemical Industries, catalog number: 185-01003 )
- Silwet L-77 (BMS, catalog number: BMS-SL7755 )
- 5% H2O2 (Wako Pure Chemical Industries, catalog number: 081-04215 )
- Mannitol (Wako Pure Chemical Industries, catalog number: 133-00845 )
- L-glutamic acid (Wako Pure Chemical Industries, catalog number: 074-00505 )
- KH2PO4 (Wako Pure Chemical Industries, catalog number: 169-04245 )
- NaCl (Wako Pure Chemical Industries, catalog number: 191-01665 )
- MgSO4·7H2O (Wako Pure Chemical Industries, catalog number: 131-00405 )
- Agar (Wako Pure Chemical Industries, catalog number: 010-15815 )
- NaOH (Wako Pure Chemical Industries, catalog number: 192-15985 )
- Murashige and Skoog basal medium (Sigma-Aldrich, catalog number: M0404 )
- Sucrose (Wako Pure Chemical Industries, catalog number: 196-00015 )
- PhytagelTM (Sigma-Aldrich, catalog number: P8169 )
- KOH (Wako Pure Chemical Industries, catalog number: 165-21825 )
- Media
- Mannitol-glutamate (MG) medium (see Recipes)
- One-half strength Murashige and Skoog (MS) medium (see Recipes)
Equipment
- High-speed centrifuge (TOMY, model: MX305 )
- Pipette (Mettler-Toledo, model: Pipet-Lite XLS+ )
- Labo shaker (TAITEC, model: NR-2 )
- Labo incubator (AS ONE, model: ICV-300P )
- Clean work station (Hitachi Industrial Equipment Systems, model: CCV-1306E )
- Spectrophotometer (JASCO, model: V-630 )
- Plant growth chamber (NKsystem, model: LPH-411SP )
- Digital scale (AS ONE, model: IB-300 )
- Mortars and pestles (AS ONE, catalog number: 6-549-02 , φ90 mm)
Procedure
- Plant growth
- Incubate 100-200 Arabidopsis seeds in 70% ethanol for 5 min in a microcentrifuge tube.
- Allow the seeds to settle by centrifugation (500 x g for 30 sec) and remove 70% ethanol by pipetting.
- Incubate the seeds with 6% sodium hypochlorite containing 0.1% Tween 20 for 15 min with gentle shaking.
- Allow the seeds to settle by centrifugation (500 x g for 30 sec) and remove 6% sodium hypochlorite by pipetting.
- Rinse the seeds with sterile distilled water at least four times.
- Maintain the seeds in a small amount of sterile distilled water (300-500 μl).
Note: The seeds can be stored at 4 °C. The cold treatment for 2 days improves the rate and synchrony of germination. - Plant individual seeds in low density (50-80 seeds/plate) using a 1 ml pipette tip on one-half strength MS plates (Figure 1A and Video 1).
Note: The one-half strength MS plates need to be dried overnight in a sterile hood work station with closed lids before transferring the surface-sterilized seeds. If the one-half strength MS plates are not dried enough, the vitreous and wet plants are observed. These plants are more sensitive to pathogen inoculation.
Figure 1. Procedure of Pseudomonas syringae flood-inoculation method in Arabidopsis. A. Planting the seeds on one-half strength MS plates. B. Transferring the seedlings to new one-half strength MS plates at one week after germination with forceps. C. Incubation of the seedlings at 24 °C with a light intensity of 150-200 μE m-2 sec-1 and a 12 h light/12 h dark photoperiod. D. Two-week-old Arabidopsis seedlings for pathogen assays. E. Preparation of P. syringae pv. tomato DC3000 (Pto DC3000) inoculum. F. Flood-inoculation by dispensing the bacterial suspension onto the plates. G. Removal of the bacterial suspension by decantation. H. Disease phenotypes of Arabidopsis seedlings 4 days after inoculation with Pto DC3000.Video 1. Procedure of planting the seeds on one-half strength MS plates - Incubate the seeds at 24 °C with a light intensity of 150-200 μE m-2 sec-1 and a 12 h light/12 h dark photoperiod for one week.
- Transfer the healthy seedlings at one-week post-germination to fresh new one-half strength MS plates with forceps (Figure 1B).
Note: The roots need to be put on one-half strength MS plate without damaging and seven seedlings per plate are good for healthy plant growth. - Incubate the seedlings at 24 °C with a light intensity of 150-200 μE m-2 sec-1 and a 12 h light/12 h dark photoperiod for one week (Figure 1C). Use the two-week old post-germination seedlings for the pathogen assays (Figure 1D).
- Bacterial growth and inoculum preparation
- Streak a -80 °C stock of bacterial culture on MG agar supplemented with rifampicin (50 μg/ml). Incubate the plate at 28 °C until bacterial colonies grow (24-48 h).
- Inoculate a single colony of Pto DC3000 onto new MG agar with rifampicin (50 μg/ml) and streak all over the plate. Incubate the plate at 28 °C (24-48 h).
- Collect as many bacterial colonies as possible with inoculating loop and suspend all colonies in an appropriate volume (20-30 ml) of sterile distilled water (Figure 1E).
Note: Be careful not to take small pieces of MG agar medium. - Measure the OD (A600) of the bacterial suspension and dilute to 4 x 106 CFU (colony forming units)/ml using sterile distilled water.
Note: 1.0 OD measurement approximately corresponds to 2 x 108 CFU/ml. - Add Silwet L-77 to a final concentration of 0.025% to the bacterial suspension and mix gently. Use the bacterial suspensions immediately after preparation.
- Flood-inoculation method
- Dispense 40 ml of bacterial suspension onto the plate containing the 2-week-old Arabidopsis seedlings and incubate for 2-3 min at room temperature (Figure 1F).
Note: One bacterial plate normally allows to make more than 800 ml of bacterial suspension. - Remove the bacterial suspension by decantation and seal the plates with 3M Micropore 2.5 cm surgical tape (Figure 1G).
- Incubate the inoculated seedlings at 24 °C with a light intensity of 150-200 μE m-2 sec-1 and a 12 h light/12 h dark photoperiod.
- Observe the symptom development at 2-4 days-post-inoculation (dpi) (Figure 1H).
- Dispense 40 ml of bacterial suspension onto the plate containing the 2-week-old Arabidopsis seedlings and incubate for 2-3 min at room temperature (Figure 1F).
- Evaluation of bacterial growth
- Collect four seedlings from a single plate by cutting the hypocotyls to separate the above agar parts (whole rosette) from the Phytagel plate at 0, 1, 2 and 3 dpi and measure the seedlings’ weight.
Note: Avoid collecting vitreous and wet plants. These plants are more sensitive to pathogen inoculation. - Surface-sterilize the seedlings in a beaker with 5% H2O2 for 3 min to determine the internal bacterial population and rinse the seedlings three times by dipping in sterile distilled water.
Note: The bacterial populations at 0 dpi is estimated from seedlings collected 1 h post-inoculation without surface-sterilization by H2O2. - Homogenize the seedlings completely with mortars and pestles for at least 1 min, and add 5 ml of sterile distilled water.
- Dilute the samples from 10-2 to 10-6 and plate on MG agar with rifampicin (50 μg/ml).
Note: 10-2 to 10-3 dilution is suitable for 0 dpi and 10-4 to 10-6 is for 1, 2 and 3 dpi. - Incubate the plates for 2-3 days and count the bacterial CFU using properly diluted samples.
Note: Yellow non-mucoid colonies are observed after 2-3 days. - Normalize the CFU as CFU/mg of tissue using the total weights of the inoculated seedlings.
Note: The bacterial populations are evaluated from at least three biological replicates and each replicate represents a pooled sample of four independent seedlings from a single experiment grown in a single plate.
- Collect four seedlings from a single plate by cutting the hypocotyls to separate the above agar parts (whole rosette) from the Phytagel plate at 0, 1, 2 and 3 dpi and measure the seedlings’ weight.
Data analysis
Student’s t-test was used for comparing two averages of bacterial growth, while ANOVA with post-hoc Tukey’s honest significant difference correction for multiple correction was applied when considering more than two averages of bacterial growth.
Recipes
- Mannitol-glutamate (MG) medium
10 g mannitol
2.0 g L-glutamic acid
0.5 g KH2PO4
0.2 g NaCl
0.2 g MgSO4·7H2O
15 g agar
Adjust pH to 7.0 with 3 N NaOH and make up the volume to 1 L with distilled water prior to autoclaving
Add rifampicin after autoclaving and pour MG medium into plastic Petri plates (100 x 15 mm) - One-half strength Murashige and Skoog (MS) medium
2.2 g Murashige and Skoog basal medium
10 g sucrose
3.0 g PhytagelTM (final 0.3%)
Adjust pH to 5.8 with 1 N KOH and make up the volume to 1 L with distilled water prior to autoclaving
Pour one-half strength MS medium into plastic Petri plates (100 x 20 mm)
Acknowledgments
This work was supported, in part, by the Program to Disseminate Tenure Tracking System, MEXT, Japan and JST ERATO NOMURA Microbial Community Control Project. This protocol was modified from previous work (Ishiga et al., 2011).
References
- Ishiga, Y. and Ichinose, Y. (2016). Pseudomonas syringae pv. tomato OxyR is required for virulence in tomato and Arabidopsis. Mol Plant Microbe Interact 29(2): 119-131.
- Ishiga, Y., Ishiga, T., Ikeda, Y., Matsuura, T. and Mysore, K. S. (2016). NADPH-dependent thioredoxin reductase C plays a role in nonhost disease resistance against Pseudomonas syringae pathogens by regulating chloroplast-generated reactive oxygen species. Peer J 4: e1938.
- Ishiga, Y., Ishiga, T., Uppalapati, S. R. and Mysore, K. S. (2011). Arabidopsis seedling flood-inoculation technique: a rapid and reliable assay for studying plant-bacterial interactions. Plant Methods 7: 32.
- Ishiga, Y., Ishiga, T., Wangdi, T., Mysore, K. S. and Uppalapati, S. R. (2012). NTRC and chloroplast-generated reactive oxygen species regulate Pseudomonas syringae pv. tomato disease development in tomato and Arabidopsis. Mol Plant Microbe Interact 25(3): 294-306.
Article Information
Copyright
© 2017 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ishiga, Y., Ishiga, T., Ichinose, Y. and Mysore, K. S. (2017). Pseudomonas syringae Flood-inoculation Method in Arabidopsis. Bio-protocol 7(2): e2106. DOI: 10.21769/BioProtoc.2106.
Category
Plant Science > Plant immunity > Disease bioassay
Microbiology > Microbe-host interactions > Bacterium
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link














