- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Highly Accurate Real-time Measurement of Rapid Hydrogen-peroxide Dynamics in Fungi
Published: Vol 6, Iss 24, Dec 20, 2016 DOI: 10.21769/BioProtoc.2080 Views: 10402
Reviewed by: Valentine V TrotterEmilia Krypotou Sadri Znaidi

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Determination of the Cellular Ion Concentration in Saccharomyces cerevisiae Using ICP-AES
Louise Thines [...] Pierre Morsomme
Aug 20, 2020 3451 Views

Determination of Fungal Tolerance Index to Heavy Metals and Heavy Metal Resistance Tests
Rosa Paulina Calvillo-Medina
Nov 5, 2021 2913 Views

High-throughput Growth Measurements of Yeast Exposed to Visible Light
Katarina Logg [...] Mikael Molin
Jan 20, 2022 3179 Views
Abstract
Reactive oxygen species (ROS) are unavoidable by-products of aerobic metabolism. Despite beneficial aspects as a signaling molecule, ROS are principally recognized as harmful agents that act on nucleic acids, proteins and lipids. Reactive oxygen species, and, in particular, hydrogen peroxide (H2O2), are deployed as defense molecules across kingdoms, e.g., by plants in order to defeat invading pathogens like fungi. Necrotrophic plant pathogenic fungi themselves secrete H2O2 to induce host cell death and facilitate infection. Hydrogen peroxide is, to a certain extent, freely diffusible through membranes. To be able to monitor intracellular hydrogen peroxide dynamics in fungi, we recently established the versatile HyPer-imaging technique in the filamentous plant pathogen Fusarium graminearum (Mentges and Bormann, 2015). HyPer consists of a circularly permuted yellow fluorescent protein (cpYFP) inserted into the regulatory domain (RD) of the prokaryotic H2O2-sensing protein, OxyR. The OxyR domain renders the sensor highly specific for H2O2. Oxidation of HyPer increases fluorescence of cpYFP excited at 488 nm and decreases fluorescence excited at 405 nm, thereby facilitating ratiometric readouts (Belousov et al., 2006). HyPer turned out to be pH-sensitive. A single amino acid mutation in the H2O2-sensing domain of HyPer renders the sensor insensitive to H2O2. This reporter is called SypHer and serves as a control for pH changes.
By using the HyPer-imaging technique, we could demonstrate that: i) HyPer imaging enables the specific and accurate detection of rapid changes in the intracellular H2O2 balance, ii) F. graminearum reacts on external stimuli with the transient production of H2O2, and iii) faces increased H2O2 level during initial infection of wheat.
The aim of this protocol is to guide the user through the basic setup of an in vitro HyPer imaging experiment in basically any fungus. It will provide the specific parameter for the fluorescence imaging as well as the construction of customized flow chambers for in vitro applications.
Background
HyPer is a genetically encoded, highly specific H2O2 sensor protein enabling the real-time detection of fluctuations in the intracellular H2O2-level, e.g., in response to external stimuli. Genetically encoded sensors are advantageous over classical staining methods like 2’,7’-dichlorodihydrofluorescein diacetate (H2DCFDA), 3,3’-Diaminobenzidine (DAB), or boronate-based H2O2 probes since they lack the major disadvantages of the latter ones as for example technically sophisticated application, requirement for chemical fixation of cells, insufficient uptake, inadequate intracellular distribution of stains, and occasionally irreversibility (reviewed in Guo et al., 2014). A genetically encoded sensor widely used for in-vivo detection of redox states in a cell is the redox-sensitive GFP (roGFP) system. The roGFP system is, however, not specific to a certain subtype of oxidative agent, i.e., H2O2, but (indirectly) monitors the redox status of a cell.
Materials and Reagents
- Microplates, 96 well, black, F-bottom (Greiner Bio One, catalog number: 655076 )
- Cover slips (24 x 40 mm) (Carl Roth, catalog number: 1870.2 )
- Standard microscope slides (Carl Roth, catalog number: 0656.1 )
- 10 ml disposable syringes (Carl Roth, catalog number: 0058.1 )
- 0.35 x 25 mm endo needle for root canal rinsing (Vedefar, Dilbeek, catalog number: 99010 )
- 92 mm Petri dish (SARSTEDT, catalog number: 82.1473 )
- Whatman paper (Sigma-Aldrich, catalog number: WHA10347511 ) (Optional)
- Plastic disposal bags (Carl Roth, catalog number: E706.1 )
- Double-sided adhesive frame (Gene Frame) (Thermo Fisher Scientific, Thermo Fisher ScientificTM, catalog number AB0576 )
- Pipette tips (1,000 µl, 200 µl, 10 µl)
- PCR tubes
- Conidia of F. graminearum (i.e., of F. graminearum expressing HyPer-2 or SypHer; preferably fresh, not frozen)
- pC1-HyPer-2 (Addgene, catalog number: 42211 )
- pC1-HyPer-C199S (SypHer) (Addgene, catalog number: 42213 )
- PCR primer for HyPer and SypHer amplification: forward primer 5’-ATG GAG ATG GCA AGC CCA GCA GGG CGA GAC GAT GT-3’; reverse primer 5’-GCT TTT AAA CCG CCT GTT-3’
- Immersion oil, Immersol W 2010 (Pulch + Lorenz, catalog number: 444969-0000-000 )
- Hydrogen peroxide 30% (Carl Roth, catalog number: 9681.4 )
- 1,4-dithiothreitol (DTT) (Sigma-Aldrich, catalog number: 000000010197777001 )
- Calcium nitrate tetrahydrate, Ca(NO3)2·4H2O (Carl Roth, catalog number: X886.1 )
- Potassium dihydrogen phosphate, KH2PO4 (Carl Roth, catalog number: P018.1 )
- Magnesium sulfate heptahydrate, MgSO4·7H2O (Sigma-Aldrich, catalog number: 230391-500G )
- Sodium chloride, NaCl (Carl Roth, catalog number: 9265.1 )
- Boric acid, H3BO3 (Carl Roth, catalog number: 6943.1 )
- Copper(II) sulfate pentahydrate, CuSO4·5H2O (Sigma-Aldrich, catalog number: 209198-250G )
- Potassium iodide, KI (Carl Roth, catalog number: 6750.1 )
- Manganese(II) sulphate monohydrate, MnSO4·H2O (Carl Roth, catalog number: 7347.2 )
- Ammonium molybdate tetrahydrate, (NH4)6Mo7O24·4H2O (Sigma-Aldrich, catalog number: 09880-100G )
- Zinc sulphate heptahydrate, ZnSO4·7H2O (Carl Roth, catalog number: 7316.1 )
- Iron(III) chloride hexahydrate, FeCl3·6H2O (Carl Roth, catalog number: 7119.1 )
- Chloroform
- Sucrose (Carl Roth, catalog number: 9097.2 )
- Granulated agar (BD, catalog number: 214530 )
- Solution A (see Recipes)
- Solution B (see Recipes)
- Suspension D (see Recipes)
- Minimal medium (see Recipes)
Equipment
- Microplate multimode reader (e.g., Berthold Technologies Multimode Microplate Reader Mithras² LB 943, BERTHOLD TECHNOLOGIES, model: Mithras2 LB 943 ), equipped with fluorescence excitation filters (380 x 10 nm and 485 x 14 nm, e.g., BERTHOLD TECHNOLOGIES, catalog numbers: 40087-01 and 40271-01 ), fluorescence emission filter (520 x 10 nm, e.g., BERTHOLD TECHNOLOGIES, catalog number: 38836-01 ), and injectors (e.g., BERTHOLD TECHNOLOGIES, model: 54116 )
- Neubauer counting chamber improved (Carl Roth, catalog number: T729.1 )
- Confocal laser scanning microscope (e.g., LSM 780 mounted on a Carl Zeiss Axio Imager.Z2 microscope with motorized stage)
- 40x objective (e.g., Carl Zeiss C-apochromat Carl Zeiss 40x/1.20 W Korr M27)
- Incubator at 28 °C (Thermo Fisher Scientific, model: Heraeus B20/UB20 )
- Solid-state laser 405 nm, 50 mW
- Argon ion laser 458, 488 and 514 nm, 30 mW
- Multi-channel pipette (Eppendorf, catalog number: 3122000035 )
- Pipettors: 10-100 µl (Eppendorf, catalog number: 4920000059 ), 100-1,000 µl (Eppendorf, catalog number: 4920000083 )
- 300 mm Heidelberger extension (Dezember, Fresenius Kabi Deutschland, catalog number: 2873112 )
Software
- Plate reader software (e.g., Berthold MikroWin Lite software [Berthold Technologies])
- Spreadsheet software program (e.g., Excel [Microsoft])
- ImageJ (Version 1.46r, http://imagej.net/)
Procedure
- Generation of HyPer and SypHer reporter mutants
- Amplify sequences for HyPer and SypHer from pC1-HyPer-2 and pC1-HyPer-C199S (SypHer) using forward primer 5’-ATG GAG ATG GCA AGC CCA GCA GGG CGA GAC GAT GT-3’ and reverse primer 5’-GCT TTT AAA CCG CCT GTT-3’ (one may add restriction enzyme recognition sites to the 5’ end of the primer to facilitate cloning in the desired binary vector).
- Transform the fungus according to standard protocols. For F. graminearum, the authors refer to Maier et al. (2005).
Note: It is recommended to check HyPer and SypHer expression by fluorescence microscopy or RT-PCR.
- Amplify sequences for HyPer and SypHer from pC1-HyPer-2 and pC1-HyPer-C199S (SypHer) using forward primer 5’-ATG GAG ATG GCA AGC CCA GCA GGG CGA GAC GAT GT-3’ and reverse primer 5’-GCT TTT AAA CCG CCT GTT-3’ (one may add restriction enzyme recognition sites to the 5’ end of the primer to facilitate cloning in the desired binary vector).
- 96-well plate assay
The following paragraph describes an assay using a 96-well plate and multimode reader to analyze the response of F. graminearum mycelia to H2O2, DTT and test substances A and B, which might be any substance of choice at a given concentration to be assayed for its potential to cause an H2O2 burst in F. graminearum.- Fill each well of a black 96-well plate with 100 µl minimal medium and wait 30 min until solidified.
- Inoculate the 96-well plate with 200 conidia (i.e., 10 µl of a 20 conidia µl-1 suspension) according to the pipetting scheme shown in Figure 1.

Figure 1. Scheme for loading and inoculation of a 96-well assay plate to be analyzed in a fluorescence plate reader. Each well is filled with 100 µl minimal medium and inoculated with conidia of the wild type (columns 3 and 4), SypHer (columns 5 and 6), and HyPer (columns 7-12). Columns 1 and 2 remain uninoculated as control. - Cover the 96-well plate, place it into a plastic bag moistened with sterile ddH2O to prevent dehydration and incubate it at 28 °C in permanent darkness for 2 days.
- On day 3, the HyPer assay shall be performed. Prolonged incubation leads to an accumulation of aerial hyphae that perturb measurement.
- Start the multimode reader and allow for the interior chamber to reach 28 °C.
- Prime the injectors according to the manufacturer’s instructions. By way of example, injector 1 could be primed with a 150 mM H2O2 solution, injector 2 with a 150 mM DTT solution, injector 3 with test substance A and injector 4 with test substance B at a concentration threefold the final concentration in the well.
- Program the measuring cycles in the plate reader software:
- Choose excitation filter ‘380 x 10’ and emission filter ‘520 x 10’ for measuring the fluorescence emitted by HyPer in its reduced state and excitation filter ‘485 x 14’ (emission filter ‘520 x 10’ for the oxidized state).
- Configure the injectors. For example, injector 1 shall inject 50 µl of the test substance at cycle 31 and injector 2 shall inject 50 µl of another substance at cycle 61. If possible, choose a slow to medium injection speed in both cases to prevent spilling.
Note: The user should avoid long measuring cycles because the HyPer response is typically very fast. If a substance is injected into all wells before the measurement of fluorescence resumes, the immediate response might already be over. Yet, the authors recommend measuring one well at a time immediately after injection.
- Add 100 µl ddH2O to all wells using a multi-channel pipette and let plate stand for 30 min at 28 °C.
- Insert plate to plate reader and start the measuring cycles.
- After the run, export the raw data for further analysis in a spreadsheet software program.
- For ratio calculations, divide the fluorescence emission value (FEV) measured after excitation with 485 nm in well A1 by the FEV after excitation with 380 nm in this particular well (equation 1).

Note: Equation 1 represents the calculation of the ratio of fluorescence intensities measured after excitation with 485 nm and 380 nm, respectively. - Apply this to all wells. Average the ratios for the biological replicates (i.e., wells A1 to H2 for the media control) for each time point and calculate the standard deviations. Plot the averaged ratios on the y-axis against the time on the x-axis. A measurement of fungal response to H2O2 and DTT according to the procedures described above should typically result in a curve as depicted in Figure 2.
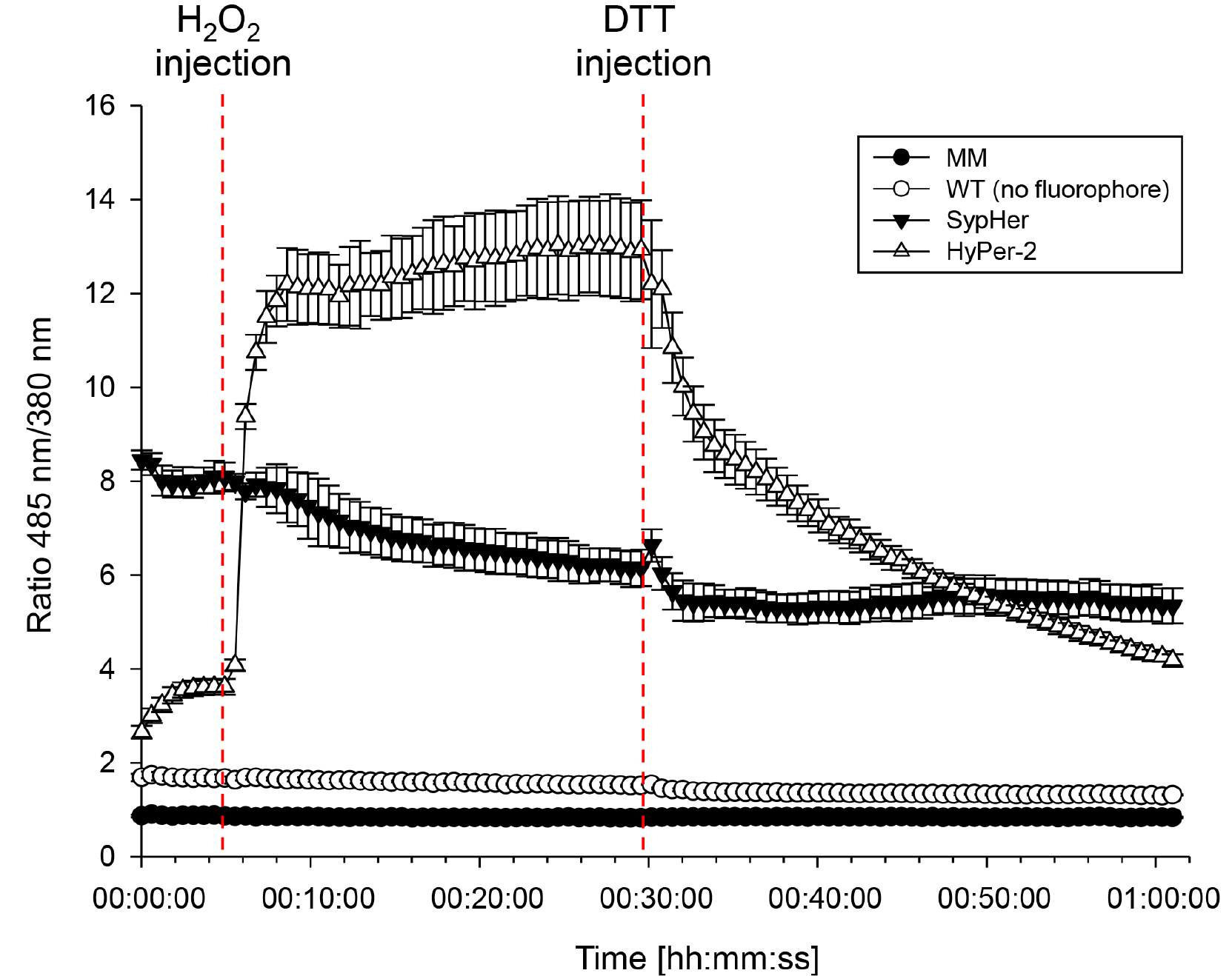
Figure 2. Ratiometric time course assay of the fungal response to external H2O2 and dithiothreitol (DTT). Timing of H2O2 (50 mM) and DTT (50 mM) induced ratio [485/380 nm] change in hyphae expressing HyPer, SypHer compared to the wild type (no fluorophore) and media control (MM). Mycelia were raised in a 96-well plate as described in the text and analyzed in a fluorometer. Error bars represent the standard deviation.
- Fill each well of a black 96-well plate with 100 µl minimal medium and wait 30 min until solidified.
- Confocal laser scanning microscopy assay
This paragraph describes the construction and usage of a fluidic chamber for in-situ microscopy of fungal response to external H2O2 or any test substance. The assembly of the fluidic chamber is illustrated in Figure 3 and Video 1.- Surface-sterilize microscope slides, e.g., in ethanol or UV light.
- Remove the cover foil that covers a gene frame and the window and adhere it to the slide.
- Fill 40 µl MM into the frame and immediately cover it with a second slide to obtain an even surface within the chamber.
- After 30 min, remove the upper cover slide by carefully moving it sideways and inoculate the agar surface with 300 conidia (15 µl of a 20 conidia µl-1 suspension) of mutants expressing HyPer or SypHer.
- Incubate in a sterile Petri dish (add 500 µl sterile ddH2O and use pipette tips as elevation) for 24 h at 28 °C in permanent darkness.
- On the next day, remove the cover foil of the second gene frame that covers the frame and the window and the remaining cover foil from the first gene frame. Take the second gene frame and attach it to the first frame on the slide.
- Cut a 1 mm and a 5 mm piece out of both gene frames to facilitate injection and drainage of a test substance, respectively (see Figure 3).
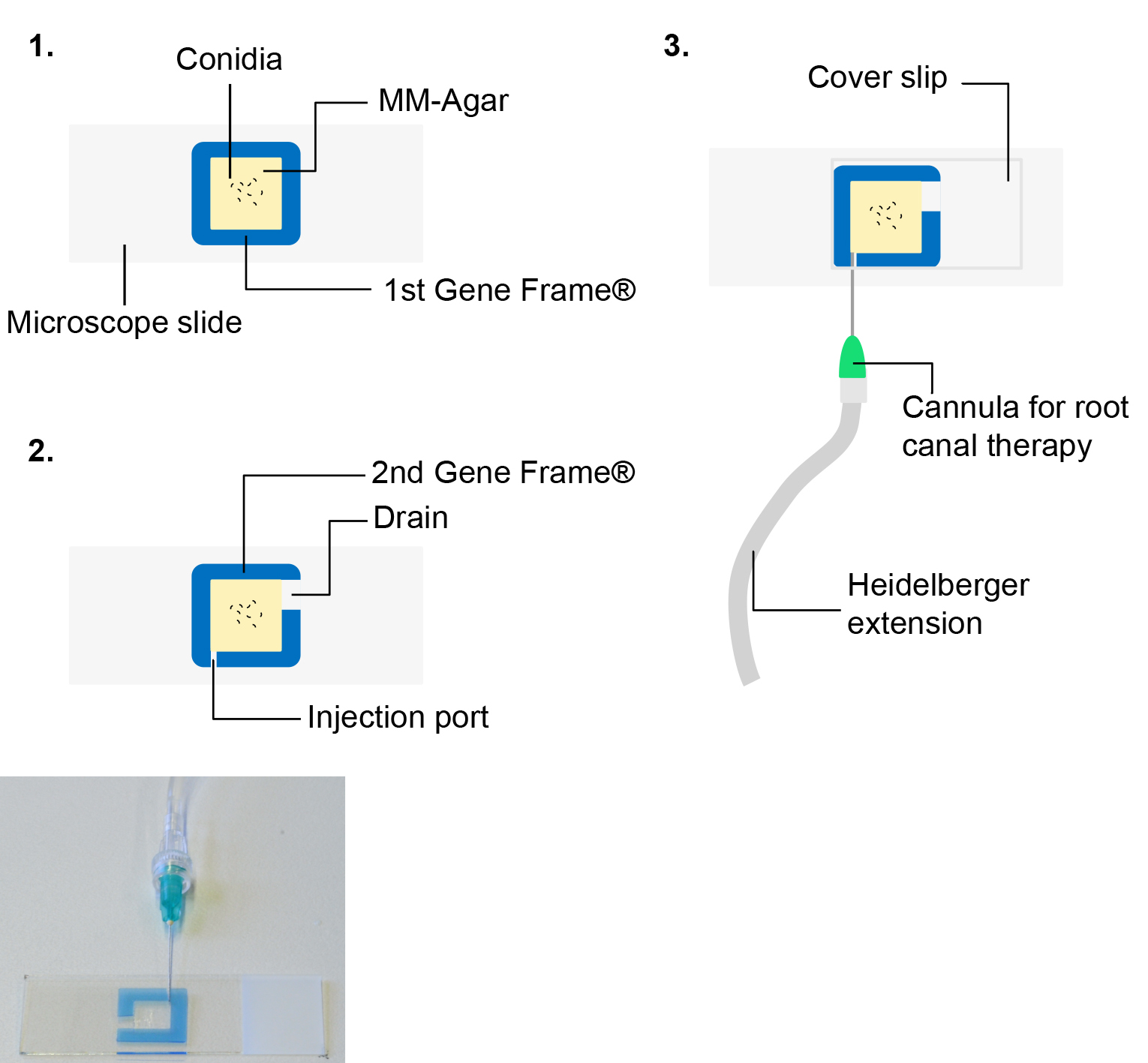
Figure 3. Assembly of a customized fluidic chamber for the simultaneous image acquisition and injection of test substances. Detailed description in the main text (see also Video 1).Video 1. Assembly of a fluidic chamber - Remove the remaining cover foil from the gene frame.
- Place a cover slip on the second frame with an overhang to the drainage port. It serves as a small reservoir for excess volume.
- Rinse the fluidic chamber with ddH2O using a syringe with an endo needle for root canal rinsing attached.
- Prefill a Heidelberger extension with a test substance (e.g., 50 mM H2O2) using a syringe and attach it to another endo needle.
- Mount the fluidic chamber to the stage of the CSLM.
- Plug the endo needle into the injection port and secure the position of the Heidelberger extension and endo needle with adhesive tape.
- Program a sequential acquisition of xyt-frames (lateral resolution 512 x 512, color depth: 8 bit) using excitation at 405 and 488 nm over at least 40 min with the shortest possible acquisition time and repetition rate. Detect the fluorescence signal for both channels in a range from 493 to 598 nm. Use a 40x/1.2 W objective for image acquisition.
- Start the scanning. Adjust laser intensity and gain of both channels so that the fluorescence signal intensity is approximately levelled in both channels (405 and 488 nm excitation). Figure 4 represents exemplarily the HyPer fluorescence in a hypha of F. graminearum.
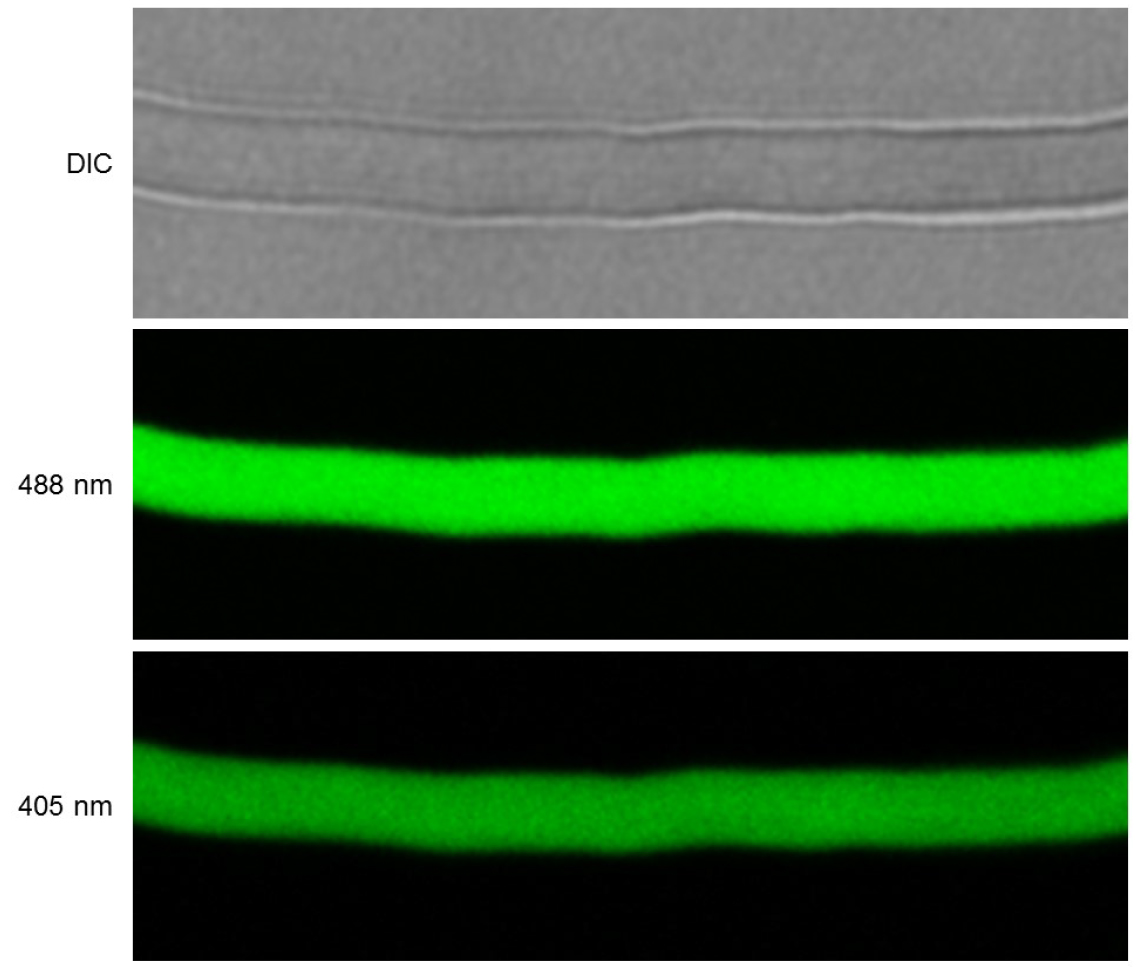
Figure 4. Fluorescence emission and bright field image of a F. graminearum hypha expressing HyPer
- Before starting the injection monitor steady-state fluorescence over 10 min.
- Carefully inject the test substance while continue scanning. In order to prevent damage of the microscope always observe the microscope slide for spillage. Inject as much volume as it takes to completely wash out the initially added ddH2O.
- Aquire xyt frames over at least 30 min.
- [Optional] Wash out the first test substance, wash the chamber once with ddH2O and repeat the injection using a new, prefilled Heidelberger extension. Take up excess volume at the drainage port using absorbent paper (e.g., Whatman paper).
- For data analysis, open xyt stacks with ImageJ.
- To obtain the FEVs of each channel over time mark and measure fluorescence intensities in one or more custom-shaped regions of interest using the ‘ROI manager’ tool and ‘measure stacks’ plugin. Calculate the 488 nm/405 nm ratio (as similar as equation 1) and plot it on the y-axis against the acquisition time on the x-axis.
- For visual representation of the 488 nm/405 nm ratio follow the instructions (link) of Kardash et al. (2011). Figure 5 shows exemplarily a 488 nm/405 nm false color ratio image of a hypha.
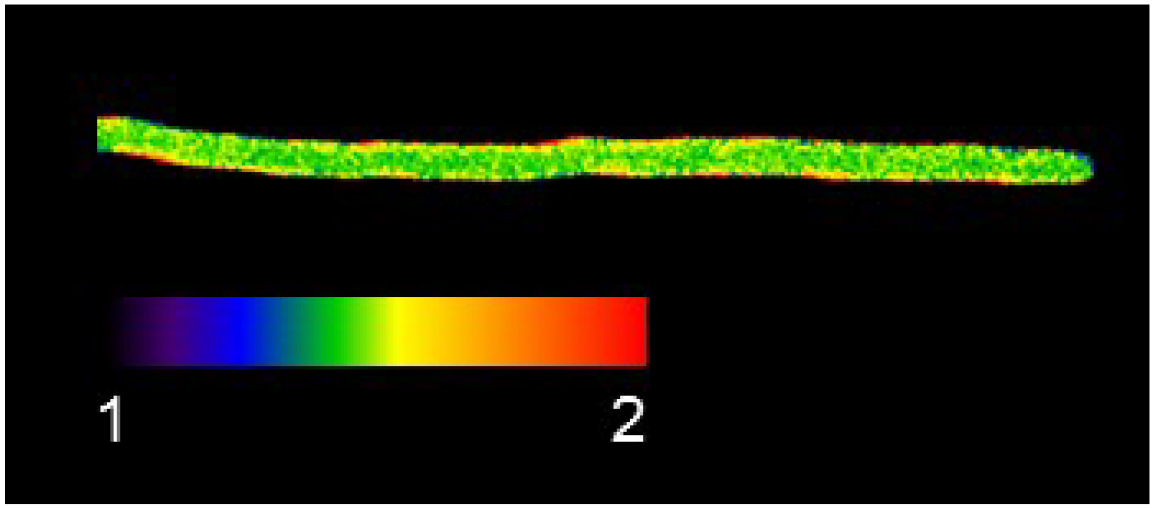

Figure 5. False color image of a 488 nm/405 nm ratio
- Surface-sterilize microscope slides, e.g., in ethanol or UV light.
Data analysis
The use of 96-well plates for the HyPer assays allows for high numbers of replicates. Each fungal colony growing in a well represents a biological replicate that is growing and monitored independently from the others. High numbers of replicates, in turn, allow for discrimination of colonies growing weakly due to an inappropriate distribution of media in the well or a low number of conidia due to pipetting mistakes. Results obtained from those wells can be excluded from the analysis.
For further details on data handling and statistics see Mentges and Bormann (2015).
Notes
HyPer measurements provide highly reproducible and reliable data on H2O2 fluctuations in hyphae of F. graminearum. The setup of a 96-well assay is easy and allows for a high number of replicates. This guarantees statistical accuracy. The CLSM analysis, although technically more sophisticated, enables monitoring intracellular H2O2 allocations, e.g., during nuclear division or directed growth (Mentges and Bormann, 2015). Critical for the CLSM assay is the plane agar surface provided by the two object slides stacked on top of each other separated by a gene frame.
Recipes
- Solution A
100 g/L Ca(NO3)2·4H2O
Sterilized by filtration - Solution B
20 g/L KH2PO4
25 g/L MgSO4·7H2O
10 g/L NaCl
Sterilized by filtration - Suspension D
60 g/L H3BO3
390 g/L CuSO4·5H2O
13 mg/L KI
60 mg/L MnSO4·H2O
51 mg/L (NH4)6Mo7O24·4H2O
5.48 g/L ZnSO4·7H2O
932 mg/L FeCl3·6H2O
Sterilized by addition of 0.1% (v/v) chloroform - Minimal medium (MM, 1 L)
10 ml solution A
10 ml solution B
10 g sucrose
1 ml suspension D
16 g agar
Acknowledgments
The authors thank Dr. V.V. Belousov for providing the HyPer and SypHer plasmids and B. Hadeler and C. Kröger for technical assistance.
References
- Belousov, V. V., Fradkov, A. F., Lukyanov, K. A., Staroverov, D. B., Shakhbazov, K. S., Terskikh, A. V. and Lukyanov, S. (2006). Genetically encoded fluorescent indicator for intracellular hydrogen peroxide. Nat Methods 3(4): 281-286.
- Guo, H., Aleyasin, H., Dickinson, B. C., Haskew-Layton, R. E. and Ratan, R. R. (2014). Recent advances in hydrogen peroxide imaging for biological applications. Cell Biosci 4: 64.
- Kardash, E., Bandemer, J. and Raz, E. (2011). Imaging protein activity in live embryos using fluorescence resonance energy transfer biosensors. Nat Protoc 6(12): 1835-1846.
- Maier, F. J., Malz, S., Lösch, A. P., Lacour, T. and Schäfer, W. (2005). Development of a highly efficient gene targeting system for Fusarium graminearum using the disruption of a polyketide synthase gene as a visible marker. FEMS Yeast Res 5(6-7): 653-662.
- Mentges, M. and Bormann, J. (2015). Real-time imaging of hydrogen peroxide dynamics in vegetative and pathogenic hyphae of Fusarium graminearum. Sci Rep 5: 14980.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Mentges, M. and Bormann, J. (2016). Highly Accurate Real-time Measurement of Rapid Hydrogen-peroxide Dynamics in Fungi. Bio-protocol 6(24): e2080. DOI: 10.21769/BioProtoc.2080.
Category
Microbiology > Microbial biochemistry > Other compound
Microbiology > Microbial physiology > Stress response
Cell Biology > Cell metabolism > Other compound
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











