- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Heterologous Expression and Purification of the Magnesium Transporter A (MgtA) in Escherichia coli
Published: Vol 6, Iss 22, Nov 20, 2016 DOI: 10.21769/BioProtoc.2001 Views: 11032
Reviewed by: Arsalan DaudiKanika GeraChijioke Joshua

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

From Llama to Nanobody: A Streamlined Workflow for the Generation of Functionalised VHHs
Lauren E.-A. Eyssen [...] Raymond J. Owens
Mar 20, 2024 6326 Views
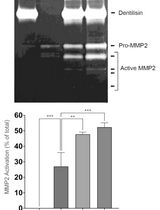
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2128 Views
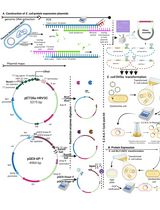
Thermus thermophilus CRISPR Cas6 Heterologous Expression and Purification
Junwei Wei [...] Yingjun Li
Jul 20, 2025 2208 Views
Abstract
The magnesium transporter A (MgtA) is a magnesium transporting P-type ATPase present in prokaryotes and plants (Subramani et al., 2016). In Salmonella typhimurium and Escherichia coli (E. coli), MgtA is expressed only in magnesium limiting conditions and plays an important role in Mg2+ homeostasis (Groisman et al., 2013). The transcription of mgtA is regulated by the two-component system PhoP/PhoQ (Soncini et al., 1996; Kato et al., 1999). The membrane bound histidine kinase, PhoQ, senses low Mg2+ concentration in the periplasmic space and phosphorylates its cognate response regulator, PhoP, which initiates mgtA transcription (Groisman et al., 2013). MgtA is targeted to the plasma membrane and facilitate the bacterial survival under low Mg2+ condition, by importing Mg2+ into the cytoplasm. The MgtA homolog in petunia (PH1) is found in the vacuolar membrane and involved with the coloration of the flower petals (Faraco et al., 2014). As a first step towards understanding the molecular details of MgtA Mg2+ transport, we describe a detailed protocol for the purification of E. coli MgtA that can be used for biochemical and biophysical studies. Recombinant E. coli MgtA with hexa histidine tag at the N-terminus was cloned from E. coli DH5α and over expressed in the E. coli C43(DE3) by fermentation to an OD > 6. Cell lysis was performed in a high pressure homogenizer and the membranes were isolated by ultracentrifugation. Membrane proteins were solubilized with the detergent dodecyl-β-D maltoside. MgtA was purified by affinity and size exclusion chromatography. Final yields of purified MgtA reach ~1 mg MgtA per 3 g of wet cell pellet.
Background
Recently we have reported that the purified MgtA from E. coli is highly dependent on lipids for its function and studied the enzyme kinetics in vitro (Subramani et al., 2016). The protocol described here is the detailed description going through every single step of the purification that should yield monodisperse detergent solubilized MgtA, earlier described in Subramani et al. (2016). The protocol will in addition present notes that describe critical points and observations made in the process.
Materials and Reagents
- Disposable cup, 100 ml, PP (polypropylene) (SARSTEDT, catalog number: 75.562.105 )
Note: It is used to store cell pellet. - 70 ml polycarbonate ultracentrifugation tube
- C43(DE3) (Lucigen, catalog number: 60446 )
- PfuUltra II Fusion HS DNA polymerase (Agilent Technologies, catalog number: 600670 )
- LB broth powder (Sigma-Aldrich, catalog number: L3022 )
- Kanamycin (Sigma-Aldrich, catalog number: K1377 )
- Polypropylene glycol P2000 (Sigma-Aldrich, catalog number: 81380 )
- Gistex® LS Ferm - yeast extract (Fermia AB, Sweden)
- Glycerol (VWR, catalog number: 24388.295 )
- Isopropyl β-D-thiogalactopyranoside (IPTG) (Biosynth, catalog number: I-8000 )
- Sodium dodecyl sulfate (SDS) (Sigma-Aldrich, Catalog number: L4509 )
- β-mercaptoethanol (Sigma-Aldrich, catalog number: M3148 )
- Bromophenol blue (Thermo Fisher Scientific, Fisher Scientific, catalog number: BP115-25 )
- Penta·His HRP Conjugate Kit (QIAGEN, catalog number: 34460 )
Note: This is antibody against the His-tag and we used it for Western-blot analysis of 6x His-tagged MgtA. This antibody is conjugated to horseradish peroxidase (HRP), so there is no requirement for the use of secondary antibody. - HEPES (Sigma-Aldrich, catalog number: H3375 )
- Potassium sulfate (K2SO4) (Sigma-Aldrich, catalog number: P9458 )
- Deoxyribonuclease I (Sigma-Aldrich, catalog number: DN25 )
- Phenylmethylsulfonyl fluoride (Sigma-Aldrich, catalog number: P7626 )
- Dodecyl-β-D maltoside (Chemical point, catalog number: CP69227-93-6-BULK )
- Imidazole (Sigma-Aldrich, catalog number: 56750 )
- Dithiothreitol (Biosynth, catalog number: D-8200 )
- Potassium hydroxide (KOH) (Sigma-Aldrich, catalog number: 221473 )
- Trizma® base (biotechnology performance certified) (Sigma-Aldrich, catalog number: T6066 )
- PageRulerTM prestained protein ladder (Thermo Fisher Scientific, Thermo ScientificTM, catalog number: 26616 )
- LB broth (see Recipes)
- LB kanamycin plate (see Recipes)
- Growth media (see Recipes)
- 1x SDS-PAGE loading buffer (see Recipes)
- Buffer A (see Recipes)
- Buffer B (see Recipes)
- Buffer C (see Recipes)
- Buffer D (see Recipes)
- Buffer E (see Recipes)
Equipment
- Incubator
- Baffled bottom Erlenmeyer flask (500 ml) (Saveen & Werner, catalog number: 00125-54 )
Note: These flasks have four baffles (flow directing ridges). It interrupts the circular flow of media and increases its gas exchange surface, thereby increasing the oxygenation of culture media when compared to a normal Erlenmeyer flask. - Bottle with screw caps, used for fermentation (Saveen & Werner, catalog number: 88-2L )
- LEX (Large-scale Expression) 48 bioreactor system (Epiphyte Three, Harbinger Biotechnology and engineering, model: LEX-48 Bioreactor )
- High pressure homogenizer (used to lyse cells) (Avestin, model: EmulsiFlex C3 )
- Potter-Elvehjem-type tissue homogenizer (30 ml) (WHEATON, catalog number: 358049 )
- 70 ml polycarbonate ultracentrifugation tube assembly (Beckman Coulter, catalog number: 355622 )
- 45 Ti rotor (Beckman Coulter, catalog number: 339160 )
- OptimaTMXE (Beckman Coulter, catalog number: A99833 )
- Column HP Histrap, 5 ml (GE Healthcare, catalog number: 17-5248-02 )
- HiLoad 16/600 Superdex 200 pg column (GE Healthcare, catalog number: 28-9893-35 )
- Vivaspin 20 MWCO 50,00 PES (Saveen & Werner, catalog number: VS2032 )
Software
- Image LabTM software
- UNICORN software associated with ÄKTA purifier
Procedure
- Overexpression of MgtA
The mgtA gene was amplified from E. coli DH5α genomic DNA using PfuUltra II Fusion HS DNA polymerase with the primers described earlier (Subramani et al., 2016). The reaction mixture and PCR conditions were made as suggested by the PfuUltra II Fusion HS DNA polymerase manual. The amplified gene was inserted between NcoI and XhoI restriction sites in pETM11 vector (EMBL). Thus the expressed MgtA will include 6x His tag followed by a tobacco etch virus (TEV) protease site at the N-terminus.- Transform the pETM11(mgtA) plasmid into C43(DE3) competent cells and plate on LB kanamycin plate and incubate at 37 °C for 16 h.
- Select 5 colonies at random and inoculate in a 500 ml baffled bottom Erlenmeyer flask containing 200 ml LB media with 50 µg/ml of kanamycin. Incubate the flask at 37 °C with 150 rpm for 16 h.
- For large scale over expression of MgtA in C43(DE3) cells, we used the LEX 48 bioreactor system (Harbinger Technology LEXTM).
Note: The LEX system delivers filtered compressed air through a sparger that passes through the screw cap. It provides both aeration and mixing of bacterial culture in the screw cap bottle. The culture flasks are placed in a temperature controlled water bath and the LEX 48 can accommodate 24 x 2 L standard screw cap bottles. Normally only 75% of the bottle capacity is used to avoid overflow of media during aeration. - Inoculate 1% of overnight culture into 2 L screw cap bottles containing 1.5 L of growth media, 50 µg/ml of kanamycin and 1.5 ml of polypropylene glycol (PPG) P2000 (antifoaming agent).
- Place the bottles in the Lex 48 bioreactor water bath. Set the temperature of the water bath at 37 °C and initiate airflow through the spargers.
- Measure the optical density at 600 nm (OD600) every hour. The antifoaming agent PPG P2000 can become cloudy under these conditions and can be cleared upon incubation in ice for 5 min. Therefore, prior to the measurement at OD600, withdraw 1 ml of culture from the 2 L bottles and incubate on ice for 5 min.
- When the OD600 is ~0.6-2, stop the airflow and cool the water bath to 17 °C by adding ice. Leave the culture bottles at 17 °C for 30 min to ensure an equivalent temperature of the bacterial culture. In order to check the level overexpression by Western blot (see step 12), pellet 1 ml of bacterial culture at 12,000 x g, 4 °C and mark as SUI (uninduced).
- Add 1 mM of IPTG and an additional 1.5 ml of PPG P2000 to each bottle. Begin the airflow and leave the bottles at 17 °C for 16 h with continuous airflow.
- To terminate the induction, transfer the culture bottles to ice. Pellet 1 ml of bacterial culture from each bottle at 12,000 x g, 4 °C and mark as SI (induced) for Western blot analysis.
- Centrifuge rest of the bacterial culture at 7,000 x g for 20 min at 4 °C and divide the resulting cell pellet into 50 g aliquots in 100 ml disposable cups and store at -20 °C until further use.
- Suspend the SUI and SI sample pellets in 1x SDS-PAGE loading buffer to normalize the OD600 to 10 and incubate at RT for 5 min. Centrifuge the lysed cells at 20,000 x g for 20 min at 4 °C. For Western blot analysis use 10 µl of the supernatant and blot against Penta·His HRP conjugated antibody to check MgtA overexpression.
Note: Confirmation of MgtA overexpression by Western blot against Penta·His HRP antibodies is highly recommended before beginning protein purification process as the detergents used during purification process are expensive and any problem associated with overexpression can be identified at this stage. Do not boil the samples used for SDS-PAGE and Western blot analysis.
- Transform the pETM11(mgtA) plasmid into C43(DE3) competent cells and plate on LB kanamycin plate and incubate at 37 °C for 16 h.
- Cell lysis and solubilisation
- Thaw the frozen cell pellet on ice and suspend in buffer A at 1:10 (wt/vol) ratio.
- Lyse the cells using EmulsiFlex-C3, a high pressure homogenizer (HPH) by passing the cell suspension three times at 11,000 psi. Always store the samples on ice unless otherwise stated.
Note: We did not test the other lysis methods like sonication and bead beater. But we believe, that they could also be used. The protein yield should be expected to vary depending on the lysis method. - Centrifuge the cell lysate at 20,000 x g for 20 min at 4 °C to remove unlysed cells and inclusion bodies.
- Collect the supernatant and centrifuge at 100,000 x g for 2 h at 4 °C to pellet both the inner and outer membrane (mixed membranes).
Note: This step requires ultracentrifuge with the rotors and tubes selected according to the volume of the sample. For example, if the sample size is 70 ml, then 45 Ti rotor and 70 ml polycarbonate ultracentrifugation tube can be used with ultracentrifuge OptimaTMXE. It is important to fill the tubes used for ultracentrifugation until the neck of the tube with possibly no gap between the liquid and cap of the tube to avoid the implosion of tubes during centrifugation. - Suspend the resulting mixed membrane pellet in buffer B using Potter-Elvehjem-type tissue homogenizer with 10 strokes. Use 1:10 (wt/vol) mixed membrane to buffer B ratio. Before adding the required amount of buffer B to membrane pellet, remove 10 ml to dissolve the detergent.
Note: The membrane suspension can be flash frozen with liquid nitrogen and stored at -80 °C if not used immediately. - We used 1% β-dodecyl maltoside (β-DDM) to solubilize the membrane proteins. Weigh the required amount of detergent powder and dissolve in the reserved 10 ml of buffer B.
- Add the dissolved β-DDM drop wise into the mixed membrane suspension while stirring at 150 rpm, 4 °C. The suspension is left to stir for another 60 min at 4 °C.
- Thaw the frozen cell pellet on ice and suspend in buffer A at 1:10 (wt/vol) ratio.
- Purification of MgtA
- Prior to use, wash the Histrap HP 5 ml column with 5 column volume (CV) of degassed MQ water and equilibrate with 10 CV of buffer C.
- Add 20 mM imidazole (pH 7.6) to the sample prepared in step B7 and load on the Histrap column at 2 ml/min.
- After loading solubilized membrane sample, wash the Histrap column with 10 CV of buffer C and 3 CV of 15% buffer D at 3 ml/min. We observed a small peak (Figure 1A, peak 1) while washing with 15% buffer D.
- Elute MgtA from the Histrap column by passing 50% buffer D at 3 ml/min. We observed a relatively large peak (Figure 1A, peak 2) while washing with buffer D.
- Take 5 µl sample from each fraction and perform SDS-PAGE electrophoresis with 12% polyacrylamide gels.
- We observed that Peak 1 contained a small proportion of MgtA along with other impurities, whereas peak 2 contained > 80% pure MgtA, as assessed by SDS-PAGE (Figure 1A).
- Pool the fractions corresponding to peak 2 and concentrate to 6 mg/ml using a vivaspin 20 (protein concentrator with 50 kDa cutoff).
- While concentrating the fractions, prepare a HiLoad 16/600 Superdex 200 pg column connected to the ÄKTA purifier by washing with 1.5 CV of degassed MQ and 1.5 CV of buffer E at 0.5 ml/min.
- Load 1 ml of concentrated MgtA (6 mg/ml) on the HiLoad 16/600 Superdex 200 pg column at 0.5 ml/min and pass 1.5 CV of buffer E.
- We observed a small peak at the void volume (44 ml), followed by a major monodisperse peak (Figure 1B, peak S1) at ~55 ml.
- For SDS-PAGE analysis, load 5 µl of fractions corresponding to the observed peaks on 12% polyacrylamide gels. We observed that the void peak contained mostly impurities, whereas peak S1 contained MgtA at > 95% purity as analyzed by SDS-PAGE (Figure 1B).
- Pool the fractions from peak S1 and concentrate to 3 mg/ml using vivaspin 20.
- Divide the concentrated MgtA as 50 µl aliquots and flash freeze with liquid nitrogen and store at -80 °C.
- In our experience, the protein was stable under these conditions for more than 1 year.
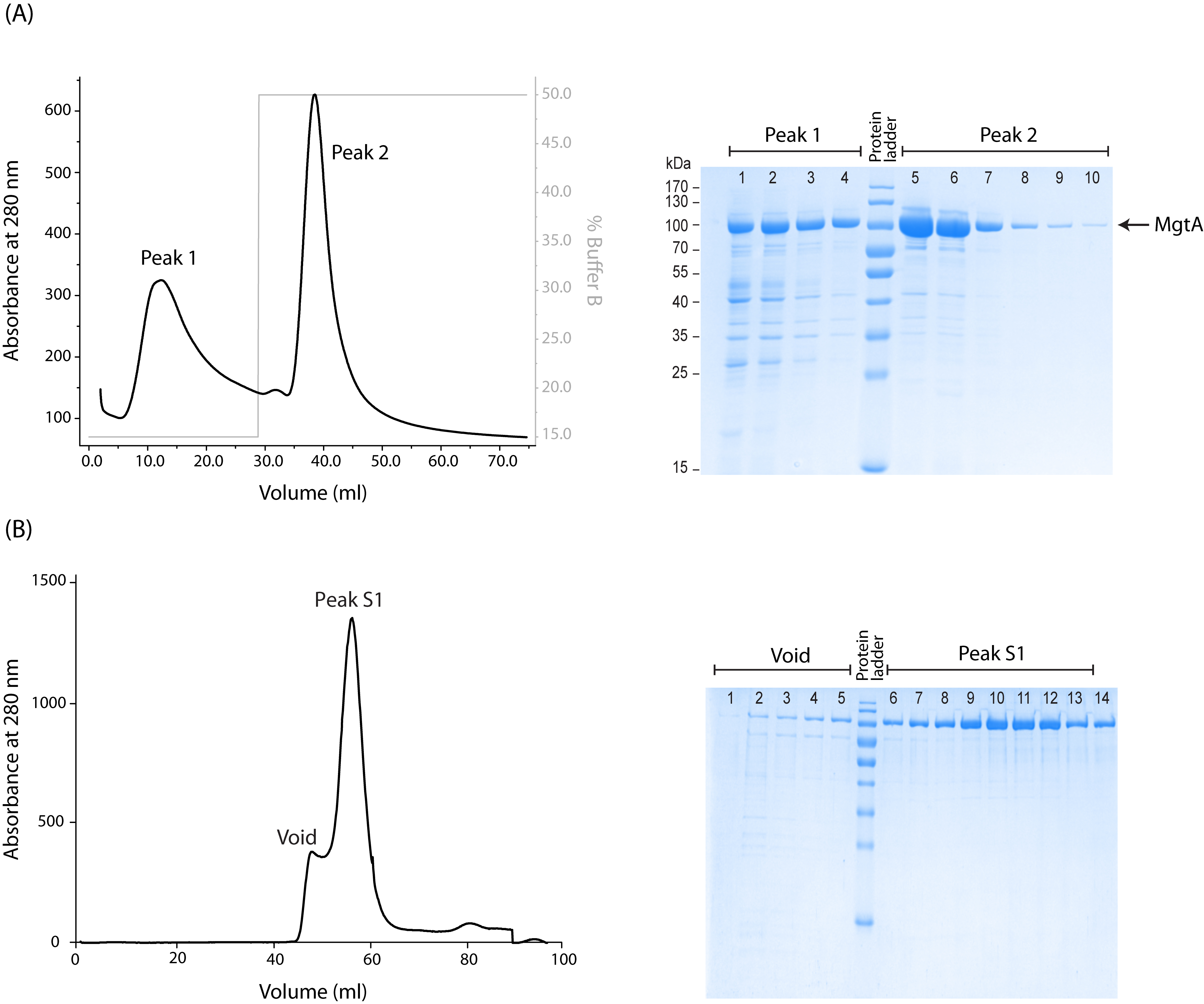
Figure 1. Representative figures of MgtA purification profile. A. Chromatogram of Histrap FF affinity column (left). The fractions from Peak 1 (lanes 1-4) and Peak 2 (lanes 6-11) were analyzed using SDS-PAGE and the Coomassie stained gel (right) shows the purity of the respective peaks. B. Chromatogram of size exclusion chromatography (left). The fractions from void (lanes 1-5) and Peak S1 (lanes 7-14) were analyzed using SDS-PAGE and the Coomassie stained gel (right) shows the purity of the respective peaks. In both SDS-PAGE gel pictures, MgtA runs at 100 kDa.
- Prior to use, wash the Histrap HP 5 ml column with 5 column volume (CV) of degassed MQ water and equilibrate with 10 CV of buffer C.
Data analysis
The data for chromatograms in Figure 1 were obtained from UNICORN software associated with ÄKTA purifier and the graphs were plotted using GraphPad Prism6. All the SDS-PAGE gels were scanned using Bio-Rad ChemiDoc XRS+ system and analyzed using Image LabTM software.
Recipes
- LB broth
Mix 40 g of LB broth powder in 1 L of MQ and autoclave - LB kanamycin plate
Mix 40 g of LB broth powder and 7.5 g of LB agar and autoclave
Prepare LB plate with 50 µg/ml kanamycin - Growth media
40 g of LB broth powder from Sigma-Aldrich (see Materials and Reagents)
10 g of Fistex LS Ferm yeast extract
Add 1 L of MQ and autoclave. Before inoculation, add 10 ml of 1 M Tris-Hcl pH 7.4 and 5 ml of glycerol - 1x SDS-PAGE loading buffer
2% SDS
10% glycerol
5% β-mercaptoethanol
0.002% bromophenol blue
63 mM Tris-HCl, pH 6.8 - Buffer A
50 mM HEPES, pH 7.0 (KOH)
100 mM K2SO4
10% glycerol
1 mM PMSF
5 mM β-mercaptoethanol
1 μg/ml DNase - Buffer B
25 mM HEPES, pH 7.0 (KOH)
100 mM K2SO4
5% glycerol
1 mM PMSF
5 mM β-mercaptoethanol - Buffer C
25 mM HEPE, pH 7.0 (KOH)
100 mM K2SO4
5% glycerol
1 mM PMSF
5 mM β-mercaptoethanol
20 mM Imidazole, pH 7.6
3 CMC β-DDM - Buffer D
25 mM HEPES, pH 7.0 (KOH)
100 mM K2SO4
5% glycerol
1 mM PMSF
5 mM β-mercaptoethanol
300 mM Imidazole pH 7.6
3 CMC β-DDM - Buffer E
25 mM HEPES, pH 7.0 (KOH)
100 mM K2SO4
5% glycerol
1 mM dithiothreitol
3 CMC β-DDM
Note: The buffers used for chromatography were always filtered using the NalgeneTM reusable bottle top filter connected to vacuum pump. Buffers were degassed with the same set up, but by closing the filter chamber while the vacuum still on and with constant stirring. Buffers were degassed until no bubbles were observed in the solution.
Acknowledgments
The Norwegian Research Council Funding (F-RIMEDBIO) #ES486454 and NCMM core Funding supported this study.
References
- Faraco, M., Spelt, C., Bliek, M., Verweij, W., Hoshino, A., Espen, L., Prinsi, B., Jaarsma, R., Tarhan, E., de Boer, A. H., Di Sansebastiano, G. P., Koes, R. and Quattrocchio, F. M. (2014). Hyperacidification of vacuoles by the combined action of two different P-ATPases in the tonoplast determines flower color. Cell Rep 6(1): 32-43.
- Groisman, E. A., Hollands, K., Kriner, M. A., Lee, E. J., Park, S. Y. and Pontes, M. H. (2013). Bacterial Mg2+ homeostasis, transport, and virulence. Annu Rev Genet 47: 625-646.
- Kato, A., Tanabe, H. and Utsumi, R. (1999). Molecular characterization of the PhoP-PhoQ two-component system in Escherichia coli K-12: identification of extracellular Mg2+-responsive promoters. J Bacteriol 181(17): 5516-5520.
- Soncini, F. C., Garcia Vescovi, E., Solomon, F. and Groisman, E. A. (1996). Molecular basis of the magnesium deprivation response in Salmonella typhimurium: identification of PhoP-regulated genes. J Bacteriol 178(17): 5092-5099.
- Subramani, S., Perdreau-Dahl, H. and Morth, J. P. (2016). The magnesium transporter A is activated by cardiolipin and is highly sensitive to free magnesium in vitro. Elife 5.
Article Information
Copyright
Subramani and Morth. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Subramani, S. and Morth, J. P. (2016). Heterologous Expression and Purification of the Magnesium Transporter A (MgtA) in Escherichia coli. Bio-protocol 6(22): e2001. DOI: 10.21769/BioProtoc.2001.
- Subramani, S., Perdreau-Dahl, H. and Morth, J. P. (2016). The magnesium transporter A is activated by cardiolipin and is highly sensitive to free magnesium in vitro. Elife 5.e11407.
Category
Microbiology > Microbial biochemistry > Protein > Isolation and purification
Biochemistry > Protein > Expression
Biochemistry > Protein > Isolation and purification
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









