- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measuring Auxin Transport Capacity in Seedling Roots of Medicago truncatula
Published: Vol 6, Iss 12, Jun 20, 2016 DOI: 10.21769/BioProtoc.1842 Views: 10076
Reviewed by: Marisa RosaScott A M McAdamMoritz Bomer

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Analysis of Monosaccharides from Arabidopsis Seed Mucilage and Whole Seeds Using HPAEC-PAD
Gillian H. Dean [...] George W. Haughn
Dec 20, 2019 5756 Views

Extraction and Quantification of Plant Hormones and RNA from Pea Axillary Buds
Da Cao [...] Christine A. Beveridge
Oct 5, 2022 2969 Views
Targeting Ultrastructural Events at the Graft Interface of Arabidopsis thaliana by A Correlative Light Electron Microscopy Approach
Clément Chambaud [...] Lysiane Brocard
Jan 20, 2023 2981 Views
Abstract
Measurement of auxin transport capacity provides quantitative data on the physiological machinery involved in auxin transport within plants. This technique is easy to perform and gives quick results. Radiolabelled auxin (indole-3-acetic-acid) is fed into the roots of Medicago truncatula via an agar block. The resulting radioactivity from radiolabelled auxin uptake in the roots is measured with a liquid scintillation counter. Here, we describe the measurement of auxin transport capacity around the nodulation susceptible zone in young seedling roots of M. truncatula in response to rhizobia inoculation. Similar assays could be adapted in other plant species and to answer other biological questions.
Keywords: Indole-3-acetic acidMaterials and Reagents
- Petri dish (150 x 15 mm) (BD, catalog number: 351058 )
Note: Currently, it is “Corning, catalog number: 351058”. - Petri dish (35 x 10 mm) (BD, catalog number: 351008 )
Note: Currently, it is “Corning, catalog number: 351008”. - SandBlaster fine 320 grit sandpaper (3M, catalog number: 70-0710-1785-2 )
- 50 ml Falcon tube (Greiner Bio One International GmbH, catalog number: 227261 )
- Waste bottle
- Aluminium foil (Confoil)
- 1.5 ml Eppendorf tube
- Kimwipes
- 6 ml scintillation vials (Pico Prias vial) (PerkinElmer, catalog number: 6000192 )
- Parafilm strips
- Medicago truncatula seeds (cultivar Jemalong A17) (SARDI)
- Sinorhizobium meliloti (rhizobia) culture
- Sodium hypochlorite solution (Bleach) (Pelikan)
- Sterilised distilled water
- Ethanol (EMD Millipore Corporation, EMSURE®)
- Agarose (AMRESCO, catalog number: 0710 )
- Emulsifier-safe, 2 x 5 L (scintillation fluid) (PerkinElmer, catalog number: 6013389 )
- CaCl2.2H2O (Sigma-Aldrich, catalog number: C3306 )
- Magnesium sulfate heptahydrate (MgSO4.7H2O) (Sigma-Aldrich, catalog number: M1880 )
- KH2SO4 (Sigma-Aldrich, catalog number: P0772 )
- Na2HPO4.2H2O (Sigma-Aldrich, catalog number: 71643 )
- Fe citrate (Sigma-Aldrich, Fluka®, catalog number: 44941 )
Note: It is also named “Iron(III) citrate tribasic monohydrate” on Sigma-Aldrich website. - H3BO3 (Sigma-Aldrich, catalog number: B6768 )
- MnSO4.4H2O (Merckmillipore, catalog number: 102786 )
- ZnSO4.7H2O (Sigma-Aldrich, catalog number: Z4750 )
- CuSO4.5H2O (Sigma-Aldrich, catalog number: C7631 )
- H2MoO4 (Sigma-Aldrich, catalog number: 232084 )
- Na2MoO4.2H2O (Sigma-Aldrich, catalog number: 331058 )
- Agar (TEKNOVA, catalog number: A7777 )
- Tritium-labelled indole-3-acetic acid (3H-IAA) blocks (pH 4.8) (American Radiolabelled Chemicals, catalog number: ART 0340 )
- Fahraeus media (see Recipes)
- 3H-IAA blocks (see Recipes)
Equipment
- Plant growth chamber, set at 25 °C, 16/8 h day/night cycle with 150 μmol m−2 s−1 light intensity (Thermo Fisher Scientific)
- Scintillation counter (PerkinElmer, model: Tri-Carb 2800TR )
- Scintillation racks (PerkinElmer)
- Pipettes (Eppendorf AG)
- Scissors (ECKERSLEY’S)
- Fine forceps (Dumont and Fils) (D'OUTILS DUMONT S.A.)
- Rotating shaker (SeouLin Bioscience, model: MyLabTM Intelli-Mixer )
- Platform shaker (Eppendorf AG, New BrunswickTM, model: INNOVA 2100 )
- Microwave
Procedure
- Medicago truncatula cultivar Jemalong A17 seeds (or other Medicago cultivars as required) are germinated and grown in vitro on 150 mm diameter Petri dishes containing Fåhraeus media, as described in Ng et al. (2015) (Figure 1).
- Seeds are first scarified on sandpaper and surface-sterilized in 6% (w/v) sodium hypochlorite for 10 min on a rotating shaker, then rinsed with sterile distilled water at least five times.
- After incubation for 6 h in sterile distilled water in a Falcon tube on a rotating shaker, seeds are spread onto Petri dishes containing Fåhraeus agar, placed at 4 °C for 48 h to synchronize germination, and subsequently germinated at 20 °C overnight.
- Seedlings of ~1 cm length are placed onto fresh Petri dishes containing Fåhraeus agar at seven seedlings per plate in a sterile laminar flow hood, and grown for four days at 25 °C with a 16 h day, 8 h night cycle and light intensity of 150 μmol m−2 s−1 in a plant growth chamber. Petri dishes are arranged vertically so that roots will grow along the surface of the media, with a black cardboard interspersed between plates to prevent the roots from being exposed to light. Discard any seedlings that have grown underneath the agar surface.
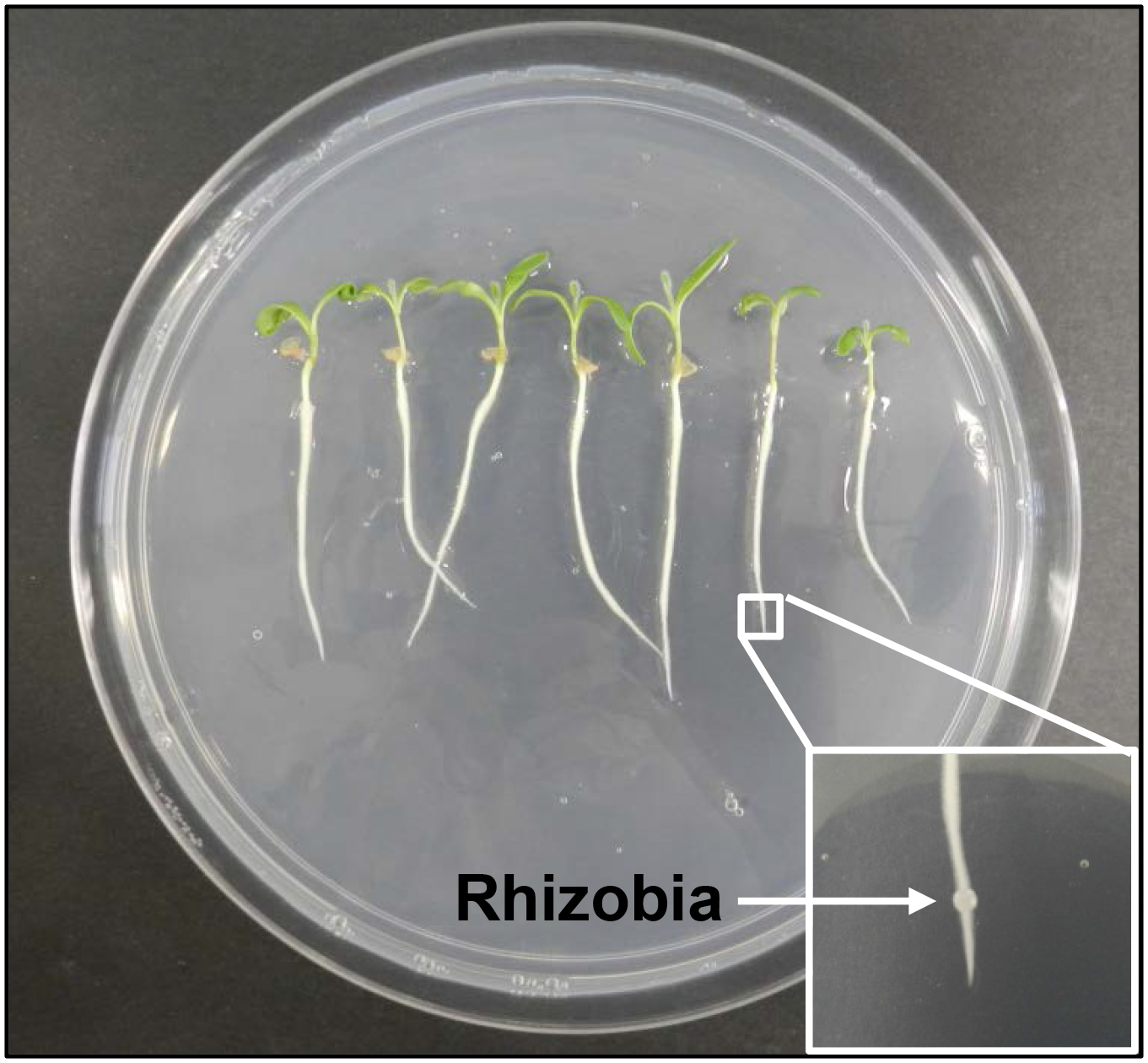
Figure 1. Growth and inoculation of Medicago truncatula seedlings with rhizobia. Multiple seedlings are grown on a single Petri dish. A fine drop of rhizobia culture (~1 μl; OD600 = 0.1) is inoculated onto the nodulation-susceptible zone on each root (inset).
- Seeds are first scarified on sandpaper and surface-sterilized in 6% (w/v) sodium hypochlorite for 10 min on a rotating shaker, then rinsed with sterile distilled water at least five times.
- If the effect of rhizobial infection is of interest, four-day-old M. truncatula seedlings are spot-inoculated with Sinorhizobium meliloti (S. meliloti) (rhizobia) at the nodulation susceptible zone, i.e. ~3 mm above the root tip, in the zone of the root where root hairs are just emerging and still elongating (Figure 1).
- Prior to inoculation, the inoculation spot is marked on the underside of the Petri dish. Note that root and agar surface need to be dry to avoid movement of the inoculum. When comparing different experiments, it is important to carry out the inoculations and auxin transport measurements at the same time of the day, as we have observed changes in total auxin transport at different times during the light/dark period.
- If the auxin transport assay is adapted for other biological questions, other treatments of the seedlings can be included at this step. For example:
- The effect of mycorrhizal infection of the roots. In this case, the roots are first locally infected with mycorrhizal spores prior to auxin transport measurement.
- The effect of natural or synthetic auxin transport modulators on auxin transport, such as flavonoids or 1-N-naphthylphthalamic acid, respectively. In this case, the roots can be treated with the chemicals in the presence or absence of rhizobia.
- In addition to acropetal auxin transport measurement described here, basipetal auxin transport (from the root tip towards the root base) could also be investigated with a slight modification to the protocol described here [please refer to Supplemental Figure 2 in Ng et al. (2015)].
- While the inoculation with rhizobia described here has been found to be optimal at a 24 h incubation time, other treatments could be adapted for shorter or longer incubation times as required, as long as the seedlings do not outgrow the length of the Petri dishes, to ensure the root tip area remains healthy and straight.
- 3H-IAA agarose blocks are prepared in a Petri dish (35 x 10 mm) and cut into blocks with dimensions of 2 mm x 2 mm x 2 mm with a sterile scalpel (Figure 2A and B).
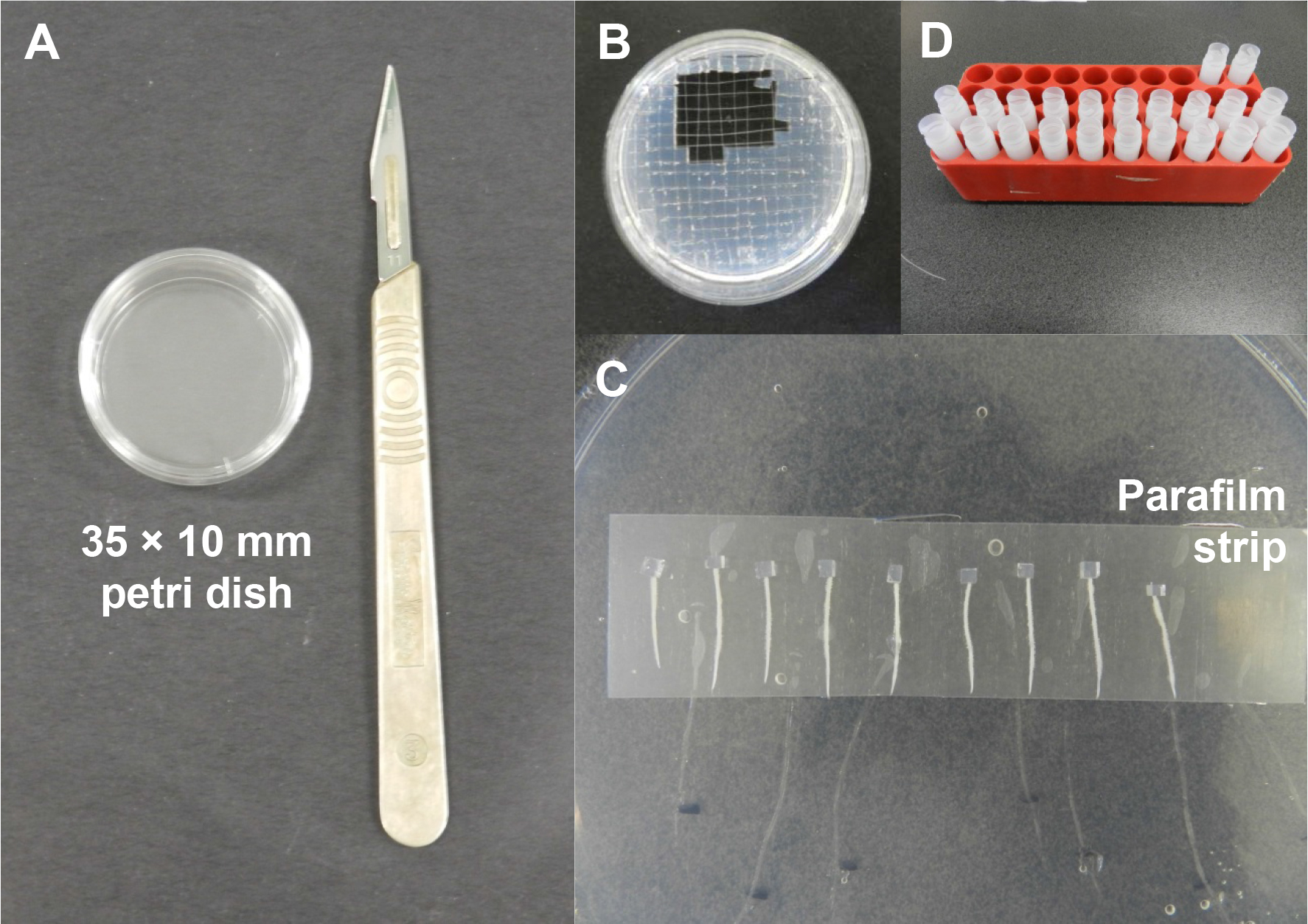
Figure 2. Preparation of radiolabelled auxin blocks and the auxin transport assay. A. A 35 x 10 mm Petri dish and a sterile scalpel for preparation of the radiolabelled auxin blocks. B. A scalpel is used to cut the cooled radiolabelled auxin agar into smaller blocks (2 mm x 2 mm x 2 mm). C. Root segments are arranged on a parafilm strip and a small radiolabelled auxin agar block is placed tightly against the cut end of each root segment using a fine forceps. D. After incubation, samples are collected in individual scintillation vials containing scintillation fluid. - To investigate auxin transport capacity at 24 h post-S. meliloti inoculation, at 18 h post-inoculation, the roots are excised 8 mm above the inoculation spot. The root segments (containing the root tip) are placed on a parafilm strip. A block of 3H-IAA agarose is positioned at the cut end of each root segment (Figure 2C). It is important to check that the cut surface of the agar block is tightly in contact with the cut root surface to ensure that there is no gap that could prevent auxin transport into the root segment. The root segments are left to incubate for 6 h in the dark at room temperature.
- 6 ml scintillation vials containing 2 ml of scintillation fluid each are prepared (Figure 2D).
- The first 4 mm of the root from the 3H-IAA agarose block is discarded. The adjacent 4 mm segment is cut with a clean scalpel and then placed in a vial containing scintillation fluid using a clean forceps, making sure the root segment is fully immersed in the fluid. The final 4 mm segment (with the excess root tip discarded) is placed in a separate scintillation vial containing scintillation fluid.
- The forceps should be regularly rinsed in ethanol and dried with Kimwipe tissues to prevent carryover of radioactivity between samples.
- The vials are covered with lids and incubated on a platform shaker at room temperature overnight to extract the radiolabelled auxin into the scintillation fluid.
- Vials containing the root samples are arranged in scintillation racks.
- Three vials with scintillation fluid but without root segments are included at the start of the run; these are used to subtract the averaged background.
- Samples are analysed in a TriCarb 2800TR scintillation counter for 1 min each using the default parameters for 3H radionuclide decay.
- Results (in counts per min) are exported as a spreadsheet and analysed. An example data set is shown in Table 1. If necessary, outliers need to be removed, i.e., those readings that lie below the average background, for example the sample with a reading of ‘9’ in the list below. These outliers are typically due to one of the agar blocks containing the radiolabelled auxin losing contact with the root.
Table 1. An example data set of Medicago truncatula roots treated with water (mock treatment) or the synthetic auxin transport inhibitor 1-N-naphthylphthalamic acid (NPA), both in the presence of rhizobia. Data are presented as counts/min.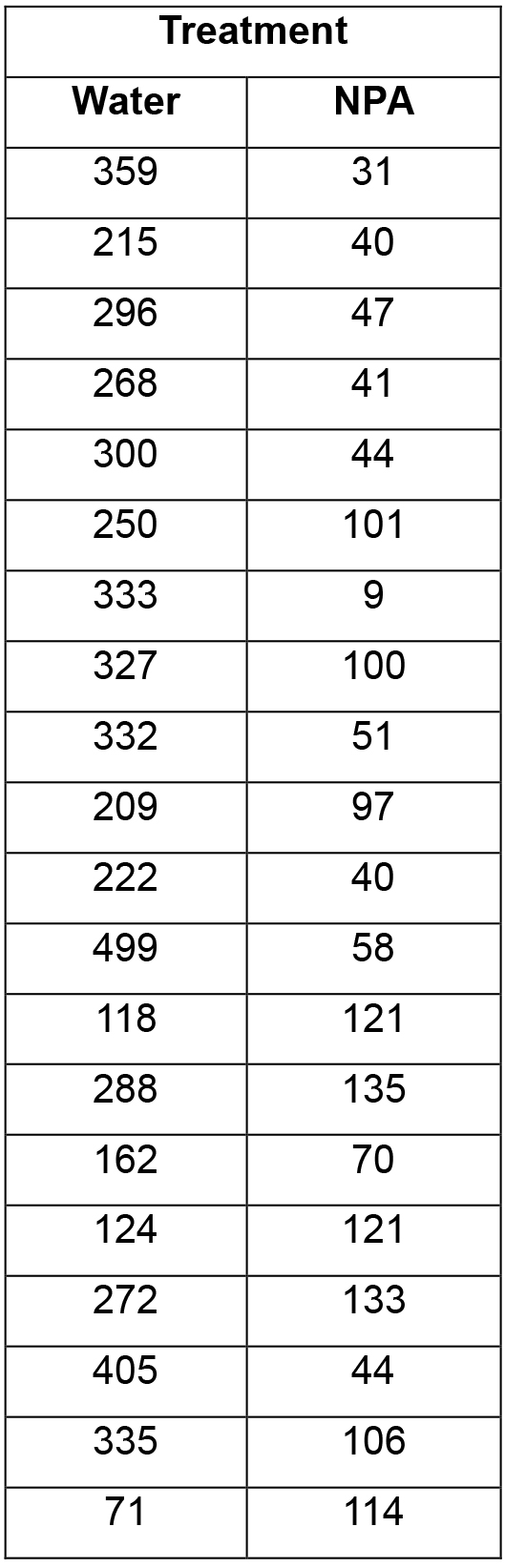
- These data sets typically result in normally-distributed data that can be analyzed using a Student’s t-test (when comparing two treatments) or analysis of variance (when comparing several treatments). We suggest a large number of seedlings (at least 15-20) for each treatment as there is some variation in auxin transport between seedlings.
- Ensure all radioactive waste is discarded in special radiation waste bins.
Recipes
- Fåhraeus media (per litre)
132 mg CaCl2·2H2O
120 mg MgSO4·7H2O
100 mg KH2SO4
75 mg Na2HPO4·2H2O
5 mg Fe Citrate
2.86 mg H3BO3
2.03 mg MnSO4·4H2O
220 μg ZnSO4·7H2O
80 μg CuSO4·5H2O
90 μg H2MoO4
121 μg Na2MoO4·2H2O
10 g agar (Gelita J3 grade)
Autoclave, cool, pour into Petri dishes (~60-70 ml in each plate)
Stock solutions are stored at 4 °C and can be used for up to one year.
In our experience, Fåhraeus media can be stored for up to six months at 4 °C. - 3H-IAA blocks
- Melt 1% water agarose in a microwave, adjusted to pH 4.8, and cool to around 50 °C.
- Prepare 7.5 μl 3H-IAA stock by diluting in 20 μl ethanol in an Eppendorf tube. Keep dark by wrapping the tube in aluminium foil.
- Carefully mix diluted 3H-IAA mixture into 2 ml melted 1% water agarose in a Petri dish (35 x 10 mm). It is important to achieve a similar amount of radiolabelled auxin in each block. This should be tested by immersing a number of individual blocks from different areas of the plate in scintillation fluid, incubating on a shaker overnight and measuring radioactivity.
- Allow mixture to air dry for 15 min.
- Cut into 2 mm x 2 mm x 2 mm blocks with a scalpel, using a printed grid as a template for block size.
- Keep blocks cool (4 °C) and in the dark until needed by wrapping the plate in aluminium foil. Always prepare blocks on the same day as carrying out the transport assays. Discard unused blocks into radiation waste.
Acknowledgments
This protocol was adapted from Lewis and Muday (2009), van Noorden et al. (2006), Wasson et al. (2006), Plet et al. (2011) and was performed by Ng et al. (2015). This work was supported by an Australian Research Council Future Fellowship awarded to Ulrike Mathesius (FT100100669).
References
- Lewis, D. R. and Muday, G. K. (2009). Measurement of auxin transport in Arabidopsis thaliana. Nat Protoc 4(4): 437-451.
- Ng, J. L., Hassan, S., Truong, T. T., Hocart, C. H., Laffont, C., Frugier, F. and Mathesius, U. (2015). Flavonoids and auxin transport inhibitors rescue symbiotic nodulation in the Medicago truncatula cytokinin perception mutant cre1. Plant Cell 27(8): 2210-2226.
- Plet, J., Wasson, A., Federico, A., Le Signor, C., Baker, D., Mathesius, U., Crespi, M. and Frugier, F. (2011). MtCRE1-dependent cytokinin signalling integrates bacterial and plant cues to coordinate symbiotic nodule organogenesis in Medicago truncatula. Plant J. 65(4): 622-633.
- van Noorden, G. E., Ross, J. J., Reid, J. B., Rolfe, B. G. and Mathesius, U. (2006). Defective long-distance auxin transport regulation in the Medicago truncatula super numeric nodules mutant. Plant Physiol 140(4): 1494-1506.
- Wasson, A. P., Pellerone, F. I. and Mathesius, U. (2006). Silencing the flavonoid pathway in Medicago truncatula inhibits root nodule formation and prevents auxin transport regulation by rhizobia. Plant Cell 18(7): 1617-1629.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ng, J. L. P. and Mathesius, U. (2016). Measuring Auxin Transport Capacity in Seedling Roots of Medicago truncatula. Bio-protocol 6(12): e1842. DOI: 10.21769/BioProtoc.1842.
Category
Plant Science > Plant biochemistry > Plant hormone
Plant Science > Plant physiology > Tissue analysis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










