- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
An in vitro Transcription/translation System for Detection of Protein Interaction
Published: Vol 6, Iss 9, May 5, 2016 DOI: 10.21769/BioProtoc.1800 Views: 17487
Reviewed by: Zhaohui LiuPablo Bolanos-VillegasAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
General Maintenance and Reactivation of iSLK Cell Lines
Ariana C. Calderón-Zavala [...] Ekaterina E. Heldwein
Jun 5, 2025 1936 Views
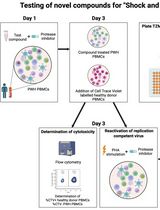
Inducible HIV-1 Reservoir Reduction Assay (HIVRRA), a Fast and Sensitive Assay to Test Cytotoxicity and Potency of Cure Strategies to Reduce the Replication-Competent HIV-1 Reservoir in Ex Vivo PBMCs
Jade Jansen [...] Neeltje A. Kootstra
Jul 20, 2025 2492 Views
Assembly and Mutagenesis of Human Coronavirus OC43 Genomes in Yeast via Transformation-Associated Recombination
Brett A. Duguay and Craig McCormick
Aug 20, 2025 3077 Views
Abstract
Studying protein-protein interaction is crucial to understand the fundamental processes of molecular biology. High-throughput screening, such as immunoprecipitation followed by proteomic analysis, allows for the identification of numerous candidate partners that might interact with a selected protein. However, experimental validation of protein-protein interaction requires conventional cloning and recombinant protein expression/purification, which are complicated and labor-intensive techniques. Here, we demonstrate an efficient experimental pipeline for verifying protein-protein interactions between a bait protein using the example of Odontoglossum ringspot virus (ORSV) capsid protein (CP) and the host CP-binding protein. These candidate CP-binding proteins were identified through high-throughput proteomic and transcriptomic approaches. Using the TOPO cloning strategy, each candidate gene was cloned into an expression vector for the expression of His-tagged recombinant proteins in a single step of an in vitro transcription/translation system. Such expressed His-tagged candidates can be used as prey with the CP bait protein in a co-immunoprecipitation (co-IP) assay to verify their physical interaction. Without the need for traditional protein expression and purification, this pipeline simplifies the validation process and provides a solution for high-throughput protein-protein interaction studies.
Keywords: Protein-protein interactionMaterials and Reagents
- TOPO cloning for the protein expression construct
- Heat shock-competent cell of E. coli strain DH5α
- pEXP5-CT/TOPO TA Expression Kit (Thermo Fisher Scientific, InvitrogenTM, catalog number: V960-06 )
- Forward/reversed primers for candidate genes
- ExTaq polymerase (Takara Bio Company, catalog number: RR001A )
- UltraPure agarose (Thermo Fisher Scientific, InvitrogenTM, catalog number: 16500 )
- QIAEX® II Gel Extraction Kit (QIAGEN, catalog number: 20051 )
- Luria-Bertani (LB) broth and LB agar plates
- Ampicillin sodium salt (Sigma-Aldrich, catalog number: A0166 )
- T7 forward primer (5’-TAATACGACTCACTATAGGG-3’)
- PrestoTM Mini Plasmid Kit (Geneaid Biotech Ltd., catalog number: PDH300 )
- Heat shock-competent cell of E. coli strain DH5α
- Expression of recombinant protein via single-step in vitro transcription/translation
- ExpresswayTM Cell-Free E. coli Expression System (Thermo Fisher Scientific, InvitrogenTM, catalog number: K9901-00 )
- Polyacrylamide gel electrophoresis (PAGE) system (Bio-Rad Laboratories, catalog number: 1610158 )
- Anti-His (C-term)-HRP monoclonal antibody (Thermo Fisher Scientific, InvitrogenTM, catalog number: R931-25 )
- COOMASSIE Brilliant Blue G-250 (VWR International, J.T.Baker, catalog number: F789 )
- ExpresswayTM Cell-Free E. coli Expression System (Thermo Fisher Scientific, InvitrogenTM, catalog number: K9901-00 )
- Protein-protein interaction and co-immunoprecipitation (co-IP)
- pGEX-4T-1 DNA vector (GE Healthcare, catalog number: 28-9545-49 )
- Bait protein with tag [herein, GST-CP (recombinant of ORSV CP protein with the N-terminus fused to glutathione S-transferase)
- Anti-bait antibody [herein, anti-CP antibody (Lee and Chang, 2008)]
- Protein G PLUS-Agarose (Santa Cruz Biotechnology, catalog number: sc-2002 )
- Anti-tag antibody (anti-GST mouse monoclonal antibody) (Bioman, catalog number: GST001M )
- Tris base (J.T.Baker, catalog number: 4109 )
- NaCl (Sigma-Aldrich, catalog number: S9888 )
- Glycerol (Sigma-Aldrich, catalog number: G5516 )
- Triton X-100 (Sigma-Aldrich, catalog number: T8787 )
- 2-mercaptoethanol (Sigma-Aldrich, catalog number: M6250 )
- Sodium dodecyl sulfate (SDS) (Sigma-Aldrich, catalog number: L3771 )
- Bromophenol blue (Sigma-Aldrich, catalog number: B0126 )
- Acrylamide/bis-acrylamide (37.5:1) (Bio-Rad Laboratories, catalog number: 1610158 )
- Ammonium persulfate (Sigma-Aldrich, catalog number: A3678 )
- TEMED (Sigma-Aldrich, catalog number: T9281 )
- Glycine (Sigma-Aldrich, catalog number: G8898 )
- Methanol (Sigma-Aldrich, catalog number: 322415 )
- Co-precipitation buffer (see Recipes)
- Wash buffer (see Recipes)
- 2x sample buffer (see Recipes)
- SDS-PAGE separating gel (see Recipes)
- SDS-PAGE stacking gel (see Recipes)
- PAGE running buffer (see Recipes)
- Western blot transfer buffer (see Recipes)
- pGEX-4T-1 DNA vector (GE Healthcare, catalog number: 28-9545-49 )
Equipment
- Thermal cycler (Biocompare, Biometra, catalog number: T3000 )
- Horizontal gel electrophoresis device (Bio-Rad Laboratories)
- 42 °C waterbath (FIRSTEK, catalog number: B101 )
- 37 °C Orbital Shaking Incubator (FIRSTEK, catalog number: S300R )
- Polyacrylamide gel electrophoresis system (Bio-Rad Laboratories, catalog number: 1658003 )
- Western blot transfer apparatus (Bio-Rad Laboratories, catalog number: 1703930 )
- Mixer for Eppendorf tube (ELMI North America, catalog number: RM-2L )
- Microcentrifuge (Eppendorf AG, catalog number: 05400002 )
Procedure

Figure 1. Illustration of the in vitro detection of the protein interaction procedure. A. Genes encoding prey proteins are cloned into the pEXP5-CT/TOPO vector using the TOPO cloning system. B. Expression of C-terminal His-tagged recombinant proteins by cell-free in vitro transcription/translation. C. Co-immunoprecipitation is performed to analyze interaction between the His-tagged prey and GST-tagged bait using a bait-specific antibody. An anti-His antibody was used for prey detection and an anti-GST antibody for examining bait precipitation.
- TOPO cloning for His-tagged protein construct (see Figure 1)
- Primer design: The forward primer of the candidate gene should include “CACC” upstream of the “ATG” start codon at the 5'-end of the primer: e.g., 5'-CACCATG(N)25-3'. The reverse primer is designed without stop codon for the His-tag fusion at the C-terminus of recombinant protein.
- Amplify the candidate gene. We recommend the use of the ExTaq polymerase, which generates DNA products with an adenine overhang at the 3'-end.
- Purify the DNA product via 0.8-1% agarose gel electrophoresis followed by gel-extraction. The QIAEX® II Gel Extraction Kit was recommended.
- Clone the extracted DNA product into the pEXP5-CT/TOPO® vector (Table 1).
Table 1. Components for the cloning reactionComponent Amount Purified DNA (0.05-1 μg) x μl Salt solution 0.5 μl pEXP5-CT/TOPO® vector 0.5 μl H2O To 3 μl - Incubate the reaction for 20 min at room temperature.
- Transform the plasmid into E. coli strain DH5α at 42 °C for 60 sec.
- Grow the transformed bacteria on LB agar containing 100 μg/ml ampicillin at 37 °C for 16 h.
- Screen the positive clones by colony polymerase chain reaction (PCR) with the T7 forward primer and gene-specific reverse primer.
- Grow the bacteria in LB liquid culture containing 100 μg/ml ampicillin at 37 °C with vigorous shaking (220 rpm) for 16 h.
- Extract plasmid DNA by PrestoTM Mini Plasmid Kit following the manufacturer's instructions.
- Confirm the selected clones by DNA sequencing.
- Primer design: The forward primer of the candidate gene should include “CACC” upstream of the “ATG” start codon at the 5'-end of the primer: e.g., 5'-CACCATG(N)25-3'. The reverse primer is designed without stop codon for the His-tag fusion at the C-terminus of recombinant protein.
- Expression of the recombinant protein via single-step in vitro transcription/translation (see Figure 1)
Note: The reaction volume is suggested as 100 μl (50 μl initial reaction and 50 μl feed buffer); however, it can be reduced to 25 μl (12.5 μl initial reaction and 12.5 μl feed buffer) for increasing the utility of the protein expression kit by 4 times in our screening protocol.- Prepare the initial in vitro transcription/translation reaction in a 1.7-ml tube using the ExpresswayTM Cell-Free E. coli Expression System (Table 2). Each recombinant protein was expressed individually in an independent reaction with its plasmid (1 μg).
Table 2. Components for initial in vitro transcription/translationComponent Amount E. coli slyD- extract 5 μl 2.5x IVPS buffer 5 μl 50 mM amino acids (without met) 0.3125 μl 75 mM methionine 0.25 μl T7 enzyme mix 0.25 μl pEXP5 plasmid DNA 1 μg H2O To 12.5 μl - Incubate the reaction for 30 min at 37 °C with vigorous shaking (220 rpm).
- Apply the feed buffer (Table 3) to the initial reaction and gently mix for several times by inverting the tube.
Table 3. Feed buffer componentsComponent Amount 2x IVPS feed buffer 6.25 μl 50 mM amino acid 0.3125 μl 75 mM methionine 0.25 μl H2O To 12.5 μl - Continue the reaction at 37 °C with shaking (220 rpm) for another 2.5 h (total 3 h).
Note: The yield increases alone with incubation time from 3-6 h, as indicated in manufacturer’s protocol; however, in our tested system, protein degradation can occur for at least two expressed proteins when the reaction time is extended to 5 h (Figure 2). Therefore, we recommend a total incubation time of 3 h. - Examine the expressed protein by Western analysis (Figure 2).
- Sample preparation: Save 5 μl expressed protein from 25 μl reaction and add equal volume of 2x sample buffer.
- Boil the protein sample at 100 °C for 10 min and cool down on ice for 5 min.
- Apply 5 μl boiled protein extract into SDS-PAGE analysis.
- Perform Western blotting with the anti-His (C-term)-HRP antibody.
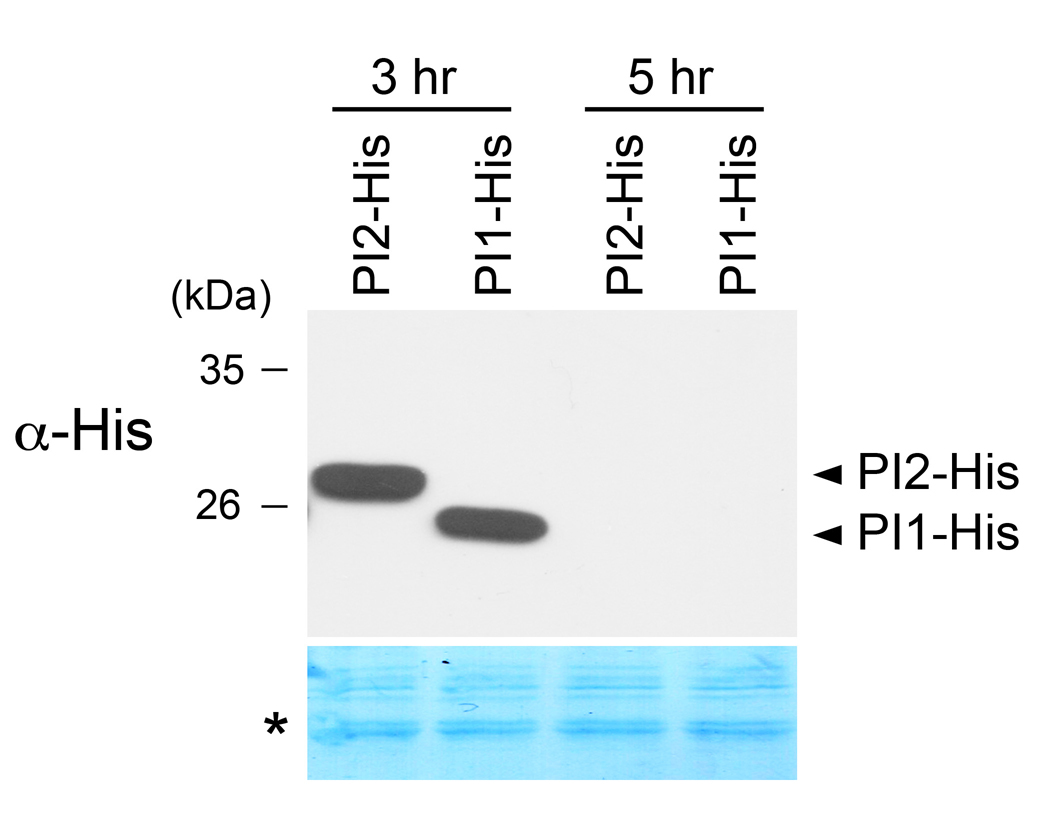
Figure 2. Example of detection of protein after in vitro transcription/translation. Tobacco proteins PI1 and PI2 (Lin et al., 2015) fused to a 6x His tag (PI1-His and PI2-His) were expressed via single-step in vitro transcription/translation with an incubation time of 3 h or 5 h. Coomassie blue staining (*) was used as loading control.
- Sample preparation: Save 5 μl expressed protein from 25 μl reaction and add equal volume of 2x sample buffer.
- Prepare the initial in vitro transcription/translation reaction in a 1.7-ml tube using the ExpresswayTM Cell-Free E. coli Expression System (Table 2). Each recombinant protein was expressed individually in an independent reaction with its plasmid (1 μg).
- Protein-protein interaction and co-immunoprecipitation (see Figure 1)
- Bait protein: A recombinant protein expressed in and purified from E. coli can be used for screening interacting partners (prey proteins). The desired tag [herein, a GST-tagged CP (GST-CP) protein] can be added when producing the bait protein. The CP gene was cloned into the pGEX-4T-1 vector by BamHI and XhoI digestion and ligation.
- Prey protein: His-tagged recombinant protein produced via in vitro transcription/translation.
Note: A negative control is required to avoid the false positive reactions. Replace the His-tagged interacting candidate product with a GFP-His in vitro transcription/translation product as a negative control. - Incubate 4 μl of crude reaction mixture from the in vitro transcription/translation reaction (prey) with 2 μg bait protein (herein, GST-CP recombinant protein) in 1 ml co-precipitation buffer for 1 h at room temperature with gentle rotation on an Eppendorf mixer with 5 rpm (mode 01).
- Balance the Protein G PLUS-agarose beads. Spin the agarose beads (30 μl suspension per co-IP reaction) at 2,000 x g for 1 min at room temperature, then remove the supernatant without disturbing the resin. Wash the beads with 0.5 ml H2O and repeat the spin procedure. Finally suspend the beads in 0.3 ml co-precipitation buffer.
- Add 10 μg of the anti-bait antibody (herein, anti-CP antibody) and balanced Protein G PLUS-agarose beads to the 1 h reaction mixture and incubate at room temperature on an Eppendorf mixer with 5 rpm (mode 01) for another 1 h.
- Centrifuge the reaction at 2,000 x g for 1 min at room temperature to sediment (pull down) the immunoprecipitant.
- Remove the supernatant and wash the immunoprecipitant twice with 0.5 ml wash buffer. Centrifuge the reaction at 2,000 x g for 1 min at room temperature for each wash and then remove the supernatant.
- Suspend the precipitant in 20 μl 2x sample buffer.
- Boil the protein sample at 100 °C for 10 min and then cool down on ice for 5 min.
- Perform Western analysis (apply 10-20 μl protein on the SDS-PAGE) for prey detection using the anti-His (C-term)-HRP antibody; and an anti-tag antibody (herein, anti-GST monoclonal antibody) was used for detection of the bait GST-CP. The official interacting data can be found in Lin et al. (2015).
- Bait protein: A recombinant protein expressed in and purified from E. coli can be used for screening interacting partners (prey proteins). The desired tag [herein, a GST-tagged CP (GST-CP) protein] can be added when producing the bait protein. The CP gene was cloned into the pGEX-4T-1 vector by BamHI and XhoI digestion and ligation.
Recipes
- Co-precipitation buffer
50 mM Tris (pH 7.5)
100 mM NaCl
0.2% glycerol
0.6% Triton X-100
0.5 mM mercaptoethanol - Wash buffer
50 mM Tris (pH 7.5)
100 mM NaCl
0.6% Triton X-100 - 2x sample buffer
50 mM Tris-HCl (pH 6.8)
2% SDS
10% glycerol
1% 2-mercaptoethanol
0.05% bromophenol blue - SDS-PAGE separating gel
375 mM Tris (pH 8.8)
10-15% acrylamide/bis-acrylamide (37.5:1)
0.1% SDS
0.05% ammonium persulfate
0.1% TEMED - SDS-PAGE stacking gel
375 mM Tris (pH 6.8)
4% acrylamide/bis-acrylamide (37.5:1)
0.1% SDS
0.05% ammonium persulfate
0.1% TEMED - PAGE running buffer
5 mM Tris (pH 8.3)
40 mM glycine
0.02% SDS - Western blot transfer buffer
25 mM Tris (pH 8.3)
192 mM glycine
10 % methanol
0.1% SDS
Acknowledgments
This work was supported by grants from the Ministry of Science and Technology, Taiwan (NSC-102-2313-B-002-068-MY3 and NSC-102-2313-B-002-066-B-MY3) to S.-S. Lin and (NSC-99-2313-B-002-043-MY3) to Y.-C. Chang.
References
- Katzen, F., Chang, G. and Kudlicki, W. (2005). The past, present and future of cell-free protein synthesis. Trends Biotechnol 23(3): 150-156.
- Lee, S. C. and Chang, Y. C. (2008). Performances and application of antisera produced by recombinant capsid proteins of Cymbidium mosaic virus and Odontoglossum ringspot virus. European Journal of Plant Pathology 122(2): 297-306.
- Lin, P. C., Hu, W. C., Lee, S. C., Chen, Y. L., Lee, C. Y., Chen, Y. R., Liu, L. Y., Chen, P. Y., Lin, S. S. and Chang, Y. C. (2015). Application of an integrated omics approach for identifying host proteins that interact with Odontoglossum ringspot virus capsid protein. Mol Plant Microbe Interact 28(6): 711-726.
- Shimizu, Y., Inoue, A., Tomari, Y., Suzuki, T., Yokogawa, T., Nishikawa, K. and Ueda, T. (2001). Cell-free translation reconstituted with purified components. Nat Biotechnol 19(8): 751-755.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Lin, P., Chang, Y. and Lin, S. (2016). An in vitro Transcription/translation System for Detection of Protein Interaction. Bio-protocol 6(9): e1800. DOI: 10.21769/BioProtoc.1800.
Category
Molecular Biology > Protein > Protein-protein interaction
Plant Science > Plant molecular biology > Protein
Microbiology > Microbe-host interactions > Virus
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link