- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Expression and Partial Purification of His-tagged Proteins in a Plant System
Published: Vol 5, Iss 17, Sep 5, 2015 DOI: 10.21769/BioProtoc.1572 Views: 14386
Reviewed by: Arsalan DaudiShyam SolankiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
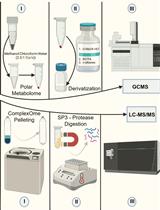
Streamlining Protein Fractional Synthesis Rates Using SP3 Beads and Stable Isotope Mass Spectrometry: A Case Study on the Plant Ribosome
Dione Gentry-Torfer [...] Federico Martinez-Seidel
May 5, 2024 2869 Views
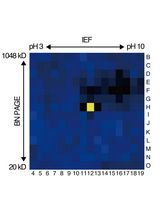

An Activity-Based Proteomics with Two-Dimensional Polyacrylamide Gel Electrophoresis (2D-PAGE) for Identifying Target Proteases in Arabidopsis Apoplastic Fluid
Sayaka Matsui and Yoshikatsu Matsubayashi
Mar 5, 2025 1989 Views
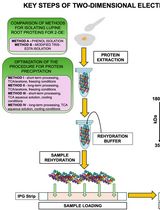
Advancing 2-DE Techniques: High-Efficiency Protein Extraction From Lupine Roots
Sebastian Burchardt [...] Emilia Wilmowicz
Oct 5, 2025 1772 Views
Abstract
Plant protein expression can be a challenging enterprise in any biochemical or molecular biology research project. Several heterologous systems like bacteria, yeast, insect cells and cell free systems have been used to produce plant proteins for in vitro experiments and structural characterization. However, due to particularities of plant proteins, for example the specific type and abundance of post-translational modifications (e.g. glycosylation), a plant system to express plant proteins is extremely desirable. The use of Nicotiana benthamiana (N. benthamiana) plants for protein expression has proven to be quick and reliable. To illustrate the robustness and rapidity of this system, recent efforts to produce the first protein based drug against the Ebola virus was conducted in N. benthamiana protein expression systems (Choi et al., 2015).
This protocol describes a simple system for the expression and enrichment (affinity purification) of plant apoplastic proteins in N. benthamiana leaves, which was successfully used in the characterization of the Arabidopsis thaliana pectin acetylesterases, PAE8 and PAE9 (de Souza et al., 2014).
Materials and Reagents
- Nicotiana benthamiana seeds
- Agrobacterium strain GV3101 (obtained from the Lab of Dr. Markus Pauly at UC Berkeley’s Plant and Microbial Biology departmenty)
- PRO-MIX® HP MYCORRHIZAE™ soil mix (Promix, catalog number: 20381RG )
- Miracle-Gro® Water Soluble All Purpose Plant Food (Scotts)
- pART27 expression vector (Gleave, 1992)
- Tryptone (MP Biomedicals, catalog number: 1010817 )
- Yeast extract (U.S. Biotech Sources, catalog number: Y01PD-500 )
- NaCl (Thermo Fisher Scientific, catalog number: S271-3 )
- 4'-Hydroxy-3', 5'-dimethoxyacetophenone (acetosyringone) (150 mM in DMSO) (Sigma-Aldrich, catalog number: D134406-1G )
- Aluminum foil (Reynolds wrap 76.2 m x 304 mm) (Reynolds Consumer Products Inc.)
- Liquid nitrogen
- 100x Halt™ Protease Inhibitor Cocktail (Thermo Fisher Scientific, catalog number: 78429 )
- β-mercaptoethanol (Sigma-Aldrich, catalog number: M6250 )
- Bradford reagent (Bio-Rad Protein Assay Dye reagent concentrate) (Bio-Rad Laboratories, AbD Serotec®, catalog number: 500-0006 )
- Small columns for Ni-NTA bead wash and elution (any that will fit a 1.5 ml Eppendorf tube, e.g. miniprep column)
- 500 µl Vivaspin Column MWCO of 5,000 (Sartorius stedim biotech, catalog number: VS0111 )
- Ni-NTA beads (QIAGEN, catalog number: 1018240 )
- 96 well plates for Bradford assay (Thermo Fisher Scientific, catalog number: 80040LE0910 )
- 2-(N-morpholino)ethanesulfonic acid (MES) (Sigma-Aldrich, catalog number: M2933 )
- MgCl2 (Thermo Fisher Scientific, catalog number: M33-500 )
- Sodium phosphate monobasic (Thermo Fisher Scientific, catalog number: S369-500 )
- Sodium phosphate dibasic (Thermo Fisher Scientific, catalog number: S374-500 )
- Imidazole (Sigma-Aldrich, catalog number: I-2399 )
- Lennox LB media (see Recipes)
- Infiltration buffer (see Recipes)
- Sodium phosphate buffer (see Recipes)
- Extraction buffer (see Recipes)
- Wash buffer (see Recipes)
- Elution buffer (see Recipes)
Equipment
- Plant pots (400 ml volume or similar, Gage Durapot) (Merrill's Packaging, catalog number: 03GA-0350S )
- Plant growth trays (T.O. Plastics, catalog number: 710245C )
- Tall covers that won’t touch the leaves (Acrodome) (Drader Manufactoring Industries Ltd., catalog number: 69973 )
- Mortar and pestle
- 500 ml culture flasks
- Immersion recipient for dipping N. benthamiana plants (e.g. 250 ml beakers)
- Metal beads (2.38 mm) (Tool Supply, catalog number: 6230 )
- Ball mill (Mixer Mill MM 400 ) (RETSCH, catalog number: MM 400)
- Plant growth chambers capable of sustaining 26 °C under long-day conditions (16 h light/8 h dark) with 170-190 µmol m-2 s-1 light intensity.
- Spray bottle with water.
- Incubator/orbital shaker, capable of 30 °C at 230 rpm incubation for 500 ml culture flaks.
- Spectrophotometer capable of OD600 measurements.
- Large centrifuge capable of spinning down 250 ml or greater volumes at 5,000 x g for 10 min
- Desiccator or vacuum chamber
- Vacuum pump (Savant Systems LLC, catalog number: Gel Pump-GP110 )
- -80 °C Freezer
- Rotating agitator/circular shaker
- Table top centrifuges (500-20,800 x g, 4 °C)
- Spectrophotometer capable of 96 well plate measurements at 595 nm
Procedure
- Plant growth conditions
The preparation of N. benthamiana plants is a key step in obtaining satisfactory protein expression; plants should be as vigorous as possible to help in their recovery after infiltration and consequent protein production. In this protocol ~6 week old N. benthamiana plants are used for Agrobacterium tumefaciens infiltration.- Grow plants at 26 °C under long-day conditions (16 h light/8 h dark) with 170-190 µmol m-2 s-1 light intensity and optimal humidity of 70%.
- Sow seeds in water-soaked soil mix (Promix HP mycorrhizae; 400 ml pots; Video 1) and grow for 2 weeks before transplanting to final destination pots (400 ml).Video 1. Detailed description of the procedures for sowing seeds (0 sec), transplanting seedlings (50 sec) and performing the vacuum infiltration (1 min 45 sec)
- Transplant seedlings carefully to preserve as much of the root as possible (Video 1). After transplantation fertilize once with Miracle Grow All-purpose Plant Food (Scotts) according to manufacturer’s recommendation and water as needed. These pots should be grown for another 4 weeks until they are ready for transformation. During the first week of growth young seedlings are kept covered. Multiple fertilization rounds at this stage can cause excessive growth which will produce plants that are too big for infiltration. It can also increase overlap between pots in the trays which will generate problems in the manipulation of the material. Starting at the fifth week it is particularly important to monitor plant development in order to plan for the infiltration procedure. Ideally plants will have 4-5 fully expanded leaves ~7 cm in diameter at 5-6 weeks for infiltration (Video 1).
- When ready for infiltration, if possible, transport plants a few hours in advance to the work site and spray the leaves with abundant water, keeping the plants covered to prevent leaf wilting.
- Grow plants at 26 °C under long-day conditions (16 h light/8 h dark) with 170-190 µmol m-2 s-1 light intensity and optimal humidity of 70%.
- Constructs, Agrobacterium cultures and vacuum infiltration
The proteins of interest described in this protocol were tagged with 6 histidines at their C- terminus. This construct was cloned into the pART27 binary vector under the control of the 35S promoter (Gleave, 1992). Vectors were transformed into the Agrobacterium strain GV3101 for N. benthamiana transient transformation.- Agrobacterium preparation
- Agrobacterium cultures (construct of interest, empty vector and P19) should be started from fresh colonies or glycerol stocks in LB media supplemented with appropriate antibiotics and cultured at 30 °C and 230 rpm. Volumes of at least 200 ml should be used and cultured until reaching OD600 of 1-2 (~ 48 h).
- When cultures present increased turbidity, measure OD600 and calculate the necessary dilution so that a final volume of at least 250 ml of OD600 0.7 will be obtained in infiltration buffer for each construct. If necessary allow cultures to grow longer to reach the required amount of cells. When co-infiltrating with P19, a suppressor of gene silencing (Voinnet et al., 2003), the final calculated OD600 of each individual construct should be at least 0.7 (total OD600 will be the sum of both individual ODs). Ideally the final OD600 ratio between construct of interest and P19 should be of 0.7:1.
- Spin down cells at 22 °C and 5,000 x g for 10 min.
- Discard LB media supernatant by decanting, eliminating as much of the supernatant as possible.
- Resuspend cell pellet using infiltration buffer.
- Add 500 µl of acetosyringone (4'-Hydroxy-3',5'-dimethoxyacetophenone; 150 mM in DMSO) for every 500 ml of suspended cells in infiltration buffer.
- Incubate bacteria for 3-4 h at room temperature before infiltration.
- Agrobacterium cultures (construct of interest, empty vector and P19) should be started from fresh colonies or glycerol stocks in LB media supplemented with appropriate antibiotics and cultured at 30 °C and 230 rpm. Volumes of at least 200 ml should be used and cultured until reaching OD600 of 1-2 (~ 48 h).
- Vacuum infiltration procedure (see Video 1 for details on this section)
- Cover the top part of the N. benthamiana pots with aluminum foil. This is to prevent excessive soil loss during the procedure and to keep infiltration buffer as clean as possible, for multiple infiltrations. A square sheet of foil, larger than the plant pot, slit from one of the sides to the center makes an easy to use, disposable cover.
- Place the infiltration buffer with cells in a beaker or container that will allow full immersion of the N. benthamiana leaves when dipped.
- Dip the plant aerial part into the cell suspension making sure all the leaves are immersed in the infiltration buffer. Be careful not to break petioles or damage the plant.
- Place beaker with plant in a vacuum chamber or desiccator capable of tolerating vacuum pressures. Depending on the size of the container available, multiple plants can be infiltrated simultaneously.
- Use a vacuum pump strong enough to produce a vacuum that will pull most of the gas out of the leaves (at least 90 kPa). When vacuum is applied gas bubbles can be observed coming out of the leaves. For every infiltration apply vacuum for 3 min, releasing the vacuum gently, and repeat operation for a total of three times.
- Remove the plants from the infiltration buffer. Leaves that were successfully infiltrated should have a translucent or water-soaked appearance. Remove any leaves that were not successfully infiltrated. Leaves that were not fully submerged usually don’t infiltrate very well.
- Place plants in a covered tray with plenty of water at the bottom. Return plants to growth chambers.
- Remove covers from trays 24-48 h after infiltration, according to how well plants recovered.
- Cover the top part of the N. benthamiana pots with aluminum foil. This is to prevent excessive soil loss during the procedure and to keep infiltration buffer as clean as possible, for multiple infiltrations. A square sheet of foil, larger than the plant pot, slit from one of the sides to the center makes an easy to use, disposable cover.
- Agrobacterium preparation
- Protein extraction and partial purification
The partial protein purification described here is based on the affinity of the 6x histidine C-terminal tag to nickel-containing resins. The affinity is based on the charges of the two mentioned groups. In plants, this approach has proven to be useful; however it is difficult to obtain pure preparations of the protein of interest using this technique alone. The method described here is able to enrich for the protein of interest but does not produce completely pure fractions. For this reason an “empty vector” protein extraction should be done in parallel to ensure that every downstream experiment using the partially purified protein has a proper negative control.- Protein Extraction
- In 3-5 days harvest N. benthamiana leaves for protein extraction by flash-freezing them in liquid nitrogen. Depending on the protein being expressed, shorter incubation periods after infiltration could be attempted.
- Pre-grind leaves (empty vector control and construct of interest co-transformed with P19) in a mortar and pestle with liquid nitrogen. This powder can be stored at -80 °C. Pre-grinding can facilitate further processing for protein extraction.
- Add 3 metal beads (2.38 mm) to a 2 ml tube.
- Without allowing pre-ground leaves to thaw (work on dry ice and in a cold room if available), place ~ 1 ml of ground material into a 2 ml tube. This volume can be scaled up proportionally if necessary.
- Using a bead beater (Retsch ball mill) grind material frozen in liquid nitrogen for 2.5 min, at 25 Hz.
- To 1 ml of extraction buffer add 10 μl of 100x Halt™ Protease Inhibitor Cocktail (0.99x) and 0.14 μl of β-mercaptoethanol (1.98 mM) (final concentrations of 49.5 mM sodium phosphate, 0.99 M NaCl and 9.9 mM imidazole).
- Incubate at 4 °C with gentle agitation for 1 h. Use rotating agitator, or similar device that allows the buffer to move in the tube and completely mix the sample.
- After one hour remove metal beads with a magnet, always keeping samples on ice.
- Pellet plant debris by centrifugation at 4 °C and 20,800 x g for 10 min.
- Collect ~ 1.1 ml of supernatant into a new 2 ml tube.
- Centrifuge again at 4 °C and 20,800 x g for 10 min, to pellet any carryover leaf debris.
- Collect 1 ml of supernatant and transfer to a fresh tube. Depending on the volume being processed it can be a 2 ml tube or larger. Always keep samples on ice.
- Measure protein content of the collected supernatant using Bradford assay (Bio-Rad Protein Assay Dye reagent concentrate). It is recommended to use a 96 well plate format with a bovine serum albumin standard curve. The protein measurement here is important to normalize the amount of protein loading onto the affinity beads, 2-3 mg of total protein/ml should be expected. At this stage protein crude extracts can be tested for the presence of the protein of interest using immunoblotting techniques (westerns or dot blots). This is recommended when setting up conditions for protein expression.
- In 3-5 days harvest N. benthamiana leaves for protein extraction by flash-freezing them in liquid nitrogen. Depending on the protein being expressed, shorter incubation periods after infiltration could be attempted.
- Nickel NTA bead preparation
- Re-suspend Ni-NTA beads and collect 100 µl into a 1.5 ml tube (50 µl of resin, resin usually compose half the volume of the product).
- Spin down at 500 x g for 1 min and remove supernatant.
- Wash 3 times with 500 µl extraction buffer, using same centrifugation conditions described above. Final suspension is done in 500 µl extraction buffer.
- After washes add 10 µl of resin (110 µl of suspension) to every 1 ml of crude protein supernatant.
- Re-suspend Ni-NTA beads and collect 100 µl into a 1.5 ml tube (50 µl of resin, resin usually compose half the volume of the product).
- Partial protein purification
- Incubate for 1 h at 4 °C under gentle agitation to allow tagged proteins to bind to the Ni-NTA beads. During this time undesired precipitation of proteins may occur, in this case an alternative procedure is to bind the tagged proteins to the resin by running the supernatant multiple times through a column containing the beads instead of the batch procedure described.
- Spin down to collect beads at 4 °C and 500 x g for 1 min. The beads will form a pellet on the bottom of the tube.
- Collect 250 µl of beads and supernatant and place into a small spin column for table top centrifuge with a 2 ml collection tube. Any column that will fit in an Eppendorf-like tube can be used here, since its purpose is just to serve as a support for the Ni-NTA beads. Column material used shouldn’t bind proteins. This step greatly facilitates the procedure and speeds up the washes and elution steps.
- Spin down at 4 °C and 500 x g for 1 min.
- Wash beads 5 times with 250 µl extraction buffer + protease inhibitors and β-mercaptoethanol (see step C1f). The wash consists of adding the referred volume and discarding the flow through after centrifugation (4 °C and 500 x g for 1 min). Alternatively flow through of the washes can be kept to monitor the presence of the protein of interest using immunoblotting techniques.
- Wash 4 times with 200 µl washing buffer
- Elute 6 times in 50 µl of elution Buffer into a fresh tube.
- Place elution fraction (~ 300 µl) in a 500 µl Vivaspin Column (MWCO of 5,000) for buffer exchange. In this case buffer exchange was necessary due to incompatibility of imidazole and downstream assays, this procedure might not always be necessary.
- Spin down at 4 °C and 20,800 x g for 5 min and add 300 µl of 50 mM ammonium formate (pH 4.5) to ~100 µl of sample (4 times dilution, each time). Repeat the procedure for a total of 4 times resulting in the recovery of 200 µl of material containing less than 1 mM imidazole. The protein concentration yield is approximately 0.025 mg/ml for every 1 ml of plant tissue starting material.
- Incubate for 1 h at 4 °C under gentle agitation to allow tagged proteins to bind to the Ni-NTA beads. During this time undesired precipitation of proteins may occur, in this case an alternative procedure is to bind the tagged proteins to the resin by running the supernatant multiple times through a column containing the beads instead of the batch procedure described.
- Protein Extraction
Recipes
- Lennox LB media
Dissolve in 1 L water, 10 g of tryptone, 5 g yeast extract and 5 g NaCl - Infiltration buffer
10 mM MES
10 mM MgCl2
pH 5.6 - Sodium phosphate buffer (pH 8)
93.2 ml of 1 M sodium phosphate dibasic
6.8 ml of 1 M sodium phosphate monobasic
Add double distilled water to 1 L - Extraction buffer
1 M NaCl
50 mM sodium phosphate (pH 8)
10 mM imidazole - Wash buffer
300 mM NaCl
50 mM sodium phosphate (pH 8)
20 mM imidazole - Elution buffer
300 mM NaCl
50 mM sodium phosphate (pH 8)
150 mM imidazole
Acknowledgments
This protocol is an expansion of that described in de Souza et al. (2014). I would like to thank Marta L. Bjornson for aiding in the revision of the manuscript and Dr. Katayoon Dehesh for laboratory logistical support in the revision process. This work was supported by the Energy Biosciences Institute at UC Berkeley.
References
- Choi, W. Y., Hong, K. J., Hong, J. E. and Lee, W. J. (2015). Progress of vaccine and drug development for Ebola preparedness. Clin Exp Vaccine Res 4(1): 11-16.
- de Souza, A., Hull, P. A., Gille, S. and Pauly, M. (2014). Identification and functional characterization of the distinct plant pectin esterases PAE8 and PAE9 and their deletion mutants. Planta 240(5): 1123-1138.
- Gleave, A. P. (1992). A versatile binary vector system with a T-DNA organisational structure conducive to efficient integration of cloned DNA into the plant genome. Plant Mol Biol 20(6): 1203-1207.
- Voinnet, O., Rivas, S., Mestre, P. and Baulcombe, D. (2003). An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J 33(5): 949-956.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Souza, A. D. (2015). Expression and Partial Purification of His-tagged Proteins in a Plant System. Bio-protocol 5(17): e1572. DOI: 10.21769/BioProtoc.1572.
Category
Plant Science > Plant biochemistry > Protein > Isolation and purification
Biochemistry > Protein > Expression
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










