- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Micro-scale NMR Experiments for Monitoring the Optimization of Membrane Protein Solutions for Structural Biology
Published: Vol 5, Iss 14, Jul 20, 2015 DOI: 10.21769/BioProtoc.1539 Views: 8245
Reviewed by: Arsalan DaudiMarc-Antoine SaniAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
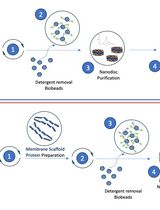
Optimizing Transmembrane Protein Assemblies in Nanodiscs for Structural Studies: A Comprehensive Manual
Fernando Vilela [...] Dorit Hanein
Nov 5, 2024 3097 Views
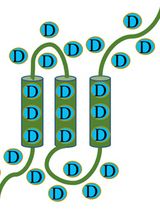
Protein Structural Characterization Using Electron Transfer Dissociation and Hydrogen Exchange-Mass Spectrometry
Rupam Bhattacharjee and Jayant B. Udgaonkar
Jun 20, 2025 1782 Views
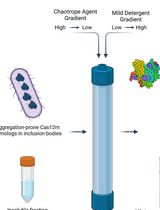
On-Column Dual-Gradient Refolding for Efficient Recovery of Insoluble Affinity-Tagged Recombinant Proteins
Anna Vlaskina [...] Maxim Patrushev
Feb 5, 2026 66 Views
Abstract
Reconstitution of integral membrane proteins (IMP) in aqueous solutions of detergent micelles has been extensively used in structural biology, using either X-ray crystallography or NMR in solution. Further progress could be achieved by establishing a rational basis for the selection of detergent and buffer conditions, since the stringent bottleneck that slows down the structural biology of IMPs is the preparation of diffracting crystals or concentrated solutions of stable isotope labeled IMPs. Here, we describe procedures to monitor the quality of aqueous solutions of [2H, 15N]-labeled IMPs reconstituted in detergent micelles. This approach has been developed for studies of β-barrel IMPs, where it was successfully applied for numerous NMR structure determinations, and it has also been adapted for use with α-helical IMPs, in particular GPCRs, in guiding crystallization trials and optimizing samples for NMR studies (Horst et al., 2013). 2D [15N, 1H]-correlation maps are used as “fingerprints” to assess the foldedness of the IMP in solution. For promising samples, these “inexpensive” data are then supplemented with measurements of the translational and rotational diffusion coefficients, which give information on the shape and size of the IMP/detergent mixed micelles. Using microcoil equipment for these NMR experiments enables data collection with only micrograms of protein and detergent. This makes serial screens of variable solution conditions viable, enabling the optimization of parameters such as the detergent concentration, sample temperature, pH and the composition of the buffer.
Keywords: Micro-scale NMRMaterials and Reagents
Studies of IMPs
- Phosphocholine-detergents (Avanti Polar Lipids)
- Tris Base (Thermo Fisher Scientific, catalog number: BP1521 )
- HCl (Thermo Fisher Scientific, catalog number: A144212 )
- Urea (Thermo Fisher Scientific, catalog number: BP169212 )
- Ethylenediaminetetraacetic acid (EDTA) (Thermo Fisher Scientific, catalog number: BP1201 )
- L-Arginine (L-Arg) (Sigma-Aldrich, catalog number: A5131100G )
- Phosphate buffer (Sigma-Aldrich, catalog number: P7994 )
- Sodium Azide (NaN3) (Sigma-Aldrich, catalog number: S803225G )
- NaCl (Thermo Fisher Scientific, catalog number: BP358212 )
- Stock solutions of unfolded protein (see Recipes)
- Refolding buffer (see Recipes)
- NMR buffer (see Recipes)
Equipment
NMR data collection
- NMR experiment set-ups used in this protocol are either part of the Bruker standard pulse sequence library or are described in the Appendix. These experiments were implemented on a Bruker DRX spectrometer equipped with microprobes (Bruker Corporation,model:1 mm TXI, 1.7 mm TXI )
- The following experiments were used to monitor the quality of aqueous solutions of [2H, 15N]-labeled IMPs: 2D [15N, 1H]-TROSY experiments (Pervushin et al., 1997), 1H-TRO-STE (Horst et al., 2011) and TRACT (Lee et al., 2006).
Procedure
- NMR sample preparation
- Add 100 µl of freshly prepared stock solution of unfolded protein to 600 μl refolding buffer and stir overnight at 4 °C.
- Exchange with NMR buffer by repeated dilution/concentration cycles. ‘‘Upconcentration’’ of the detergent during these dilution/concentration cycles was taken into account when adjusting the detergent concentration in the NMR sample. For details on how to adjust these cycles, see Stanczak et al. (2009). For example, 10 mM 30-Fos in the NMR buffer results in approximately 160 mM 30-Fos in the NMR sample (Stanczak et al., 2012).
- Added 5 μl of D2O and 1 μl of a 100 mM solution of 4, 4-dimethyl-4-silapentane-1-sulfonic acid (DSS) as an internal reference for the 1H chemical shifts to 45 μl of the protein solution.
- Add 100 µl of freshly prepared stock solution of unfolded protein to 600 μl refolding buffer and stir overnight at 4 °C.
- Evaluation of different sample conditions based on 2D [15N, 1H]-TROSY experiments. The following criteria were used (see Zhang et al., 2008; Stanczak et al., 2012 for examples):
- Completeness of NMR observation of the IMP by comparison of the number of observed backbone 15N–1H correlation peaks with the number expected from the amino acid sequence.
- High average peak intensity and uniform distribution of peak intensities.
- Analysis of the peak line shapes to support the interpretation of the peak intensity measurements.
- Completeness of NMR observation of the IMP by comparison of the number of observed backbone 15N–1H correlation peaks with the number expected from the amino acid sequence.
- Evaluation of the hydrodynamic radius of IMP/detergent mixed micelles, using 1H-TRO-STE (Horst et al., 2011) and TRACT (Lee et al., 2006) experiments.
- Determine the translational diffusion coefficient, Dt, using the 1H-TRO-STE experiment.
- Determine the rotational diffusion coefficient, Dr, using the TRACT experiment.
- Calculate the hydrodynamic radius, Rh, using the formula Rh = (3Dt/4Dr)1/2.
- Optimize sample conditions towards small Rh values, which favors the recording of high-quality NMR data.
- Determine the translational diffusion coefficient, Dt, using the 1H-TRO-STE experiment.
Data analysis
Processing and analysis of NMR datasets:
Process all NMR experiments with the Bruker standard software Topspin. For all experiments, the data matrices are multiplied with an exponential window function in the 1H-dimension, and for 2D [15N, 1H]-TROSY a 75°-shifted sine bell window (De Marco and Wüthrich, 1976) is applied in the 15N-dimension The 2D [15N, 1H]-TROSY data sets were analyzed using the XEASY module (Bartels et al., 1995) of the CARA release 1.5.5 (www.cara.nmr.ch). The 1H-TRO-STE datasets are analyzed using the Bruker T1/T2-software package, as described in the Topspin DOSY application manual, chapter 3.2. The TROSY and anti-TROSY components of the TRACT data set are first separated using the Bruker standard AU program split, and then individually integrated using the Bruker integration module. The integrals were fitted to a mono-exponential decay, using the program XMGRACE (http://plasma-gate.weizmann.ac.il).
Notes
Optimization of NMR acquisition parameters for the 2D [15N, 1H]-TROSY, 1H-TRO-STE and TRACT experiments:
- For all experiments
- Determine the lengths of high power radio-frequency pulses as described in the user manual for the spectrometer. Typical pulse lengths are 8 μs and 35 μs for 1H and 15N, respectively.
- Set the carrier frequency to the water line by minimizing the residual water signal in the Bruker standard pulse sequence zggppr.
- Optimize the water flip-back pulses sp2 and sp3, using the experiment shown in Listing 1 of the Appendix to interactively minimize the residual water signal in the topspin gs acquisition mode.
- Determine the lengths of high power radio-frequency pulses as described in the user manual for the spectrometer. Typical pulse lengths are 8 μs and 35 μs for 1H and 15N, respectively.
- 2D [15N, 1H]-TROSY and TRACT experiments (listings 2 and 4 of the Appendix)
- Adjust the WATERGATE soft pulse sp1, using the Bruker standard pulse sequence zggpwg by minimizing the residual water signal interactively in the topspin gs acquisition mode.
- Adjust the [15N, 1H]-INEPT transfer delay, d2, to maximize the signal from the protein amide moieties in the first increment of the TRACT experiment. Typical values for d2 are between 2.0 and 2.5 ms.
- Adjust the number of complex points and the maximum evolution time, t1max, for the 15N-dimension in the 2D [15N, 1H]-TROSY experiment. Typical values for IMP’s reconstituted in detergent micelles are 100 points and 35 ms, respectively. Use the same t1max value for the TRACT experiment.
- Adjust the WATERGATE soft pulse sp1, using the Bruker standard pulse sequence zggpwg by minimizing the residual water signal interactively in the topspin gs acquisition mode.
- 1H-TRO-STE experiment (listing 3 in the Appendix)
- Calibrate the pulsed-field gradient strengths with the residual 1H signal of 99.9% D2O, using the Bruker standard pulse sequence ledgp2s1d ( Dt = (1.902 ± 0.09) × 10‒9 m2s‒1 for HDO at 25 °C).
- Optimize the [15N,1H]-CRINEPT transfer delay, d2, by maximizing the signal from the protein amide moieties in the first increment of the spectrum.
- Optimize the diffusion delay and the gradient duration, d20 and p30, respectively, using the standard procedures described in the Topspin DOSY application manual, Chapter 2.1.
- Calibrate the pulsed-field gradient strengths with the residual 1H signal of 99.9% D2O, using the Bruker standard pulse sequence ledgp2s1d ( Dt = (1.902 ± 0.09) × 10‒9 m2s‒1 for HDO at 25 °C).
Recipes
- Stock solutions of unfolded protein
[2H, 15N]-labeled protein (12 mg/ml) in 20 mM Tris-HCl at pH 8.0 and 6 M urea
- Refolding buffer
20 mM Tris-HCl at pH 7.5
5 mM EDTA
600 mM L-Arg
45 mM detergent
- NMR buffer
5 mM phosphate buffer at pH 6.8
10 mM NaCl, 0.3 % (v/w) NaN3
10 mM detergent
Acknowledgments
This work was adapted from previously published studies on the E. coli outer membrane protein X (OmpX) (Stanczak et al., 2009), and was used as a platform for the structure determination of E. coli OmpW (Horst et al., 2014). The procedures described in this protocol were also used to characterize E. coli OmpA in lipid bilayer nanodiscs and detergent micelles (Susac et al., 2014). This work was supported by the Roadmap initiative grant P50 GM073197 for technology development. Kurt Wüthrich is the Cecil H. and Ida M. Green Professor of Structural Biology at the Scripps Research Institute.
References
- Bartels, C., Xia, T. H., Billeter, M., Guntert, P. and Wuthrich, K. (1995). The program XEASY for computer-supported NMR spectral analysis of biological macromolecules. J Biomol NMR 6(1): 1-10.
- De Marco, A. and Wüthrich, K. (1976). Digital filtering with a sinusoidal window function: an alternative technique for resolution enhancement in FT NMR. J Magn Reson 24(2): 201-204.
- Horst, R., Horwich, A. L. and Wuthrich, K. (2011). Translational diffusion of macromolecular assemblies measured using transverse-relaxation-optimized pulsed field gradient NMR. J Am Chem Soc 133(41): 16354-16357.
- Horst, R., Stanczak, P. and Wuthrich, K. (2014). NMR polypeptide backbone conformation of the E. coli outer membrane protein W. Structure 22(8): 1204-1209.
- Horst, R., Stanczak, P., Stevens, R. C. and Wuthrich, K. (2013). beta2-Adrenergic receptor solutions for structural biology analyzed with microscale NMR diffusion measurements. Angew Chem Int Ed Engl 52(1): 331-335.
- Lee, D., Hilty, C., Wider, G. and Wüthrich, K. (2006). Effective rotational correlation times of proteins from NMR relaxation interference. J Magn Reson 178(1): 72-76.
- Pervushin, K., Riek, R., Wider, G. and Wuthrich, K. (1997). Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci U S A 94(23): 12366-12371.
- Stanczak, P., Horst, R., Serrano, P. and Wuthrich, K. (2009). NMR characterization of membrane protein-detergent micelle solutions by use of microcoil equipment. J Am Chem Soc 131(51): 18450-18456.
- Stanczak, P., Zhang, Q., Horst, R., Serrano, P. and Wuthrich, K. (2012). Micro-coil NMR to monitor optimization of the reconstitution conditions for the integral membrane protein OmpW in detergent micelles. J Biomol NMR 54(2): 129-133.
- Susac, L., Horst, R. and Wuthrich, K. (2014). Solution-NMR characterization of outer-membrane protein A from E. coli in lipid bilayer nanodiscs and detergent micelles. Chembiochem 15(7): 995-1000.
- Zhang, Q., Horst, R., Geralt, M., Ma, X., Hong, W. X., Finn, M. G., Stevens, R. C. and Wuthrich, K. (2008). Microscale NMR screening of new detergents for membrane protein structural biology. J Am Chem Soc 130(23): 7357-7363.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Horst, R. and Wüthrich, K. (2015). Micro-scale NMR Experiments for Monitoring the Optimization of Membrane Protein Solutions for Structural Biology. Bio-protocol 5(14): e1539. DOI: 10.21769/BioProtoc.1539.
Category
Biochemistry > Protein > Structure
Biochemistry > Protein > Labeling
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









