- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
An Evaluation of Cellulose Degradation Affected by Dutch Elm Disease
Published: Vol 5, Iss 14, Jul 20, 2015 DOI: 10.21769/BioProtoc.1535 Views: 9169
Reviewed by: Arsalan DaudiRenate WeizbauerAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Enzymatic Starch Quantification in Developing Flower Primordia of Sweet Cherry
Nestor Santolaria [...] Afif Hedhly
Apr 5, 2025 1896 Views
New Approach to Detect and Isolate Rhamnogalacturonan-II in Arabidopsis thaliana Seed Mucilage
Dayan Sanhueza and Susana Saez-Aguayo
Sep 5, 2025 1248 Views
Detailed Method for the Purification of Rhamnogalacturonan-I (RG-I) in Arabidopsis thaliana
Liang Zhang [...] Breeanna R. Urbanowicz
Feb 5, 2026 144 Views
Abstract
The pathogenic fungus Ophiostoma novo-ulmi spreads within the secondary xylem vessels of infected elm trees, causing the formation of vessel plugs due to tyloses and gels, which ultimately result in Dutch elm disease. Foliage discoloration, wilting and falling from the tree are typical external leaf symptoms of the disease followed by the subsequent death of sensitive trees. Cellulolytic enzymes produced by the fungus are responsible for the degradation of medium molecular weight macromolecules of cellulose, resulting in the occurrence of secondary cell wall ruptures and cracks in the vessels but rarely in the fibers (Ďurkovič et al., 2014). The goal of this procedure is to evaluate the extent of cellulose degradation by a highly aggressive strain of O. novo-ulmi ssp. americana × novo-ulmi. Size-exclusion chromatography (SEC) compares molecular weight distributions of cellulose between the infected and the non-infected elm trees, and reveals changes in the macromolecular traits of cellulose, including molecular weights, degree of polymerization, and polydispersity index. 13C magic angle spinning nuclear magnetic resonance (13C MAS NMR) spectra help to identify and also to quantify the loss of both crystalline and non-crystalline cellulose regions due to degradation. The procedure described herein can also be easily used for other woody plants infected with various cellulose-degrading fungi.
Keywords: Ophiostoma novo-ulmiMaterials and Reagents
- Absolute ethanol (99.5%) (EMD Millipore, catalog number: 107017 )
- Toluene (99.5%) (Merck KGaA, catalog number: 107019 )
- Acetylacetone (99%) (Merck KGaA, catalog number: 109600 )
- Dioxane (99.5%) (Merck KGaA, catalog number: 109671 )
- Fuming hydrochloric acid (37%) (Merck KGaA, catalog number: 101834 )
- Methanol (99.8%) Merck KGaA, catalog number: 107018 )
- Ultrapure water
Note: It is produced by Millipore Simplicity® 185 (UV) ultrapure water purification system (EMD Millipore). - Pyridine (99.5%) (Merck KGaA, catalog number: 109728 )
- Phenyl isocyanate (99%) (Merck KGaA, catalog number: 107255 )
- Tetrahydrofuran (99.8%) (Merck KGaA, catalog number: 109731 )
Equipment
- Chainsaw, bandsaw, abrasive belt machine and woodworking lathe
- Analytical balance (accurate to 1 mg or 0.1 mg)
- Desiccator and oven for drying of samples (set to 50 ± 3 °C, 70 ± 3 °C, and to 105 ± 3 °C)
- Polymix (Kinematica, model: PX-MFC 90D )
- Analysette 3 vibratory sieve shaker (Fritsch)
- Soxhlet extraction apparatus (Sigma–Aldrich, catalog number: 64825 )
- Boiling flasks (50 ml) (Sigma–Aldrich, catalog number: Z418773 )
- Water bath (Harry Gestigkeit Gmbh, model: W 16 )
- Fritted-glass filtering crucible of medium porosity (16–40 µm) (VWR International, catalog number: 511-2403 )
- Dropping flasks (50 ml) (Smith Scientific Limited, catalog number: 8029/50 )
- High performance liquid chromatography system (degasser, pump, autosampler, heater and diode-array ) (Agilent Technologies, model: 1200 series )
- Captiva Premium Syringe Filter (0.45 mm PTFE membrane, 15 mm) (Agilent Technologies, catalog number: 5190-5085 )
- PLgel (10 μm, 7.5 x 300 mm, two pieces) (Agilent Technologies, model: MIXED-B column )
- PLgel (10 μm, 7.5 x 50 mm) (Agilent Technologies, model : Guard-column )
- Solid-state NMR spectrometer (Varian, model: 400-MHz ) with the following assembly (Figure 1):
Magnet: superconducting actively shielded magnet, B = 9.4 T, wide bore – 89 mm
Console: three high-power linear RF channels, all with spin lock and decoupling capability, output power up to 1000 W,- channel – high band RF, narrow band amplifier 375–400 MHz
- channel – broad band amplifier 18–240 MHz
- channel – broad band amplifier 10–130 MHz
a: 1H – 19F / 31P – 79Br / 23Na – 15N , 4 mm T3-HXY for solids, double and triple resonance, MAS up to 18 kHz
b: 1H – 19F / 31P – 79Br / 23Na – 15N, 7.5 mm T3-HXY for solids, double and triple resonance, MAS up to 7 kHz
c: Double channels goniometric static HX probe
X channel – 5, 10 mm coils, resonance from 31P to 15N
Accessories:
Variable temperature control unit, range –150 °C ↔ +250 °C
Accessory for low-band measurements – down from 15N
Accessories for sample preparation and measurements:
a.ZrO rotors with diameters of 4 and 7.5 mm
b. Rotor packing and cleaning tools, isopropyl alcohol, liquid nitrogen
c. Assembly for hydration and dehydration of samples
d. Primary and secondary standards for calibration of NMR ppm scale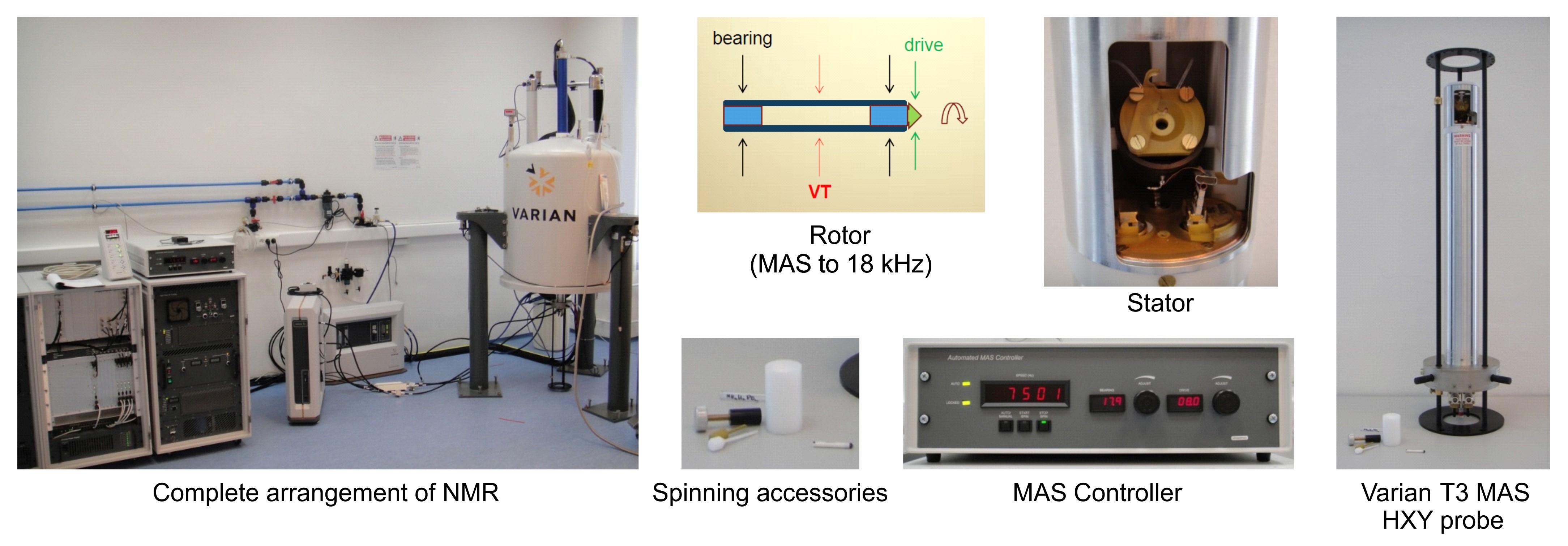
Figure 1. Solid-state 13C MAS NMR equipment - channel – high band RF, narrow band amplifier 375–400 MHz
Software
- ChemStation for LC 3D systems (Agilent Technologies)
- Clarity, GPC module, version 5.0.3.180 (DataApex)
- VnmrJ 3.2 (Varian)
- Mnova 8.1, or a higher version (Mestrelab Research)
Procedure
- Cellulose isolation from the stems of infected trees according to Seifert's method
- Select elm trees affected by Dutch elm disease (Figures 2A and 2D) and saw several wood discs, approximately 4 cm thick, from the stems at breast height 130 cm above a tree base. If forking does not allow disc sampling from the stem at this height, sample wood discs at the highest possible position on the stem below forking. Distinct infection zones are characterized by discoloration of the wood (Figures 2B and 2E) where the fungal hyphae grow and spread through secondary xylem vessels (Figures 2C and 2F).
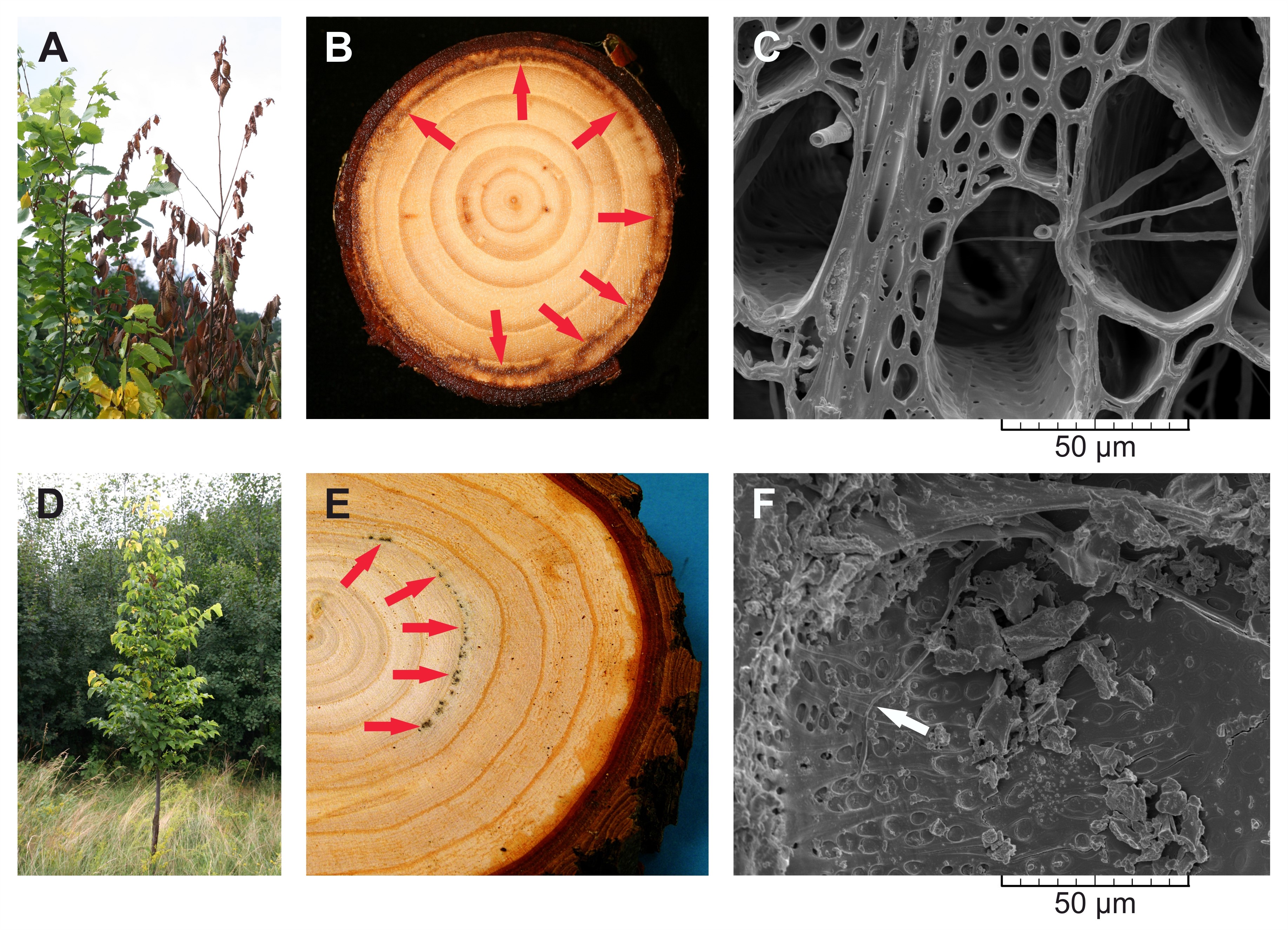

Figure 2. Dutch elm disease (DED) and the infection zones in Dutch elm hybrids 'Groeneveld' (susceptible to DED, A-C) and 'Dodoens' (tolerant to DED, D-F), which were artificially inoculated with spores of the pathogenic fungus Ophiostoma novo-ulmi ssp. americana x novo-ulmi. A. DED symptoms in an infected susceptible tree. B. Wood disc sawn from a susceptible tree, in which the infection zones (marked by red arrows) are restricted to the annual ring for current-year. Early the following growing season after an inoculation these infected trees died. C. Evidence of fungal hyphae growth inside latewood vessels within the infection zone. Cross-section in scanning electron microscopy, scale bar = 50 μm. D. Diminished symptoms of DED in an infected tolerant tree. E. Wood disc sawn from a tolerant tree which survived the infection. Infected trees continued to grow without any physiological weakening in the following years. Discolored infection zones are marked by red arrows. F. Occasional occurrence of a fungal hypha (white arrow) growing inside earlywood vessel within the infection zone. Radial section in scanning electron microscopy, scale bar = 50 μm. - Separate the annual ring sections which involve the infection zones from wood discs using a bandsaw, abrasive belt machine and woodworking lathe. See Video 1 for more detail. Disintegrated and extracted wood coming only from these sections of the annual ring is subjected to size-exclusion chromatography and 13C nuclear magnetic resonance measurements. Also, separate the identical annual ring sections from the control trees to enable comparisons of macromolecular traits of cellulose between the infected and the non-infected trees.
Video 1. Separation of the annual ring sections which involve the infection zones - Disintegrate separated wood sections into sawdust using a Polymix PX-MFC 90D mill.
- Sieve sawdust to a desirable fraction (size 0.50-1.00 mm) and dry in a desiccator.
- Extract the dry sawdust (5 g per 250 ml of the extraction solution per sample) according to the ASTM International standard procedure D1107-96 (2013) in a Soxhlet extraction apparatus with an ethanol-toluene solution (1 L absolute ethanol and 427 ml toluene) for 6 h.
- Dry the extracted sawdust in air on bench top overnight, then under vacuum at 50 ± 3 °C for at least 4 h.
- Place the dry wood sawdust (1 g), acetylacetone (6 ml), dioxane (2 ml), and hydrochloric acid (1.5 ml) into a 50 ml boiling flask.
- Heat the flask under reflux using a boiling water bath for 30 min, then allow the mixture to cool slowly to near room temperature and add methanol (30-40 ml) in the fume hood.
- Dry the filtering crucibles in an oven at 105 ± 3 °C for 2 h, then cool down in a desiccator to room temperature and weigh.
- Filter the mixture through the previously weighed filtering crucibles. Slowly rinse the solids using the following sequence: methanol (100 ml), followed by hot water (40 ml), dioxane (40 ml), and finally methanol (50 ml). During filtration, gently apply a vacuum to remove any liquids. Each rinse step should take approximately 2 min.
- Dry the crucible and acid insoluble residue (i.e., Seifert’s cellulose) at 105 ± 3 °C until a constant weight is achieved, usually a minimum of 90 min.
- Select elm trees affected by Dutch elm disease (Figures 2A and 2D) and saw several wood discs, approximately 4 cm thick, from the stems at breast height 130 cm above a tree base. If forking does not allow disc sampling from the stem at this height, sample wood discs at the highest possible position on the stem below forking. Distinct infection zones are characterized by discoloration of the wood (Figures 2B and 2E) where the fungal hyphae grow and spread through secondary xylem vessels (Figures 2C and 2F).
- Size-exclusion chromatography (SEC) of cellulose tricarbanilates
- Put Seifert’s cellulose (50 mg), pyridine (8.0 ml) and phenyl isocyanate (1.0 ml) into a 50 ml dropping flask.
- Place the sealed flask in an oil bath and heat in the oven at 70 ± 3 °C for 72 h.
- Cool to room temperature and add methanol (2.0 ml) to eliminate any excess phenyl isocyanate.
- Add the yellow solution dropwise into a rapidly magnetic stirring methanol: water (7:3) mixture (150 ml).
- Filtrate the precipitate and wash with methanol:water (7:3) mixture (1 x 50 ml), then with water (2 x 50 ml) to a neutral reaction (pH = 7.0).
- Dry the cellulose tricarbanilate (CTC) in air on bench top overnight, then under vacuum at 50 ± 3 °C.
- Dissolve CTC (2.0 mg) in tetrahydrofuran (THF) (2.0 ml) and filtrate through a Puradisc 25 NYL syringe filter (pore size 0.45 μm) into an autosampler vial using a 5 ml syringe.
- Inject the sample (10 μl) into a chromatograph and analyse by SEC at the following conditions: temperature of 35 °C, mobile phase (THF) flow rate of 1.0 ml/min on two PLgel, 7.5 x 300 mm, MIXED-B columns preceded by a PLgel, 7.5 x 50 mm, Guard-column.
- Acquire data with Chemstation software, and then import the data from Chemstation into the Clarity software. Calculate the molecular weights (Mn, Mw, Mz, Mz+1, Mp), degree of polymerization (DPw) and polydispersity index (PDI) of cellulose samples, and compare molecular weight distributions of cellulose tricarbanilates between the infected and the non-infected trees (Figure 3).
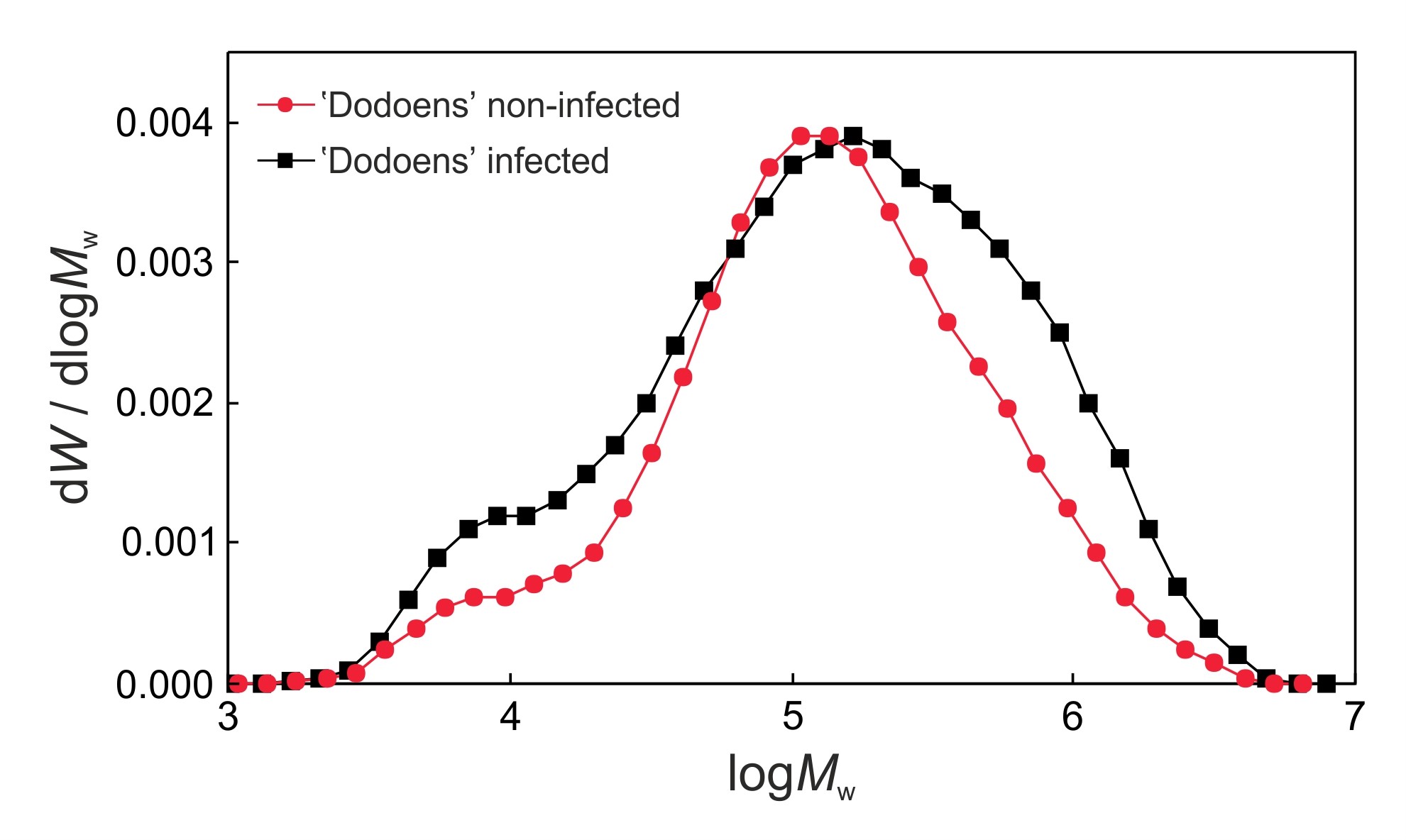
Figure 3. Size-exclusion chromatography of molecular weight distributions of cellulose tricarbanilates prepared from the non-infected and infected trees of the Dutch elm hybrid 'Dodoens'. Cellulolytic enzymes produced by O. novo-ulmi ssp. americana x novo-ulmi are responsible for the degradation of mostly medium molecular weight macromolecules of cellulose (DPw values of approximately 235 to 238) which results in an substantial increase in low molecular weight macromolecules (left side of the chromatogram). At the same time, however, the infected tree responds to the infection with a significant increase in the biosynthesis of high molecular weight macromolecules of cellulose (right side of the chromatogram), followed by a shift in the peak (Mp value) to the high molecular weight area. Co-occurring biosynthetic and biodegradation processes result in changes to the macromolecular traits of cellulose (Mn, Mw, Mz, Mz+1, DPw, PDI) in the infected elm trees. - Put Seifert’s cellulose (50 mg), pyridine (8.0 ml) and phenyl isocyanate (1.0 ml) into a 50 ml dropping flask.
- Solid-state 13C MAS NMR
- For the 13C MAS NMR measurement, start with a calibration of the chemical shift scale (ppm) and measure the 13C MAS NMR spectrum of adamantane.
- Fill the 4 mm ZrO rotor (volume of 52 μl) with a pulverized and compacted extracted sawdust sample.
- Insert the rotor into the stator of T3-HXY probe for solids, insert the probe into the magnet bore, and spin the rotor to the rate of 10 kHz.
- Tune 1H and 13C channels of the probe.
- Set up the parameters of the “onepul” pulse sequence (Figure 4) - π/2-pulse width, transmitter and decoupler offsets, spectral width, decoupling type and power, and acquisition time.
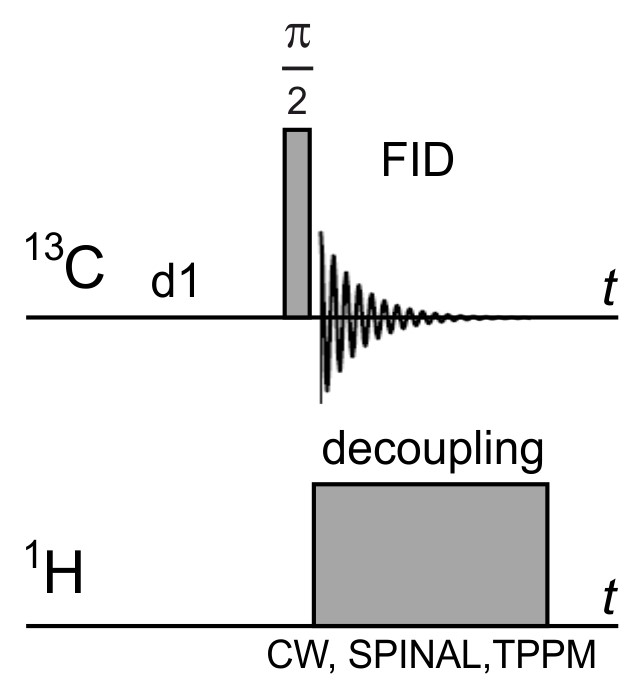
Figure 4. Scheme of the used pulse sequence - one pulse + dipolar decoupling + sample rotation at a magic angle- Measure and record the free induction decay (FID).
- Use Mnova 8.1 software, or a higher version, for the Fourier transformation of FIDs, as well as FID and NMR spectra processing. The software and the accompanying help manual can both be downloaded from the Mestrelab Research website: http://mestrelab.com/software/ and http://mestrelab.com/software/mnova/manuals/.
- Compare signal intensities of 13C MAS NMR spectra at both 83 ppm (corresponding to C4 carbon atoms of amorphous cellulose) and 89 ppm (corresponding to C4 carbon atoms of crystalline cellulose) between the infected and the non-infected trees (Figure 5). Based on peak heights, calculate crystallinity index and the percentage loss for both amorphous and crystalline regions, respectively.
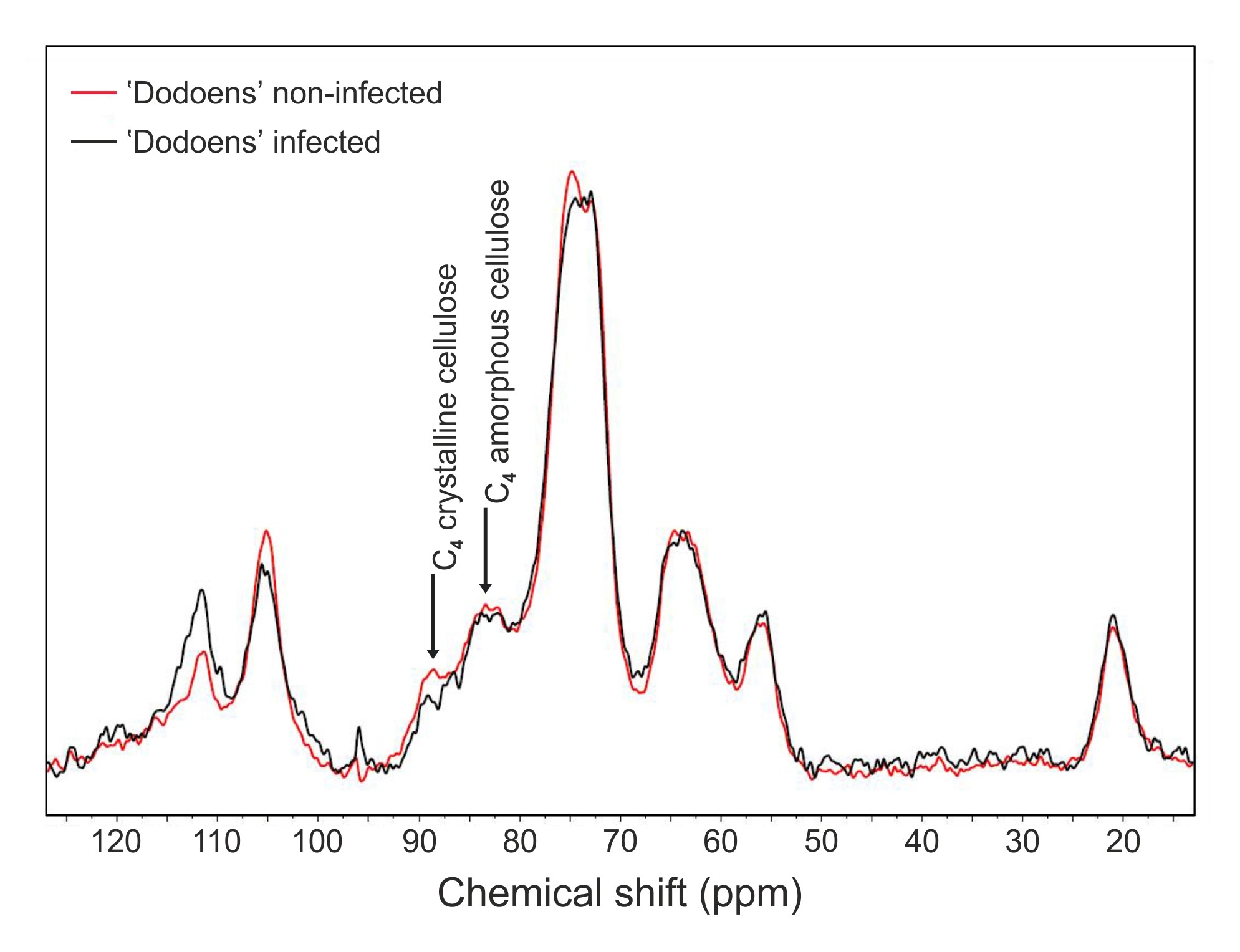
Figure 5. Solid-state 13C MAS NMR spectra of extractives-free samples from the non-infected and infected trees of the Dutch elm hybrid 'Dodoens'. Signals at 83 ppm and 89 ppm correspond to C4 carbon atoms of amorphous and crystalline cellulose, respectively. The decrease in signal intensities at both resonances reveals that losses in crystalline (26.09% drop) and non-crystalline (5.26% drop) cellulose regions have occurred in parallel. - For the 13C MAS NMR measurement, start with a calibration of the chemical shift scale (ppm) and measure the 13C MAS NMR spectrum of adamantane.
Notes
We have found that syringyl to guaiacyl (S/G) ratio in lignin affected the cellulose degradability by O. novo-ulmi in the infected elm trees (Ďurkovič et al., 2014). Other recent studies also revealed that an S/G ratio has a significant influence on the cross-linking between lignin and other cell wall components, thus modifying the microscopic structure and topochemistry of the cell wall, the cell wall degradability during chemical and hot-water pretreatments, and the successive hydrolysis of cellulose to glucose (Li et al., 2010; Studer et al., 2011; Papa et al., 2012). Therefore, we suggest using standard analytical methods such as alkaline nitrobenzene or cupric oxidations, NMR, pyrolysis–gas chromatography–mass spectrometry (Py–GC–MS) or others to determine lignin monomer composition as quantified by the S/G ratio. Thereby, both the lignin monomer composition and the cellulose degradation data can provide a more complete view of the biodegradation process caused by cellulose-degrading fungi.
Acknowledgments
The authors thank Dr. Miloň Dvořák, Dr. Jana Krajňáková, Dr. Miroslava Mamoňová, Dr. Ingrid Čaňová, Dr. Jaroslav Ohanka and Mr. Miroslav Rusnák for their technical assistance. This work was funded by the Slovak scientific grant agency VEGA (1/0149/15). This protocol has been adapted from our previous work (Ďurkovič et al., 2014).
References
- ASTM D1107-96. (2013). Standard test method for ethanol-toluene solubility of wood. American Society for Testing and Materials International, West Conshohocken, PA.
- Ďurkovič, J., Kačík, F., Olčák, D., Kučerová, V. and Krajňáková, J. (2014). Host responses and metabolic profiles of wood components in Dutch elm hybrids with a contrasting tolerance to Dutch elm disease. Ann Bot 114(1): 47-59.
- Li, X., Ximenes, E., Kim, Y., Slininger, M., Meilan, R., Ladisch, M. and Chapple, C. (2010). Lignin monomer composition affects Arabidopsis cell-wall degradability after liquid hot water pretreatment. Biotechnol Biofuels 3: 27.
- Papa, G., Varanasi, P., Sun, L., Cheng, G., Stavila, V., Holmes, B., Simmons, B. A., Adani, F. and Singh, S. (2012). Exploring the effect of different plant lignin content and composition on ionic liquid pretreatment efficiency and enzymatic saccharification of Eucalyptus globulus L. mutants. Bioresour Technol 117: 352-359.
- Studer, M. H., DeMartini, J. D., Davis, M. F., Sykes, R. W., Davison, B., Keller, M., Tuskan, G. A. and Wyman, C. E. (2011). Lignin content in natural Populus variants affects sugar release. Proc Natl Acad Sci U S A 108(15): 6300-6305.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Ďurkovič, J., Kačík, F. and Olčák, D. (2015). An Evaluation of Cellulose Degradation Affected by Dutch Elm Disease. Bio-protocol 5(14): e1535. DOI: 10.21769/BioProtoc.1535.
Category
Plant Science > Plant biochemistry > Carbohydrate
Plant Science > Plant immunity > Perception and signaling
Plant Science > Plant metabolism > Carbohydrate
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











