- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Proximity Ligation Assay (PLA) to Detect Protein-protein Interactions in Breast Cancer Cells
Published: Vol 5, Iss 10, May 20, 2015 DOI: 10.21769/BioProtoc.1479 Views: 34989
Reviewed by: HongLok LungShannon RuppertAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
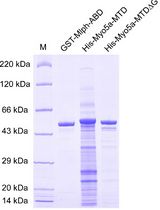
Determination of Dissociation Constants for the Interaction of Myosin-5a with its Cargo Protein Using Microscale Thermophoresis (MST)
Rui Zhou [...] Xiang-Dong Li
Feb 5, 2025 1788 Views
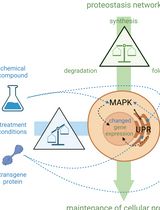
Cell-Sonar, an Easy and Low-cost Method to Track a Target Protein by Expression Changes of Specific Protein Markers
Sabrina Brockmöller [...] Simone Rothmiller
Feb 5, 2025 1682 Views
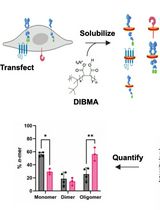
SiMPull-POP: Quantification of Membrane Protein Assembly via Single Molecule Photobleaching
Ryan J. Schuck [...] Rajan Lamichhane
Jan 5, 2026 353 Views
Abstract
Protein-protein interaction networks provide a global picture of cellular function and biological processes, and the dysfunction of some interactions causes many diseases, including cancer. The in situ proximity ligation assay (PLA) is a powerful technology capable of detecting the interactions among proteins in fixed tissue and cell samples. The interaction between two proteins is detected using the corresponding two primary antibodies raised in different species. Species-specific secondary antibodies (PLA probes), each with a unique short DNA strand attached to it, bind to the primary antibodies. When the PLA probes are in close proximity (<40 nm), the DNA strands can interact through a subsequent addition of two other circle-forming DNA oligonucleotides. Several-hundredfold replication of the DNA circle can occur after the amplification reaction, and a fluorescent signal is generated by labelled complementary oligonucleotide probes. Therefore, each detected signal is visualized as an individual fluorescent dot, which can be quantified and assigned to a specific subcellular location based on microscopy images. This revolutionary technique enables us to study the protein complex formation with high specificity and sensitivity compared to the other traditional methods, such as co-immunoprecipitation (Co-IP).
Materials and Reagents
- Antibody pair for detecting protein-protein interaction (two primary antibodies must have been raised in different species, e.g. rabbit-anti-EGFR from Cell Signaling Technology, catalog number: 2232 ; mouse-anti-DNA-PK from BD Pharmingen, catalog number: 556456 )
- DuolinkTM in situ reagents
- DuolinkTM in situ PLA reagents
- Duolink in situ complementary oligonucleotide probe MINUS and PLUS [5x, secondary antibody conjugated with a PLA oligonucleotide, the choice of PLA probes depends on the species of your primary antibodies: e.g. anti-rabbit PLUS (Sigma-Aldrich, catalog number: DUO82029 ); anti-mouse MINUS (Sigma-Aldrich, catalog number: DUO 92004)].
- Blocking solution (Sigma-Aldrich, catalog number: DUO82014 ): For blocking of the sample if you have not already optimized your primary antibody with another blocking solution, e.g. 2% BSA in PBS.
- Antibody diluent (Sigma-Aldrich, catalog number: DUO82015 ): For dilution of PLA probes and the primary antibodies, alternatively PBS solution with 1% BSA works as well.
- Duolink in situ complementary oligonucleotide probe MINUS and PLUS [5x, secondary antibody conjugated with a PLA oligonucleotide, the choice of PLA probes depends on the species of your primary antibodies: e.g. anti-rabbit PLUS (Sigma-Aldrich, catalog number: DUO82029 ); anti-mouse MINUS (Sigma-Aldrich, catalog number: DUO 92004)].
- Duolink detection reagents
- Ligation reagents (5x, contains oligonucleotides that hybridize to the PLA probes and all components needed for ligation except the Ligase) (Sigma-Aldrich, catalog number: DUO82016 )
- Ligase (1 unit/μl) (Sigma-Aldrich, catalog number: DUO82029)
- Amplification reagents (5x, contains all components needed for Rolling Circle Amplification except the Polymerase. Included are also oligonucleotide probes labelled with a fluorophore that hybridize to the RCA product): Containing far red fluorescently labelled oligonucleotides (Sigma-Aldrich, catalog number: DUO82019 )
- Polymerase (10 unit/μl): (Sigma-Aldrich, catalog number: DUO82030 )
The Duolink detection reagents are available to purchase as a kit from Sigma-Aldrich (e.g. catalog number for Duolink in situ Detection Reagents Far Red DUO92013 ).
- Ligation reagents (5x, contains oligonucleotides that hybridize to the PLA probes and all components needed for ligation except the Ligase) (Sigma-Aldrich, catalog number: DUO82016 )
- Duolink washing buffer A and B (see recipes)
- DuolinkTM in situ PLA reagents
- ProLong Gold antifade reagent with DAPI (Life Technologies, catalog number: P-36931), alternatively regular immunofluorescence mounting media and DAPI can be used separately.
- Reagents required for fixation and permeabilization of the sample (e.g. 3.7% fresh-made formaldehyde and 0.1% Triton X-100 diluted in PBS)
Equipment
- Fluorescence microscope equipped as follows (e.g. Leica TCS SP5 Microsystems)
- Excitation/emission filters compatible with fluorophore (ranging from 488-633 nm) and nuclear stain (ultraviolet) excitation/emission
- Camera and software for image acquisition
- Excitation/emission filters compatible with fluorophore (ranging from 488-633 nm) and nuclear stain (ultraviolet) excitation/emission
- Shaker
- Humidity chamber (moist chamber) (Figure 1)
- Freeze block for enzymes
- 37 °C incubator
- Pipettes (covering the range from 1 μl to 1,000 μl)
- Glass cover slips compatible with fluorescence microscopy (12 mm diameter and 0.13 to 0.16 mm thickness) (Optics, catalog number: 01 115 20 )
- MilliQ® or other equivalently high purity water
Software
- Duolink ImageTool software is highly recommended (Demo version for free-download: http://www.olink.com/products/duolink/downloads/duolink-image-tool). Alternatively, ImageJ or LAS AF Lite software from Leica Microsystems can be used for analysis.
Procedure
- The primary antibodies need to be optimized through concentration titration to make sure that they work properly before the start of proximity ligation assay (usually ranging from 1:100 to 1:500), and IgG from the same isotype and species was usually used as negative control.
- Breast cancer cells (e.g. Hs578T and MDA-MB-468) were transferred on the coverslip in plates/culture dishes and grown to 50%-70% confluency.
- The cells were washed with PBS, fixed with 3.7% formaldehyde for 15 min, permeabilized with 0.1% Triton X-100 for 10 min, washed, and then blocked using Duolink blocking buffer for 1 h (all these steps are usually performed at room temperature; if you have previously optimized your assay, use the same conditions for Duolink).
- The coverslips were transferred into a humidified chamber (Figure 1) and then a small volume of primary antibodies targeting the proteins under investigation were pipetted onto each coverslip (for a 12 mm coverslip, 20-40 µl of antibody added in antibody diluent is more than enough). Antigen-antibody interaction will mostly accomplish in an hour at room temperature, or overnight at 4 °C with gentle agitation (the residual solution can be absorbed using tissues to obtain an equal residual volume on each slide as this will affect reproducibility, However, do not allow the samples to dry before adding the primary antibodies as this will cause background).
- The two corresponding PLA probes were mixed and diluted (1:5) in the antibody diluent. Allow the mixture to sit for 20 min at room temperature (e.g. for a 50 μl reaction take 10 μl of PLA probe MINUS stock, 10 μl of PLA probe PLUS stock and 30 μl of the antibody diluent). The samples were washed with PBS+0.05% Tween twice, and then incubated with PLA probes MINUS and PLUS for 1 h at 37 °C.
- The samples were washed in 1x wash buffer A for 2 x 5 min under gentle agitation, and the probes were ligated with two other circle-forming DNA oligonucleotides by ligation-ligase solution for 30 min at 37 °C (1:40 dilution, consisting of two oligonucleotides and ligase to hybridize oligonucleotides to the two PLA probes and join to a closed circle if they are in close proximity).
- The samples were washed in 1x wash buffer A for 2 x 2 min under gentle agitation and two added oligonucleotides by enzymatic ligation were amplified via rolling circle amplification by the incubation with amplification-polymerase solution (1:80 dilution) over 90 min at 37 °C (consisting of nucleotides and fluorescently labelled oligonucleotides, which is added together with polymerase. The fluorescently labelled oligonucleotides will hybridize to the rolling-circle amplification (RCA) product using the ligated circle as a template, and the signal is easily visible as a distinct fluorescent spot by fluorescence microscope).
- The samples were washed in 1x wash buffer B for 2 x 10 min followed by 0.01x wash buffer B for 1 min by diluting 1x buffer B 1:100 in high purity water.
Note: In steps 6-8 above, the volume of wash buffers A and B depends on the size of coverslip, for a small one (12 mm), we usually wash in a 12-well plate using 1 ml wash buffer. - The samples were dried in hood for approximately 10 min at room temperature in the dark, and mounted onto a slide in a minimal volume of ProLong Gold antifade reagent with DAPI.
- Fluorescent signal amplification was used for the assay and interactions were detected using a fluorescence or confocal microscope, using at least a 20x objective. After imaging, the slides can be stored at -20 °C in the dark.

Figure 1. The chamber for incubation. The 6-well plate was covered with a piece of parafilm to support the coverslip with cells, and another lid was used to prevent sample drying. The chamber is designed to be incubated in a tissue culture incubator at 37 °C for most of the PLA reactions.
Representative data

Figure 2. Duolink proximity ligation assay for protein interactions between epidermal growth factor receptor (EGFR) and DNA-dependent protein kinase (DNA-PK) in breast cancer cell line MDA-MB-468. The cells were treated with DNA-damaging agent etoposide (20 μM) at different time points, and then Duolink assay between EGFR and IGFBP-3 performed as described (cells without treatment as negative controls). Each red spot represents for a single interaction and DNA was stained with DAPI.
Recipes
- Duolink in situ wash buffer A
8.8 g NaCl
1.2 g Tris base
0.5 ml Tween 20
Adjust pH to 7.4 with HCl
Add high purity dH2O to 1,000 ml (final concentrations 0.01 M Tris, 0.15 M NaCl and 0.05% Tween 20)
Filter sterilize (0.22 μm)
Stored at 4 °C, bring the solutions to room temperature before use - Duolink in situ wash buffer B
5.84 g NaCl
4.24 g Tris base
26.0 g Tris-HCl
Adjust pH to 7.5 using HCl
Add high purity dH2O to 1,000 ml (final concentrations 0.2 M Tris and 0.1M NaCl)
Filter the solution through a 0.22 μm filter
Stored at 4 °C. Bring the solutions to room temperature before use
Acknowledgments
This work was supported by Grant Number DP0984232 to RCB from the Australian Research Council.
References
- Soderberg, O., Gullberg, M., Jarvius, M., Ridderstrale, K., Leuchowius, K. J., Jarvius, J., Wester, K., Hydbring, P., Bahram, F., Larsson, L. G. and Landegren, U. (2006). Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat Methods 3(12): 995-1000.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Lin, M. Z., Martin, J. L. and Baxter, R. C. (2015). Proximity Ligation Assay (PLA) to Detect Protein-protein Interactions in Breast Cancer Cells . Bio-protocol 5(10): e1479. DOI: 10.21769/BioProtoc.1479.
Category
Cancer Biology > General technique > Biochemical assays > Protein analysis
Molecular Biology > Protein > Protein-protein interaction
Biochemistry > Protein > Interaction > Protein-protein interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











