- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Random DNA Binding Selection Assay (RDSA)
Published: Vol 5, Iss 8, Apr 20, 2015 DOI: 10.21769/BioProtoc.1452 Views: 10353
Reviewed by: Tie LiuAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Determination of Protein-DNA (ZMYND11-DNA) Interaction by a Label-Free Biolayer Interferometry Assay
Yuan-Yuan Li [...] Hai-Tao Li
Feb 20, 2015 13578 Views
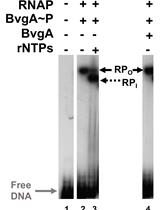
Combining Gel Retardation and Footprinting to Determine Protein-DNA Interactions of Specific and/or Less Stable Complexes
Meng-Lun Hsieh [...] Deborah M. Hinton
Dec 5, 2020 3590 Views
A Gel-Based Assay for Probing Protein Translocation on dsDNA
Christiane Brugger and Alexandra M. Deaconescu
Jul 20, 2021 4343 Views
Abstract
Protein-DNA interaction is a very important cellular process, by which regulation of DNA biological function, usually gene expression, is exerted. The method of random DNA binding selection assay (RDSA) can be used to identify DNA elements bound by proteins with DNA-binding activities. This method is based on the enrichment of the target DNA element by the immobilized recombinant protein on special beads and repeated PCR amplification.
Keywords: Protein-DNA interactionMaterials and Reagents
- The prokaryotic expression vectors: pGEX-6P-1 (GE Healthcare) or pTXB3 (New England Biolabs)
- The expression strain [Escherichia coli (E.coli) BL21 (DE3) or others]
- T-Vector pGEMT-easy and T4-DNA ligase (Promega Corporation, catalog number: A1360 )
- DNA polymerase (Takara Bio Company, catalog number: DR001 )
- Sterile ddH2O (double distilled water)
- Oligonucleotides (Pitzschke et al., 2009; Wang et al., 2014)
- Chitin resin (New England Biolabs, catalog number: S6651S )
- PBS buffer (see Recipes)
- Column buffer (see Recipes)
- RDSA buffer (Pitzschke et al., 2009) (see Recipes)
Table 1. Oligonucleotides for RDSA
Oligonucleotides for RDSA
RDSA_1
TAGTTGAGTCTCACAAACGAACAC(N20)CATTCCAAAATCCATGGCTGATA
RDSA_1 fo
TAGTTGAGTCTCACAAACGAACAC
RDSA_1 re
TATCAGCCATGGATTTTGGAATG
RDSA_2
AATGGATCCAAGCTTAAGC(N18)CGTTGAATTCCCATGGACA
RDSA_2 fo
AATGGATCCAAGCTTAAGC
RDSA_2 re
TGTCCATGGGAATTCAACG
Equipment
- Glutathione sepharose beads (GE Healthcare)
- The empty chromatography columns (Bio-Rad Laboratories)
- 4 °C refrigerator
- Centrifuge
- Whirling shaker
Procedure
- Random DNA preparation
- PCR amplification.
The double strand DNA fragments RDSA1 and RDSA2 with 20- and 18-bp random sequence regions flanked by independent sets of adaptors were amplified on the templates of RDSA_1 and RDSA_2 by PCR, using RDSA_1/2fo and RDSA_1/2re primers, respectively (Table 1). Conditions for PCR amplification are: 5 min at 94 °C; 30 cycles of 30 sec at 94 °C, 30 sec at 50 °C, and 30 sec at 72 °C; 10 min at 72 °C.
- Purification of PCR products.
- Take at least 200 μl PCR products; add equal volume of 24:1 (chloroform: isoamyl alcohol) to sample tubes; shake tubes vigorously by hand for at least 30 sec, centrifuge the sample at 15,000 x g for 10 min at 4 °C.
- Transfer the aqueous phase to fresh tubes; add equal volume of isopropanol and mix; place in -20 °C freezer for 1 h.
- Centrifuge the samples at 15,000 x g for 10 min at 4 °C.
- Discard the supernatants and wash the precipitates with 70% alcohol.
- Air dry the precipitates for at least 10 min; dissolve the precipitates in sterile ddH2O and store at -20 °C until use.
- Take at least 200 μl PCR products; add equal volume of 24:1 (chloroform: isoamyl alcohol) to sample tubes; shake tubes vigorously by hand for at least 30 sec, centrifuge the sample at 15,000 x g for 10 min at 4 °C.
- PCR amplification.
- Prokaryotic expression of target protein tagged with GST or CBD (chitin binding domain)
- Construct expression vector.
The coding region of the target gene is ligated in frame into the expression vector pGEX-6P-1 (for GST fusion protein) or pTXB3 (for CBD fusion protein).
- The constructs are transformed into E.coli expression strain BL21 (DE3) to express fusion proteins.
- Construct expression vector.
- RDSA cycles
- Harvest cells (100 ml bacterial cultures) and resuspend them in PBS buffer (for GST fusion protein) or column buffer (for CBD fusion protein).
- Break cells by sonication (950 W x 20%, 2-10 min according to the solubility of the expressed protein and with 2 sec interruption after every 2 sec sonication. The tube with sample should be placed in ice-water during manipulation) and centrifuge at 1,000 x g for 10 min at 4 °C.
- Prepare beads columns (load at least 3 ml beads in each chromatography column); equilibrate the glutathione sepharose beads column (for GST fusion protein) with PBS buffer (ten times of the bead volume), and the chitin beads column (for CBD fusion protein) with column buffer (ten times of the bead volume). In the following steps, the beads columns should be placed on ice.
- Slowly load the clarified lysate on to the columns.
- Wash the columns with at least 20 times of bed volume PBS buffer for GST fusion protein or column buffer for CBD fusion protein to thoroughly remove the unbound proteins; wash the beads at least three times with RDSA buffer; resuspend the beads in 3 ml RDSA buffer. Now the recombinant target protein is immobilized on beads.
- Divide the beads into 5-10 tubes (1.5 ml tube).
- Add the random DNA fragment RDSA1 or RDSA2 (10 μl, about 5-10 μg) into one tube; incubate for more than 2 h at 4 °C with gentle rolling.
- Centrifuge at 500 x g for 5 min at 4 °C, discard the supernatant completely.
- Wash the beads with RDSA buffer for at least four times to remove unbound DNA fragments; add 500 µl sterile ddH2O and boil the beads for 5 min to release the bound DNA; centrifuge at 500 x g for 5 min at 4 °C.
- Transfer the supernatant to a fresh tube; purify and precipitate the released DNA as described in step A2 (do not need to dissolve the DNA).
- Add PCR reagents to the tube:
Perform PCR amplification using the following parameters: 5 min at 94 °C; 30 cycles of 30 sec at 94 °C, 30 sec at 50 °C, and 30 sec at 72 °C; 10 min at 72 °C.74 µl
ddH2O
10 µl
10x PCR buffer
10 µl
2.5 mM dNTP
2.5 µl
20 μM RDSA1 fo or RDSA2 fo primer
2.5 µl
20 μM RDSA1 re or RDSA2 re primer
1.0 µl
rTaq DNA polymerase
- Purify the PCR products as described in A2; dissolve the DNA in 10 μl sterile ddH2O and transfer the DNA to another tube in C6 for the next RDSA cycle (repeat steps C7-12).
- After 5 to 10 cycles, clone the purified PCR products into pGEM-T easy vector and sequence.
- Harvest cells (100 ml bacterial cultures) and resuspend them in PBS buffer (for GST fusion protein) or column buffer (for CBD fusion protein).
Notes
- By increasing the amounts of the non-specific competitor poly (dIdC) (0, 50, 100, 150, 200, 250, 300 and 350 ng) and reducing the amounts of protein in the binding reactions (700 μl, 650 μl, 600 μl, 550 μl, 500 μl, 400 μl, 300 μl, 200 μl protein bound beads) the stringency can be increased in each RDSA cycle (Pitzschke et al., 2009; Wang et al., 2014).
- To exclude effects from the nonrandom flanking adaptors, the RDSAs must be performed in duplicates with two different sets of input random DNA fragments (RDSA1 and RDSA2) (Pitzschke et al., 2009).
- The number of total cycles may be different for different proteins. To our experience, at least 5 cycles should be performed to exclude the non-specific binding.
- The results of RDSAs should be verified by other methods, such as electrophoretic mobility shift assay (EMSA) and yeast one-hybrid system.
- The annealing temperature should be adjusted by different random primers.
Recipes
- PBS buffer (pH 7.3)
140 mM NaCl
2.7 mM KCl
10 mM Na2HPO4
1.8 mM KH2PO4
5 mM DTT
1 mM PMSF
- Column buffer
20 mM Tris-HCl (pH 8.5)
500 mM NaCl
1 mM EDTA
- RDSA buffer
5 mM Tris (pH 8.0)
75 mM NaCl
2.5 mM MgCl2
0.5 mM EDTA
5% glycerol
1% Tween
1 mM DTT
Acknowledgments
We thank Dr. Heribert Hirt for suggestions on this experiment. This work was supported by the Natural Science Foundation of China (grant no. 30971748).
References
- Pitzschke, A., Djamei, A., Teige, M. and Hirt, H. (2009). VIP1 response elements mediate mitogen-activated protein kinase 3-induced stress gene expression. Proc Natl Acad Sci U S A 106(43): 18414-18419.
- Wang, Y., Peng, W., Zhou, X., Huang, F., Shao, L. and Luo, M. (2014). The putative Agrobacterium transcriptional activator-like virulence protein VirD5 may target T-complex to prevent the degradation of coat proteins in the plant cell nucleus. New Phytol 203(4): 1266-1281.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Wang, Y. and Luo, M. (2015). Random DNA Binding Selection Assay (RDSA). Bio-protocol 5(8): e1452. DOI: 10.21769/BioProtoc.1452.
Category
Molecular Biology > DNA > DNA-protein interaction
Molecular Biology > DNA > Gene expression
Biochemistry > Protein > Interaction > Protein-DNA interaction
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









