- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Membrane Flotation Assay
Published: Vol 5, Iss 7, Apr 5, 2015 DOI: 10.21769/BioProtoc.1435 Views: 14224
Reviewed by: Fanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
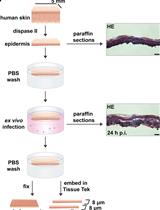
Ex vivo Human Skin Infection with Herpes Simplex Virus 1
Nydia C. De La Cruz [...] Dagmar Knebel-Mörsdorf
May 5, 2022 3064 Views
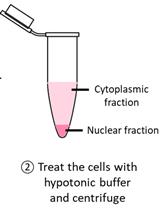
Detection of Cytoplasmic and Nuclear Circular RNA via RT-qPCR
Ke-En Tan [...] Yat-Yuen Lim
Sep 5, 2023 3405 Views

Extraction of Bacterial Membrane Vesicle and Phage Complex by Density Gradient Ultracentrifugation
Shangru Li [...] Tianyuan Jia
Aug 20, 2024 2054 Views
Abstract
Many postitive-stranded RNA viruses, such as Hepatitis C virus (HCV), highjack cellular membranes, including the Golgi, ER, mitchondria, lipid droplets, and utilize them for replication of their RNA genome or assembly of new virions. By investigating how viral proteins associate with cellular membranes we will better understand the roles of cellular membranes in the viral life cycle. Our lab has focused specifically on the role of lipid droplets and lipid-rich membranes in the life cycle of HCV. To analyze the role of lipid-rich membranes in HCV RNA replication, we utilized a membrane flotation assay based on an 10-20-30% iodixanol density gradient developed by Yeaman et al. (2001). This gradient results in a linear increase in density over almost the entire length of the gradient, and membrane particles are separated in the gradient based on their buoyant characteristics. To preserve membranes in the lysate, cells are broken mechanically in a buffer lacking detergent. The cell lysate is loaded on the bottom of the gradient, overlaid with the gradient, and membranes float up as the iodixanol gradient self-generates. The lipid content of membranes and the concentration of associated proteins will determine the separation of different membranes within the gradient. After centrifugation, fractions can be sampled from the top of the gradient and analyzed using standard SDS-PAGE and western blot analysis for proteins of interest.
Materials and Reagents
- Huh7.5 cells
- Huh7.5 cells containing replicating HCV Replicon RNA (= Replicon Cells) (Vogt et al., 2013)
- Dulbecco’s Modified Eagle’s Medium (with 4.5 g/L glucose, L-glutamine & Sodium pyruvate) (DMEM) (Corning Cellgro, catalog number: 10-013-CV )
- Fetal Bovine Serum (FBS) (Benchmark, catalog number: 100-106 )
- Penicillin/Streptomicyn Solution 100x (Pen/Strep) (Corning Cellgro, catalog number: 30-002-Cl )
- L-Glutamine (Corning Cellgro, catalog number: 25-005-CI )
- G418 sulfate (Corning Cellgro, catalog number: 25-052-CI )
- Dulbecco’s Phosphate Buffered Saline without Ca2+ and Mg2+ (PBS) (Corning Cellgro, catalog number: 21-031-CV )
- Trypsin/EDTA (Corning Cellgro, catalog number: 25-052-CI)
- Protease inhibitor cocktail (Sigma-Aldrich, catalog number: P8340 ) (use as 1:100 dilution)
- Trypan Blue Stain (Gibco, catalog number: 15250-061 )
- BioRad Protein Assay (Bio-Rad Laboratories, catalog number: 500-0006 )
- Brilliant Blue G-250 (Thermo Fisher Scientific, catalog number: 100-25 )
- Iodixanol (60%) (Sigma-Aldrich, catalog number: D1566-250ML ) (kept at 4 °C for use in the experiment )
- PBS/Sucrose (see Recipes)
- 20% iodixanol (see Recipes)
- 10% iodixanol (see Recipes)
- 2x Laemmli buffer (see Recipes)
Equipment
- Ultra clear centrifuge tubes (Beckman Coulter, catalog number: 344059 )
- 37 °C 5% CO2 cell culture incubator
- P1000 pipette
- Microscope
- Hematocytometer
- Tight fitting dounce homogenizer (7 ml) (Wheaton, catalog number: 357542 )
- Centrifuge (Beckman Coulter, model: Allegra 6R )
- Spectrophotometer
- Ultracentrifuge (Beckman Coulter, model: Optima L-80 XP with SW41T rotor)
- Protein electrophoresis apparatus
- Western blot apparatus
Procedure
- Huh7.5 or HCV replicon cells are grown in DMEM supplemented with 10% FBS, 1x Pen/Strep, 1x Glutamine for 4-5 days from 20% to a 80% final confluency in T175 flasks (will need 5 flasks per cell type). The medium for HCV replicon cells is additionally supplemented with 800 μg/ml of G418.
- Cells are harvested by washing once with 10 ml of PBS (add PBS to flask, agitate by hand 2-3 times and aspirate PBS off), then trypsinized with 4 ml of trypsin per T175 flask (incubate cells with trypsin at 37 °C for 3-5 min, take off cells with 6 ml of PBS by washing the walls of the flask and transferring to a 50 ml cornical tube). Cells are spun down for 5 min at 400 x g, the supernatant is aspirated off, and cells are washed once with 30 ml PBS by resuspending cells in PBS and spun down again for 5 min at 400 x g, then resuspended in 10 ml PBS, and counted in a hematocytometer. A total of 3 x 107 cells for each sample are resuspended in 3.5 ml pre-chilled PBS/Sucrose plus protease inhibitor cocktail (diluted 1:100) and immediately stored on ice.
- The cells are lysed with 200 passages in a tight-fitting dounce homogenizer, while keeping the homogenizer on ice.
Note: Be careful not to lift the pestle above the surface of the lysate to minimized creation of air bubbles.
- Check that cells are over 90% lysed by mixing 10 μl of the lysate with 10 μl Trypan Blue stain, then checking the sample in the microscope. Dead cells will be stained blue. If not enough cells are lysed, lyse cells with additional passages in the dounce homogenizer, and check every 50 passages that cell lysis was successful.
- The cell lysate is spun at 2,500 x g for 10 min at 4 °C to pellet cellular debris and nuclei.
- Transfer the supernatant (referred to as crude lysate) to a fresh tube and keep on ice.
- Measure total protein concentration in the crude lysate using Bio-Rad Protein Assay according to the manufacturer’s protocol.
- Take 5 mg of protein for each sample and add PBS/Sucrose to a total volume of 2 ml. Freeze the rest of the crude lysate mixed 1:1 with 2x Laemmli buffer as a control.
- Mix 2 ml of sample 1:1 with 2 ml of 60% iodixanol (pre-chilled at 4 °C) to get 4 ml of sample at a final concentration of 30% iodixanol.
- Put the 4 ml of 30% iodixanol/lysate mixture on the bottom of a centrifuge tube. Overlay carefully with 4 ml of 20% iodixanol (pre-chilled), then overlay with 4 ml of 10% iodixanol (pre-chilled). Then fill the tube with 10% until the level reaches 1 mm below the rim of the centrifuge tube. (Overlay with a P1000 pipette by cutting off 1 cm of the pipette tips, then pipette slowly 1 ml at the time right into the middle of the centrifuge tube right on top of the gradient level, to minimize mixing.) Make sure the tubes are balanced well for centrifugation, add carefully 10% Iodixanol on top to balance tubes if necessary.
- Spin at 209,000 x g (35,000 rpm) overnight (16 h) at 4 °C in an SW41T rotor.
- Next day: Collect 22 fractions (500 μl) each from top to bottom. Use cut off pipette tips of P1000 and carefully take off 500 μl from the middle by just touching the pipette tip to the surface of the gradient, then lowering the tip slowly to stay right at the surface of the gradient while taking off the sample very slowly. The increased width of the pipette tip decreases a whirlpool effect and minimizes mixing of the gradient layers.
- Mix fractions 1:1 with 2x Laemmli buffer, and freeze, preferable in aliquots, at -80 °C.
- Samples can then be analyzed via SDS-PAGE and Western blots [see results in Vogt et al. (2013)].
Notes
- This protocol could be applied to other cell lines as well. The distribution among the fractions at the end may vary depending on the cell line. Cells lines with a very low lipid content may not work as well as the Huh7.5, however, the general protocol could be applied (and could be optimized a bit) to most cell lines since they all contain cellular membranes.
- For detailed information on how to determine lipid content for whole cell lysate, readers are recommended to read the review article by Camus et al. (2013).
Recipes
- PBS/Sucrose
Dissolve 8.55 g sucrose in 100 ml of PBS to a final concentration of 0.25 M sucrose in PBS
Kept at 4 °C
- 20% Iodixanol
10 ml of Iodixanol (60%)
20 ml of PBS/Sucrose
Kept at 4 °C
- 10% Iodixanol
5 ml of Iodixanol (60%)
25 ml of PBS/Sucrose
Kept at 4 °C
- 2x Laemmli buffer
125 mM tris-HCl (pH 6.8)
20% glycerol
2.5% SDS
2.5 mg/100 ml Brilliant Blue G-250
Mix fresh before use: 950 μl of 2x Laemmli buffer with 50 μl β-mercaptoethanol
Acknowledgments
This work was supported by funds from the Gladstone Institutes, and the National Institutes of Health (R056 AI069090 (MO), and P30 DK026743 (MO) (University of California – San Francisco Liver Center). We gratefully acknowledge support through the Training Grant (T32 DK060414) from the US National Institute of Health to DAV.
References
- Camus, G., Vogt, D. A., Kondratowicz, A. S. and Ott, M. (2013). Lipid droplets and viral infections. Methods Cell Biol 116: 167-190.
- Vogt, D. A., Camus, G., Herker, E., Webster, B. R., Tsou, C. L., Greene, W. C., Yen, T. S. and Ott, M. (2013). Lipid droplet-binding protein TIP47 regulates hepatitis C Virus RNA replication through interaction with the viral NS5A protein. PLoS Pathog 9(4): e1003302.
- Yeaman, C., Grindstaff, K. K., Wright, J. R. and Nelson, W. J. (2001). Sec6/8 complexes on trans-Golgi network and plasma membrane regulate late stages of exocytosis in mammalian cells. J Cell Biol 155(4): 593-604.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Vogt, D. A. and Ott, M. (2015). Membrane Flotation Assay. Bio-protocol 5(7): e1435. DOI: 10.21769/BioProtoc.1435.
Category
Microbiology > Microbe-host interactions > Ex vivo model
Microbiology > Microbial cell biology > Organelle isolation
Cell Biology > Organelle isolation > Fractionation
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link