- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Purification of Starch Granules from Arabidopsis Leaves and Determination of Granule-Bound Starch Synthase Activity
Published: Vol 4, Iss 23, Dec 5, 2014 DOI: 10.21769/BioProtoc.1316 Views: 10018
Reviewed by: Tie LiuXiao-qing Xu

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
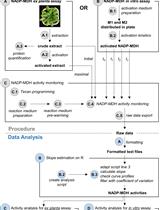
A Semi-throughput Procedure for Assaying Plant NADP-malate Dehydrogenase Activity Using a Plate Reader
Kevin Baudry and Emmanuelle Issakidis-Bourguet
Aug 20, 2023 1474 Views
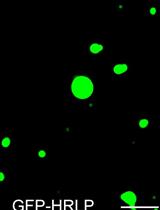
An in vitro Assay to Probe the Formation of Biomolecular Condensates
Yu Zhang and Shen Lisha
Sep 5, 2023 3209 Views
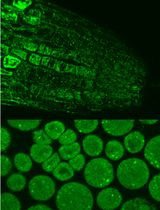
Immunofluorescence for Detection of TOR Kinase Activity In Situ in Photosynthetic Organisms
Ana P. Lando [...] Giselle M. A. Martínez-Noël
Dec 20, 2024 1824 Views
Abstract
Starch constitutes the most important carbon reserve in plants and is composed of branched amylopectin and linear amylose. The latter is synthesized exclusively by the Granule-Bound Starch Synthase (GBSS, EC 2.4.1.21). Here we report a readily reproducible, specific and highly sensitive protocol, which includes the isolation of intact starch granules from Arabidopsis thaliana leaves and the subsequent determination of GBSS activity. We have applied this method to study GBSS activity in diurnal cycles in vegetative growth and during the photoperiodic transition to flowering in Arabidopsis (Tenorio et al., 2003; Ortiz-Marchena et al., 2014).
Keywords: Starch granuleMaterials and Reagents
- Plant materials
Note: Arabidopsis thaliana (A. thaliana) were grown in controlled cabinets on peat-based compost. - Liquid N2
- HEPES (Sigma-Aldrich, catalog number: H4034 )
- Potassium hydroxide pellets (Panreac Applichem, catalog number: A0566 )
- TritonTM X-100 (Sigma-Aldrich, catalog number: X100 )
- Miracloth (Merck KGaA, catalog number: 475855 )
- Percoll® (Sigma-Aldrich, catalog number: P1644 )
- Glycogen (from rabbit liver Type III) (Sigma-Aldrich, catalog number: G8876 )
- Tricine (Sigma-Aldrich, catalog number: T0377 )
- Potassium acetate (Sigma-Aldrich, catalog number: P1147 )
- DL-Dithiothreitol (DTT) (Sigma-Aldrich, catalog number: D0632 )
- Ethylenediaminetetraacetic acid disodium salt dihydrate (EDTA) (Sigma-Aldrich, catalog number: E5134 )
- Sodium hydroxide pellets (Panreac Applichem, catalog number: 131687 )
- Maltotriose (Sigma-Aldrich, catalog number: M8378 )
- Adenosine 5’ diphospho, [D-glucose-14C(U)] ammonium salt (ADP [U-14C] glucose) (American Radiolabeled Chemicals, catalog number: ARC 3297 )
- Methanol (CARLO ERBA Reagents, catalog number: 412532 )
- Potassium chloride (AppliChem GmbH, catalog number: A2939 )
- Ecolite(+)TM liquid scintillation cocktail (MP Biomedicals, catalog number: 882475 )
- 1 M HEPES-KOH (see Recipes)
- 1 M Tricine (see Recipes)
- 0.25 M Potassium acetate (see Recipes)
- 1 M DTT (see Recipes)
- 0.5 M EDTA (see Recipes)
- 0.1 M Maltotriose (see Recipes)
- 3 M KCl (see Recipes)
- 20 mg/ml Glycogen solution (see Recipes)
- Extraction buffer (see Recipes)
- Percoll buffer (see Recipes)
- Washing buffer (see Recipes)
- Precipitation buffer (see Recipes)
Equipment
- Small mortar and pestle
- 1.5 ml microfuge tubes
- Automatic pipettes
- Precision scale
- Centrifuge (Eppendorf, model: 5810 R )
- Microcentrifuge (Eppendorf, model: 5424 )
- Scintillation Counter (Beckman Coulter, model: LS 6000 IC )
- Scintillation vials (Sigma-Aldrich, catalog number: Z376817 )
Procedure
- Starch granules isolation
Starch granules were purified following a modification of the Percoll method described by Tenorio et al. (2003).- Homogenize 600 mg (fresh weight) of Arabidopsis leaves in a mortar and pestle in the presence of liquid N2 to a fine powder and add 1 ml of extraction buffer. Leaf material was collected from fully developed rosette leaves from at least three different plants at growth stage 3.50 [according to Boyes et al. (2001)].
- Filter the homogenate through two layers of Miracloth on 15 ml Falcon-type tubes.
- Add 5 ml of Percoll buffer (4 °C) to the filtrate and mix.
- Centrifuge samples for 5 min, 805 x g at 4 °C (2,000 rpm, Eppendorf-5810 R centrifuge) and discard the supernatant.
- Wash the pellet by resuspension in 5 ml washing buffer (4 °C), centrifuge as in step A4 and discard the supernatant. Repeat three times.
- Air-dry the pellet (do not let it dry completely) containing the starch granules (Figure 1) and use it immediately for determination of GBSS activity as described below.
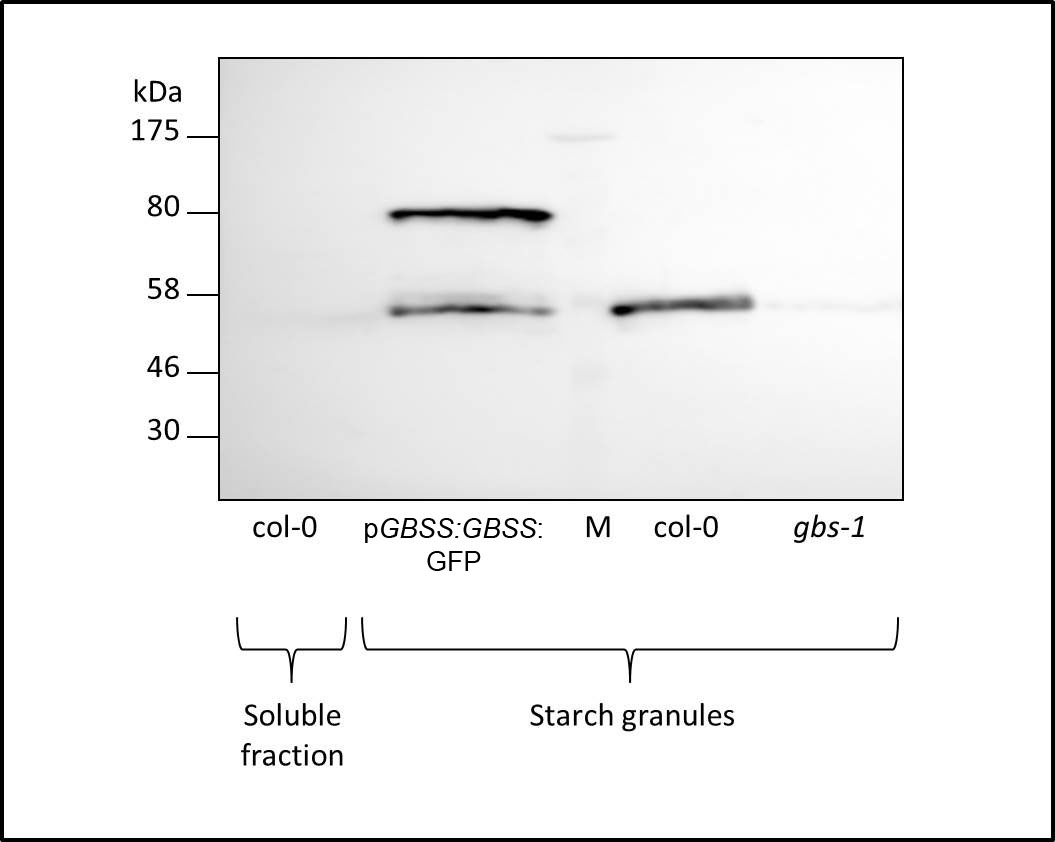

Figure 1. Immunodetection of GBSS in starch granules isolated from Arabidopsis Col-0 leaves. Western blot employing GBSS‐specific antibodies in protein extracts from starch granules isolated from wild‐type (Col-0), GBSS mutant (gbs-1) and a recombinant line in which GBSS is fused to GFP. Molecular Mass (MM) markers are shown in kDa (left). Notice GBSS MM around 55 kDa and GBSS-GFP around 80 kDa. GBSS detection is strictly associated to starch granules fractions (three rightmost lanes) and cannot be detected in a soluble protein fraction (leftmost lane).
- Homogenize 600 mg (fresh weight) of Arabidopsis leaves in a mortar and pestle in the presence of liquid N2 to a fine powder and add 1 ml of extraction buffer. Leaf material was collected from fully developed rosette leaves from at least three different plants at growth stage 3.50 [according to Boyes et al. (2001)].
- GBSS activity assay
The granule-bound starch synthase activity was immediately measured from freshly purified starch granules.- Starch granules were resuspended in 200 μl of 100 mM Tricine (pH 8.4), 25 mM potassium acetate, 10 mM DTT, 5 mM EDTA and 10 mM ADP [U-14C] glucose (specific activity 7.4 GBq/mol).
- 100 µl aliquot was quickly extracted and boiled for 5 min to represent DPMtotal of the assay.
- The rest of the reaction was incubated at 30 °C for 20-60 min.
- The reaction was stopped by boiling for 5 min.
- Starch granules were precipitated by adding 1.875 ml of precipitation buffer and 25 µl of glycogen solution as an inert carrier to increase recovery from alcohol precipitation.
- Tubes were centrifuged 3 min, 2,000 x g at 4 °C (4,615 rpm, Eppendorf-5424) and the supernatant discarded.
- The pellet was resuspended in 0.2 ml deionized water and washed by an additional precipitation with 1.750 ml precipitation buffer without glycogen.
- Tubes were centrifuged as in step B5. The pellet was air-dried for a short time, resuspended in 1 ml of deionized water and transferred into an appropriate scintillation vial.
- 5 ml of EcoLite liquid scintillation cocktail was added to enhance 14C counting efficiency.
- Finally, the radioactivity incorporated into the starch granules was determined with a scintillation counter.
- GBSS activity (see Table 1 and Figure 2) is calculated by using the following formula:
Activity (nmol/min/gfw)=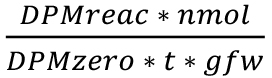
DPMreac = disintegration per minute of the reaction sample minus DPM blank (Table 1)
nmol = nmoles of ADP [U-14C]glucose (1,000 nmol according to the protocol)
DPMtotal = total DPM of added ADP[U-14C]glucose (step B2)
t = reaction time (min) (20-60 min according to the protocol)
gfw = fresh weight (g) (0.3 g according to the protocol)
Table 1. Determination of GBSS activity in starch granules from wild type (Col-0) and gbs-1 mutantPlant Blanka
(DPM)Sampleb
(DPM)DPMtotalc
(DPM)DPMreacd
(DPM)gfw
(g)Reaction time
(min)nmol ADP-gluc
(nmol)Activity
(nmol/min/gfw)Col-0 178.44 2,641.02 57,645.40 2,462.58 0.332 20 1,000 6.43 gbs-1 193.92 197.94 57,645.40 4.02 0.266 20 1,000 0.01 - DPM for the blank sample. Blank sample is identical to each sample except that it was stopped at time 0, so that the reaction was not allowed to start.
- To simplify, a single sample measurement is shown in Table 1. We recommend calculating data from three independent experiments. Sample DPM correspond to the Disintegration Per Minute at the desired reaction time for each sample (experimental data).
- DPMtotal is the total amount of DPM provided by the radiolabelled substrate. It actually represents the maximum of DPM for each sample. According to the above example it consists on the total DPM given by 1,000 nmol of ADP[U-14C]glucose determined for aliquots from step B2.
- DPMreac = DPMsample minus DPMblank
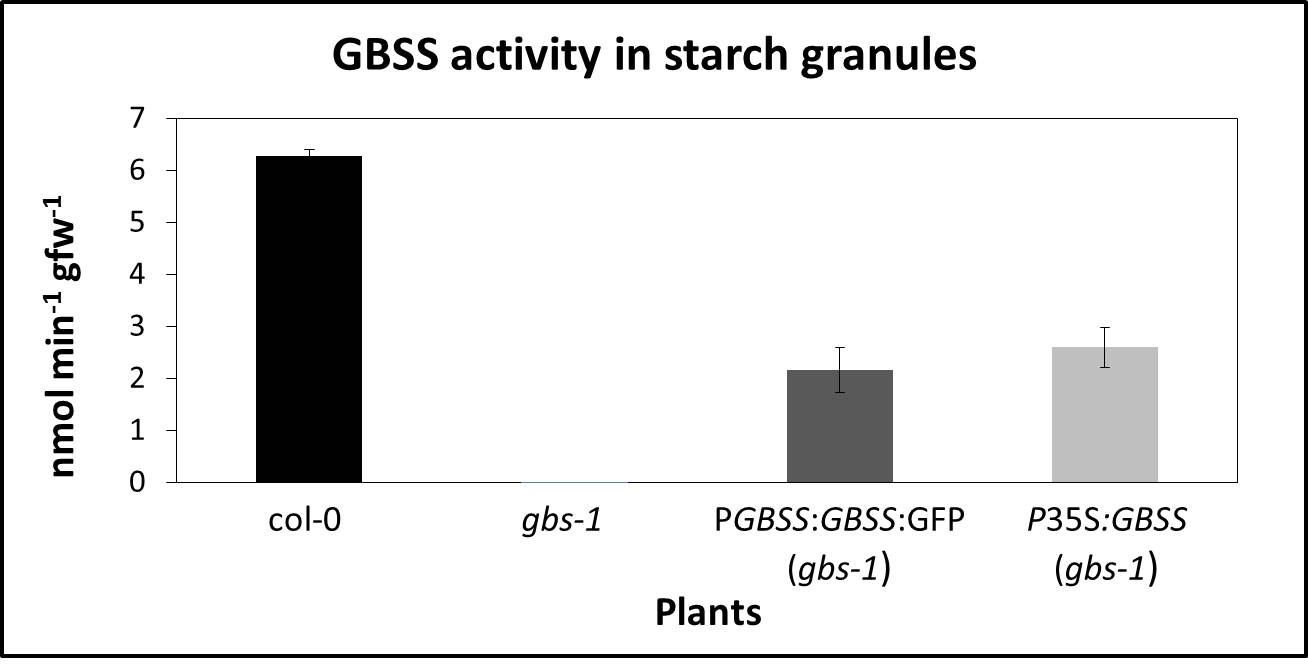
Figure 2. GBSS activity in starch granules from Arabidopsis thaliana leaves. GBSS activity (nmol/min/gfw) in starch granules from wild‐type (Col-0), GBSS mutant (gbs-1), GBSS fused to GFP under its own promoter (pGBSS:GBSS:GFP) in gbs-1 background and GBSS expressed under the 35S promoter (P35S:GBSS) in gbs-1 background lines. Values are the average of three independent experiments. Bars indicate the standard deviation (±SEM). Starch granules isolation and GBSS activity was carried out as described in text.
- DPM for the blank sample. Blank sample is identical to each sample except that it was stopped at time 0, so that the reaction was not allowed to start.
- Starch granules were resuspended in 200 μl of 100 mM Tricine (pH 8.4), 25 mM potassium acetate, 10 mM DTT, 5 mM EDTA and 10 mM ADP [U-14C] glucose (specific activity 7.4 GBq/mol).
Representative data
- As shown in Figure 1, a GBSS activity of ca. 6 nmol/min/gfw is expected in starch granules isolated from A. thaliana ecotype col-0 leaves.
- Relative SD (expressed as percentage of the mean) fell in the range of 2-14 per cent for samples from three independent experiments.
Notes
- Use dischargeable lab ware and appropriate radioactive secure installations and equipment while manipulating ADP [U-14C] glucose.
Recipes
Note: The given volumes are sufficient for 50 reactions.
- 1 M HEPES-KOH (pH 7.5)
Mix 59.58 g HEPES with 175 ml deionized water
Adjust pH to 7.5 with KOH pellets
Add deionized water to 250 ml
Sterilize through 0.2 µm pore diameter filters
Stored at 4 °C - 1 M Tricine (pH 8.4)
Mix 4.48 g Tricine with 20 ml deionized water
Adjust pH to 8.4 with KOH
Add deionized water to 25 ml
Autoclave
Stored at 4 °C - 0.25 M Potassium acetate
Mix 1.23 g Potassium acetate with 40 ml deionized water
Sterilize through 0.2 µm pore diameter filters
Stored at Room Temperature (RT) - 1 M DTT
Weigh 1.54 g DTT
Add deionized water to 10 ml
Distribute in aliquots and store at -20 °C - 0.5 M EDTA (pH 8.0)
Mix 18.61 g EDTA (disodium, dihydrate) with 80 ml deionized water
Adjust to pH 8.0 with NaOH
Note: It is not easy to dissolve the EDTA. It will not dissolve completely until the pH is around 8.0.
Autoclave (15 psi, 1-2 h, 120 °C)
Stored at RT - 0.1 M Maltotriose
Weigh 0.2655 g maltotriose
Add deionized water to 5 ml
Sterilize through 0.2 µm pore diameter filters
Distribute in aliquots and store at -20 °C - 3 M KCl
Weigh 2.237 g of KCl
Add deionized water to 10 ml
Autoclave (15 psi, 1-2 h, 120 °C)
Stored at RT - 20 mg/ml Glycogen solution
Weigh 25 mg of glycogen
Add deionized water to 1.25 ml
Stored at -20 °C - Extraction buffer
Mix 2.5 ml 1 M HEPES-KOH (pH 7.5) with 0.5 ml Triton X-100
Add deionized water to 50 ml
Stored at RT - Percoll buffer
Mix 12.5 ml 1 M HEPES-KOH (pH 7.5) with 125 ml Percoll
Add deionized water to 250 ml - Washing buffer
37.5 ml of 1 M HEPES-KOH (pH 7.5)
Add deionized water to 750 ml - Precipitation buffer
150 ml Methanol
Add 9.4 ml 3 M KCl
Add deionized water to 200 ml
Stored at RT
Acknowledgments
This work was performed with funding from projects CSD2007-00057, BIO2008-02292, and BIO2011-28847-C02-00 (Spanish Ministry of Economy and Competitiveness, MINECO) and Excellence projects P06-CVI-01450 and P08-AGR-03582 (Junta de Andalucía) partially supported by FEDER funding to F. V. and J. M. R. We also acknowledge the TRANSPLANTA consortium Project CONSOLIDER 28317 (MINECO).
References
- Boyes, D. C., Zayed, A. M., Ascenzi, R., McCaskill, A. J., Hoffman, N. E., Davis, K. R. and Gorlach, J. (2001). Growth stage-based phenotypic analysis of Arabidopsis: a model for high throughput functional genomics in plants. Plant Cell 13(7): 1499-1510.
- Ortiz-Marchena, M. I., Albi, T., Lucas-Reina, E., Said, F. E., Romero-Campero, F. J., Cano, B., Ruiz, M. T., Romero, J. M. and Valverde, F. (2014). Photoperiodic control of carbon distribution during the floral transition in Arabidopsis. Plant Cell 26(2): 565-584.
- Tenorio, G., Orea, A., Romero, J. M. and Mérida, A. (2003). Oscillation of mRNA level and activity of granule-bound starch synthase I in Arabidopsis leaves during the day/night cycle. Plant Mol Biol 51(6): 949-958.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Albi, T., Ortiz-Marchena, M. I., Ruiz, M. T., Romero, J. M. and Valverde, F. (2014). Purification of Starch Granules from Arabidopsis Leaves and Determination of Granule-Bound Starch Synthase Activity. Bio-protocol 4(23): e1316. DOI: 10.21769/BioProtoc.1316.
Category
Plant Science > Plant metabolism > Carbohydrate
Plant Science > Plant biochemistry > Protein > Activity
Biochemistry > Carbohydrate > Polysaccharide
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









