- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Semi-denaturing Detergent Agarose Gel Electrophoresis (SDD-AGE)
Published: Vol 4, Iss 22, Nov 20, 2014 DOI: 10.21769/BioProtoc.1297 Views: 25053
Reviewed by: Fanglian HeAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

From Llama to Nanobody: A Streamlined Workflow for the Generation of Functionalised VHHs
Lauren E.-A. Eyssen [...] Raymond J. Owens
Mar 20, 2024 6244 Views
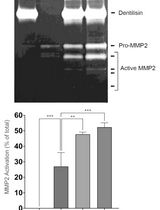
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2095 Views
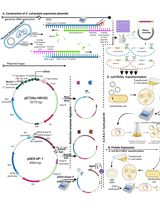
Thermus thermophilus CRISPR Cas6 Heterologous Expression and Purification
Junwei Wei [...] Yingjun Li
Jul 20, 2025 2171 Views
Abstract
Pathological proteins in neurodegenerative diseases suffer a conformational change to a misfolded amyloid state. Such pathological event leads to the aggregation of these proteins that indefinitely propagates as an altered form of itself, and harbor prion-like properties (Wickner, 1994; Prusiner, 2012). In addition to diseases, prions can also have beneficial adaptive roles in lower eukaryotes (in fungi and yeast) (Eaglestone et al., 1999; True et al., 2004; Coustou et al., 1999). Besides separating polymers from their precursor soluble monomers, another particular difficulty of the study of amyloid proteins is to resolve the heterogeneity of the aggregates, since these usually exhibit a variable degree of polymorphism. Semi-denaturating detergent agarose gel electrophoresis (SDD-AGE) is a technique that takes advantage of both the property of prions and prion-like polymers to be highly resistant to solubilization by SDS detergent, and the large pores sizes of agarose, that allow the resolution of high molecular weight complexes. In this method, we describe in detail how this technique can be used to characterize heterogeneous aggregation in bacteria and yeast (Gasset-Rosa et al., 2014; Molina-García and Giraldo, 2014), and further be applied to study the aggregation pattern of proteins that become prone to aggregation through genetic manipulation.
Materials and Reagents
- Bacteria: Gram negative Escherichia coli (E. coli) or yeast Saccharomyces cerevisiae (S. cerevisiae)
Notes:
Amyloids exist in E. coli and S. cerevisiae. In both organism amyloids can be functional. In E. coli the most well know example is the curli protein which is assembled in the extracellular medium and allow the attechemnet of the bacteria to the surface and sustain the formation of biofilms (Chiti and Dobson, 2006; Chapman et al., 2002).
In S. cerevisiae, amyloids confer to cells selctive advantage under certain physiological conditions and they aren’t harmful (Chien et al., 2004; Shorter and Lindquist, 2005; Wickner et al., 2007), a few examples are Sup35p y Ure2p, Rnq1p. The conversion of Sup35 from soluble to the amyloid state drives the reduction of translation termination activity (Wickner et al., 1995). - Protease inhibitor cocktail (Roche Diagnostics, catalog number: 04 693 159 001 )
- Silica beads (1 mm lysing matrix C) (MP Biomedicals, catalog number: 6912-050 )
- Pure agarose D2 (Pronadisa, catalog number: 8034 )
- Detergents: SDS (Bio-Rad Laboratories, catalog number: 161-0301 ) and Sarkosyl 97% N-lauroylsarcosine (Sigma-Aldrich, catalog number: L-9150 )
- PVDF membrane (Bio-Rad Laboratories)
- Antibodies
Notes:
Depends on the protein you work, the antibody may be against the protein that you are studying for aggregation. A recommended strategy if the user is going to generate a recombinant protein is to include a tag in the amino or carboxyl terminal of the protein and use a specific antibody against the tag. One example is the Histidine-tag, the corresponding antibody would be a Monoclonal anti-polyhistidine (Sigma-Aldrich, catalog number: H1029 ).
Antibodies that recognize monomer state may still be able to identify the protein in polymers even is the protein conformation has changed. Usually monomer antibodies have a broad range recognition epitopes. In our hands the antibodies used were:
For E. coli, house-made polyclonal antibodies against RepA (generated in rabbit immunized with soluble RepA), (Gasset-Rosa et al., 2014); and monoclonal anti poly-histidine against His-RepA (WH1) (Sigma-Aldrich, catalog number: H1029) (Molina-García and Giraldo, 2014). Both of them work.
For S. cerevisiae, anti-HA antibody (Roche Diagnostics, catalog number: 12CA5 ) against the HA (Human influenza hemagglutinin) tag of Sup35 protein (unpublished). - Cell culture medium
- Luria Bertani (LB) medium (see Recipes)
- Yeast extract peptone dextrose (YPD) (see Recipes)
- Luria Bertani (LB) medium (see Recipes)
- Lysis buffer (see Recipes)
- Running buffer (see Recipes)
- Loading buffer (see Recipes)
- Transference buffer (see Recipes)
Equipment
- Fast Prep-MilliPore/MP FastPrep-24 homogenizer (catalog number: 6004-500 )
- Horizontal agarose electrophoresis system (Electrophoresis power supply EPS601 General Electrics)
- Wet/Tank transfer blotting system (Bio-Rad Laboratories)
Procedure
- Growth cells in your adjusted conditions.
- Harvest cells.
- Bacteria: Harvest cells (25 ml O.D. = 2) by centrifugation (10 min, 3,500 rpm, 4 °C). And, resuspend the cells in 400 μl of lysis buffer.
- Yeast: Harvest cells (200 ml O.D. = 0.8) by centrifugation (5 min, 3,500 rpm, 4 °C). And, resuspend the cells in 500 μl of lysis buffer.
- Bacteria: Harvest cells (25 ml O.D. = 2) by centrifugation (10 min, 3,500 rpm, 4 °C). And, resuspend the cells in 400 μl of lysis buffer.
- Transfer the suspension to a tube containing silica beads (usually 1/3 of the volume is sufficient).
- Lysate the cells.
- Bacteria: Lysate the cells using the Fast-Prep 24 homogenizer (4 cycles at speed IV, 30 sec each at 4 °C). And, dilute 3 μl of cell lysate with 30 μl with lysis buffer, and then add 10 μl of 4x loading buffer to the sample.
- Yeast: Lysate the cells using the Fast-Prep 24 homogenizer (5 cycles at speed V, 30 sec each at 4 °C). And, add 10 μl of 4x loading buffer to 30 μl of the lysate.
- Bacteria: Lysate the cells using the Fast-Prep 24 homogenizer (4 cycles at speed IV, 30 sec each at 4 °C). And, dilute 3 μl of cell lysate with 30 μl with lysis buffer, and then add 10 μl of 4x loading buffer to the sample.
- Incubate the samples 10 min at room temperature.
- Load them in the agarose gel.
- Prepare 200 ml of agarose at 1.5% in 1x TAE.
Note: Heat the agarose suspension to melting and then add SDS to 0.1%. - Pour it into 20 cm x 24 cm horizontal plate mould. Make sure that the casting tray is on a flat surface.
Notes:- Eliminate all the possible bubbles.
- Agarose electrophoresis are run using horizontal agarose electrophoresis system, the samples need to be run for long distance in order to resolve polymers of high molecular weight.
- Eliminate all the possible bubbles.
- Fit a thick comb (e.g., >2 mm, 20-well) allowing to load 40 μl per sample.
- Submerge the gel in a horizontal electrophoresis container with 1x TAE-0.1% SDS at 10 °C.
- Prepare 200 ml of agarose at 1.5% in 1x TAE.
- Run electrophoresis at 100 V for 7.5 h at 10 °C.
Note: For detecting oligomers with high hydrophobic feature run it at lower voltage 50 V and room temperature for 12 h with 1% (final concentration) of Sarkosyl in the sample (4% Sarkosyl in the loading buffer). The concentration for the running buffer is TAE 1x and 0.1% SDS as described before.
Caution: Extreme care during electrophoresis to avoid any electrical hazard. - Cut the PVDF membrane adjusted to the size of the agarose gel and 2 pieces with the same dimensions of Whatman paper.
- Attach tightly the gel to the PDVF membrane and add in each side a piece of Whatman paper.
Note: Everything should be pre-wet with transference buffer. Make sure that there are no bubbles in between. - Place the cast into the Wet Transfer-Blot and cover with buffer 1x TAE-0.1% SDS.
Caution: Extreme care during electrophoresis to avoid any electrical hazard. - Run the transference at 16 V/400 mA and 10 °C for 15 h.
- Perform Western-blot with the appropriate antibodies to detect your polymers.
Recipes
- Luria Bertani (LB) medium
Bacto-tryptone 10 g/L
Yeast extract 5 g/L
NaCl 5 g/L - Yeast extract peptone dextrose (YPD)
2% bacto peptone
2% glucose
1% yeast extract - Lysis buffer
25 mM Tris-HCl (pH 6.8)
250 mM NaCl
5 mM EDTA
10% glycerol supplemented with protease inhibitors (1 tablet each 10 ml) - Running buffer
1x TAE (Tris acetate EDTA)
0.1% SDS - Loading buffer (4x)
0.5% TAE
5% glycerol
2% sarkosyl
0.1 mg/ml bromophenol blue plus protease inhibitors (1 tablet each 10 ml) - Transference buffer
1x TAE (Tris Acetate EDTA)
0.1% SDS
Acknowledgments
We thank Prof. Rafael Giraldo for his encouragement and continuous support, and Spanish MINECO for funding (BIO2012-30852 and CSD2009-00088).
References
- Coustou, V., Deleu, C., Saupe, S. J. and Begueret, J. (1999). Mutational analysis of the [Het-s] prion analog of Podospora anserina. A short N-terminal peptide allows prion propagation. Genetics 153(4): 1629-1640.
- Eaglestone, S. S., Cox, B. S. and Tuite, M. F. (1999). Translation termination efficiency can be regulated in Saccharomyces cerevisiae by environmental stress through a prion‐mediated mechanism. EMBO J 18(7): 1974-1981.
- Gasset-Rosa, F., Coquel, A. S., Moreno-Del Alamo, M., Chen, P., Song, X., Serrano, A. M., Fernandez-Tresguerres, M. E., Moreno-Diaz de la Espina, S., Lindner, A. B. and Giraldo, R. (2014). Direct assessment in bacteria of prionoid propagation and phenotype selection by Hsp70 chaperone. Mol Microbiol 91(6): 1070-1087.
- Molina-García, L. and Giraldo, R. (2014). Aggregation interplay between variants of the RepA-WH1 prionoid in Escherichia coli. J Bacteriol 01527-01514.
- Prusiner, S. B. (2012). Cell biology. A unifying role for prions in neurodegenerative diseases. Science 336(6088): 1511-1513.
- True, H. L., Berlin, I. and Lindquist, S. L. (2004). Epigenetic regulation of translation reveals hidden genetic variation to produce complex traits. Nature 431(7005): 184-187.
- Wickner, R. B. (1994). [URE3] as an altered URE2 protein: evidence for a prion analog in Saccharomyces cerevisiae. Science 264(5158): 566-569.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Molina-García, L. and Gasset-Rosa, F. (2014). Semi-denaturing Detergent Agarose Gel Electrophoresis (SDD-AGE). Bio-protocol 4(22): e1297. DOI: 10.21769/BioProtoc.1297.
Category
Microbiology > Microbial biochemistry > Protein > Isolation and purification
Microbiology > Microbial biochemistry > Protein > Structure
Biochemistry > Protein > Electrophoresis
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










