- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Fractionation by Ultracentrifugation of Gram Negative Cytoplasmic and Membrane Proteins
Published: Vol 4, Iss 21, Nov 5, 2014 DOI: 10.21769/BioProtoc.1287 Views: 20612
Reviewed by: Fanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

From Llama to Nanobody: A Streamlined Workflow for the Generation of Functionalised VHHs
Lauren E.-A. Eyssen [...] Raymond J. Owens
Mar 20, 2024 6247 Views
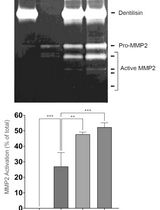
Purification of Native Dentilisin Complex from Treponema denticola by Preparative Continuous Polyacrylamide Gel Electrophoresis and Functional Analysis by Gelatin Zymography
Pachiyappan Kamarajan [...] Yvonne L. Kapila
Apr 5, 2024 2097 Views
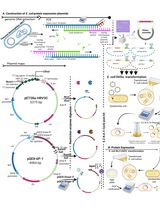
Thermus thermophilus CRISPR Cas6 Heterologous Expression and Purification
Junwei Wei [...] Yingjun Li
Jul 20, 2025 2171 Views
Abstract
Protein fractionation is a useful separation process which divides membrane proteins (including those located in the outer and inner membrane) and cytoplasmic proteins into discrete fractions. Fractionation of proteins can simplify analysis of the numbers of proteins present, and therefore make easier to characterize any environmentally or mutation induced changes in expression profiles, or changes in protein strucutre resulting from post-translational modification. This protocol is derived from Haigh et al. (2013) and it is specific to Gram negative bacteria.
Materials and Reagents
- Bacterial culture media (subject to the needs of the bacterium being studied; for Escherichia coli, Luria Broth or DMEM are suitable)
- Tris base (Thermo Fisher Scientific)
- Triton X-100 (Sigma-Aldrich) (optional)
- Tris-Triton (TT)
- HCl (Hydrochloric acid)
- TT (see Recipes)
Equipment
- Bench top ultracentrifuge (e.g. Beckman Coulter, model: TL-100 ) capable of up to 50,000 rpm for culture volumes of 25 ml or less, and a larger mainframe ultracentrifuge for larger culture volume preparations
- Bench top centrifuge capable of up to 12,225 x g
- Sonicator
- Spectrophotometer
Procedure
- The bacteria should be cultured in media appropriate for the bacterial species under investigation or the environmental condition being tested. Cultures can be at logarithmic or stationary phase, although stationary phase cultures can be harder to lyse for protein extraction.
- Harvest the bacteria by centrifugation at 6,708 x g for 10 min at 4 °C, discard the culture supernatant (unless it is needed to analyse secreted protein profiles).
- Wash the pellet by re-suspending it uniformly with buffer to a volume at 50 times that of the pellet (e.g. the pellet from a 200 ml stationary phase E. coli culture would be around 2 ml in volume and would need at least 100 ml of buffer), then re-centrifuge at 6,708 x g for 10 min at 4 °C. Repeat this wash step.
- For separation of E. coli membrane proteins the pellet should be washed in 10 mM Tris base (pH 7.5) (this buffer can be used with any proteins from any cellular source). For small volume culture protein fractionations (e.g. 25 ml) an overnight growth will give an OD600 of approximately 2; once harvested and washed, this would be re-suspended in 0.5-1 ml 10 mM Tris base (pH 7.5), to give a final cell density of OD600 of 20-30 (the suspension should have the appearance of pouring cream). For large volume culture (e.g. 200 ml) re-suspend in 5-10 ml 10 mM Tris base (pH 7.5). If proteolysis is likely to be a problem, a cocktail of broad spectrum protease inhibitors should be included at the levels recommended by the manufacturer.
- Freeze the culture suspension at -80 °C for 2 h or overnight to weaken the cell wall and increase the lysis of the bacterial cells by sonication.
- Thaw the bacterial suspension on ice and place it in a non-breakable plastic beaker sitting in an ice water bath. Set the sonicator between 6-8 microns and lyse the cells in cycles of 15 sec sonication followed by 45 sec cooling until the lysate changes from an opaque solution into a less turbid solution (usually 4-5 cycles are enough for log phase cells; stationary phase cultures may need more treatments).
- Centrifuge the lysed sample at 6,708 x g for 10 min at 4 °C to remove large debris fragments and unlysed cells (these will be contained in the pellet) and transfer the supernatant containing the total protein extract (membrane and cytoplasmic) into appropriate ultracentrifuge tubes.
- Ultracentrifuge the protein extract at 108,726 x g for 10 min at 4 °C to separate membrane proteins from cytoplasmic proteins.
- After the centrifugation the supernatant will contain the cytoplasmic proteins while the pellet contains the total membrane protein. If only a total membrane proteins fraction is required (comprising both inner and outer membrane proteins) then go to step 11, if only the outer membrane and inner membrane proteins fractions are required go to step 11 and then 10.
- The pellet consisting of total membrane proteins is re-suspended in 10 mM Tris-HCl (pH 7.5) supplemented with 2% Triton X-100 in 10 mM Tris-HCl (pH 7.5) (TT; e.g. for a small volume culture initially of 25 ml re-suspend in 100-200 μl of TT); the inner membrane proteins are solubilised by incubation at room temperature with occasional mixing for 30 min. The mixture is then again centrifuged at 108,726 x g for 10 min at 4 °C; the supernatant will contain the inner membrane protein fraction whereas the insoluble pellet will contain the outer membrane protein. To remove any residual inner membrane proteins, the outer membrane pellet is re-extracted for a further 30 min in 500 μl of TT (this step is only to purify the outer membrane proteins and so the supernatant should be discarded).
Note: The extra wash/extraction steps are needed to increase the relative purity of the different fractions. Sometime it is possible that there are carry over between the fractions especially if people do not take care to remove all supernatants between centrifugations (but that's is more a procedure error). Generally, we determined that two washes are enough to obtain a pure membrane fraction. We tested the purity of the protein fraction by separating them on SDS gel and subjecting the proteins to sequencing. However, more washes steps can be increased to enhance the purity if required.
- Wash the total membrane pellet twice or outer membrane pellet once in 10 mM Tris-HCl (pH 7.5); for a small volume culture initially of 25 ml, 500 μl of buffer would be sufficient. Finally, re-suspend in 30-50 μl of the same buffer according to the membrane pellet size.
- The concentration of the cytoplasmic and membrane protein samples should be measured using the Bradford assay (He, 2011a) or via a spectrophotometer set at 280 nm.
- For storage, proteins should be contained in capped 1.5 or 2 ml sterile plastic tubes, and stored at -20 °C for around 1 month or -80 °C for 6 months or longer. Protein extracts should be thawed on ice and always kept at 0-4 °C for the minimum amount of time to maximize stability and protein activity.
- For simple 1-dimensional SDS PAGE, normalized protein samples are mixed with adequate amounts of 2x SDS protein sample buffer and the gel electrophoresis carried out as described by He (2011b) (for 1D SDS-gel electrophoresis). The protein fractions can also be separated by 2D gel electrophoresis if required.
Recipes
- TT
20 μl Triton X-100 in 0.98 ml of 10 mM Tris-HCl (pH 7.5)
Acknowledgments
This protocol is derived from Haigh et al. (2013).
References
- He, F. (2011a). Bradford protein assay. Bio-protocol 1(6): e45.
- He, F. (2011b). Laemmli-SDS-PAGE. Bio-protocol 1(11): e80.
- Haigh, R., Kumar, B., Sandrini, S. and Freestone, P. (2013). Mutation design and strain background influence the phenotype of Escherichia coli luxS mutants. Mol Microbiol 88(5): 951-969.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Sandrini, S. M., Haigh, R. and Freestone, P. P. E. (2014). Fractionation by Ultracentrifugation of Gram Negative Cytoplasmic and Membrane Proteins. Bio-protocol 4(21): e1287. DOI: 10.21769/BioProtoc.1287.
Category
Microbiology > Microbial biochemistry > Protein > Isolation and purification
Microbiology > Microbial cell biology > Organelle isolation
Biochemistry > Protein > Isolation and purification
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









