- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Analysis of L- and D-Amino Acids Using UPLC
Published: Vol 4, Iss 17, Sep 5, 2014 DOI: 10.21769/BioProtoc.1231 Views: 13938

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Optimizing Transmembrane Protein Assemblies in Nanodiscs for Structural Studies: A Comprehensive Manual
Fernando Vilela [...] Dorit Hanein
Nov 5, 2024 3096 Views
Protein Structural Characterization Using Electron Transfer Dissociation and Hydrogen Exchange-Mass Spectrometry
Rupam Bhattacharjee and Jayant B. Udgaonkar
Jun 20, 2025 1781 Views
On-Column Dual-Gradient Refolding for Efficient Recovery of Insoluble Affinity-Tagged Recombinant Proteins
Anna Vlaskina [...] Maxim Patrushev
Feb 5, 2026 66 Views
Abstract
With the exception of glycine, α-amino acids are optically active, and two optical isomers (L- and D-) of each amino acid can be formed. Recent developments of analytical techniques have revealed that several free D-amino acids such as D-aspartate, D-serine and D-alanine exist in many kinds of organism including human and have biologically important roles. D-Aspartate regulates reproductive activity in animals and humans. D-Serine serves as a co-agonist of the N-methyl-D-aspartate receptor, which mediates glutamatergic neurotransmission. D-Alanine plays a role like osmolyte in crustaceans and mollusks. In this protocol, we describe a method for analysis of L- and D-amino acids using ultra-performance liquid chromatography (UPLC). To analyze D- and L-amino acids, the enantiomers are initially converted into diastereomers (diastereomers are stereoisomers that are not related as object and mirror image and are not enantiomers) using pre-column derivatization with o-phthaldialdehyde plus N-acylated cysteine (N-acethyl-L-cysteine or N-tert-butyloxycarbonyl-L-cysteine). The resultant derivatives are fluorescent diastereomers. This is followed by separation of the resultant fluorescent isoindol derivatives on an octadecylsilyl stationary phase using UPLC, and the fluorescence is detected by a fluorescence detector included in UPLC system. Using this method, 16 kinds of D-amino acid can be analyzed.
Materials and Reagents
- N-Acetyl-L-cysteine (NAC) (Wako Pure Chemical Industries, catalog number: 017-05131 )
- N-tert-butyloxycarbonyl-L-cysteine (NBC) (Funakoshi, catalog number: 04-621 )
- o-Phthaldialdehyde (OPA) (Nacalai Tesque, catalog number: 27824-74 )
- Methanol (Wako Pure Chemical Industries, catalog number: 132-06471 )
- Acetonitrile (Wako Pure Chemical Industries, catalog number: 015-08633 )
- Trichloroacetic acid (Wako Pure Chemical Industries, catalog number: 203-04952 )
- Sodium hydroxide (Wako Pure Chemical Industries, catalog number: 192-15985 )
- 0.4 M sodium borate buffer (see Recipes)
- 50 mM sodium acetate buffer (see Recipes)
- Acetic acid (Wako Pure Chemical Industries, catalog number: 192-01075 )
- Sodium acetate (Wako Pure Chemical Industries, catalog number: 192-01075)
- Acetic acid (Wako Pure Chemical Industries, catalog number: 192-01075 )
- 50% TCA (see Recipes)
- 1 M NaOH (see Recipes)
Equipment
- ACQUITY UPLC-FLR system (Waters)
- AccQ-Tag Ultra 2.1 x 100 mm column (Waters, catalog number: 186003837 )
- ACQUITY binary solvent manager
- ACQUITY sample manager
- ACQUITY FLR detector
- Sample vial: 12 x 32 mm glass screw neck vial, Quick Thread, LectraBond cap, PTFE/silicone septa, Total Recovery (Waters, catalog number: 186000384c )
- Empower 2 software
- AccQ-Tag Ultra 2.1 x 100 mm column (Waters, catalog number: 186003837 )
- Vortex (Taitec, catalog number: 0061271-000 )
- Polytetrafluoroethylene membrane filters (4-mm diameter, 0.2-μm pore size) (Merck KGaA, catalog number: SLLGH04NL )
- Amicon Ulta 0.5 ml centrifugal filter 3 K device (Merck KGaA, catalog number: UFC500396 )
Software
- Empower 2 software
Procedure
- D, L-Amino acids are bound with OPA, and NAC (Aswad, 1984) or NBC (Hashimoto et al., 1992) using two methanolic solutions (A and B) containing the derivatizing reagents (see Notes 1-3). This derivatization results in fluorescent diastereomers. Methanolic solution A is prepared by dissolving 8 mg of OPA and 10 mg of NAC in 1 ml of methanol, while methanolic solution B is prepared by dissolving 10 mg of OPA and 10 mg of NBC in 1 ml of methanol. These methanolic solution A and B are not light sensitive.
- The reaction mixture (250 μl) for the derivatization contains 25 μl of amino acid sample, 50 μl of methanolic solution (A or B) and 175 μl of 0.4 M sodium borate buffer (pH 10.4) in a sample vial. Vortex the reaction mixture. To choose the correct methanolic solution for your sample, see Notes 2 and 3.
- After fluorescence derivatization for 2 min at room temperature in the dark, set the sample vial on vial holder in ACQUITY sample manager of UPLC system. Introduction of an aliquot (1 μl) of the reaction mixture into a UPLC system is performed automatically. Because the fluorescence derivatives are unstable, the introduction of an aliquot into a UPLC system should be performed immediately after the incubation.
- The system is operated at a flow rate of 0.25 ml/min at 30 °C. The UPLC gradient system for analysis of OPA-NAC derivatives [A = 50 mM sodium acetate (pH 5.9) and B = methanol] is 10-20% B for 3.2 min, 20% B for 1 min, 20-40% B for 3.6 min, 40% B for 1.2 min, 40-60% B for 3.8 min, 60% B for 1 min and 60-10% B for 0.01 min, and then the system is equilibrated with 90% A and 10% B for 4.19 min. About 4.0 ml of A and about 2.7 ml of B are needed for the analysis of one sample. The gradient system for analysis of OPA-NBC derivatives [A = 50 mM sodium acetate (pH 5.9) and B = acetonitrile] is 15-21% B for 7 min, 21-27.5% B for 1.5 min, 27.5% B for 2 min, 27.5-30% B for 1 min, 30-40% B for 2 min, 40% B for 0.5 min and 40-15% B for 0.01 min, and then the system is equilibrated with 85% A and 15% B for 3.99 min. About 3.8 ml of A and about 1.8 ml of B are needed for the analysis of one sample. The excitation and emission wavelengths for fluorescent detection of the diastereoisomeric amino acid derivatives are 350 nm and 450 nm, respectively. Before the first analysis, the system is pre-equilibrated with the equilibration condition described above for 16 min.
- The data is processed using Empower 2 that is control and analysis software for a UPLC system. The peak heights and retention times are used for amino acid quantification and identification, respectively (see Note 4). In the case of analysis for biological sample, identification of amino acid is performed based on retention times of amino acids provided from analysis of a control amino acid sample run alongside.
- The standard solution for calibration curves consists of 4 mM 32 kinds of amino acids: D- and L-forms of aspartate (Asp), glutamtamate (Glu), serine (Ser), alanine (Ala), leucine (Leu), phenylalanine (Phe), valine (Val), methionine (Met), tryptophan (Trp), arginine (Arg), histidine (His), threonine (Thr), asparagine (Asn), tyrosine (Tyr) and glutamine (Gln), plus L-isoleucine (L-Ile) and D-allo-isoleucine (D-allo-Ile). The standard solution was diluted to 7 concentrations (5, 25, 50, 100, 250, 500 and 1,000 μM) in 0.05 M HCl, after which the solutions were filtered through polytetrafluoroethylene membrane filters (4-mm diameter, 0.2-μm pore size). Based on chromatograms provided from UPLC analysis of each concentration of the standard solution, calibration curves of D or L-amino acid are calculated plotting peak height versus content of amino acid. The calculation of equation of a calibration curve and quantification of amino acid in your sample are performed using Empower 2 software.
Representative data
Instability of the column temperature negatively affects reproducibility of separation of two neighboring peaks (for example, peaks of D-histidine and L-histidine in the case of OPA-NAC derivatization). We recommend that you keep the column temperature at 30 °C accurately.
Notes
- In this method, amino groups in compounds are targets for diastereomeric and fluorescent derivatization, and not only amino acids but also proteins are derivatized with the derivatizing reagents. The derivatized fluorescent proteins cause contaminated peaks, and prevent identification and quantification of amino acids. Therefore, proteins in a sample should be eliminated by filtration using an Amicon Ulta 0.5 ml centrifugal filter 3 K device or trichloroacetic acid (TCA) treatment.
- TCA treatment is performed as described below. Add 125 μl of 50% TCA (see Recipes) to the 500 μl of a sample solution such as biological sample and food sample. After incubation for 5 min at room temperature, the mixture is centrifuged (10,000 x g for 15 min at 4 °C), and an aliquot (500 μl) of the supernatant is neutralized by addition of 300 μl of 1 M NaOH (see Recipes). When samples after TCA treatment are used for UPLC analysis, filter the sample solution through polytetrafluoroethylene membrane filters (4-mm diameter, 0.2-μm pore size).
- Using OPA-NAC derivatization, D- and L-forms of Asp, Ser, Gln, His, Arg, Ala, Trp, Val, Met, Tyr, Phe, Leu, D-Thr, D-allo-Ile and L-Ile can be analyzed at one time (Figure 1A).
- Using OPA-NBC derivatization, D- and L-forms of Glu, Asn, Asp, Ser, Gln, Ala, Tyr, Val, Met, Phe, D-Thr, D-Arg, D-Leu, L-His, L-Ile and L-Trp can be analyzed at one time (Figure 1B).
- In analyses using OPA-NAC and OPA-NBC, when calibration curves were calculated plotting peak height versus content of amino acid (from 0.5 to 100 pmol), the linearity was observed in all cases with regression coefficients above 0.998.
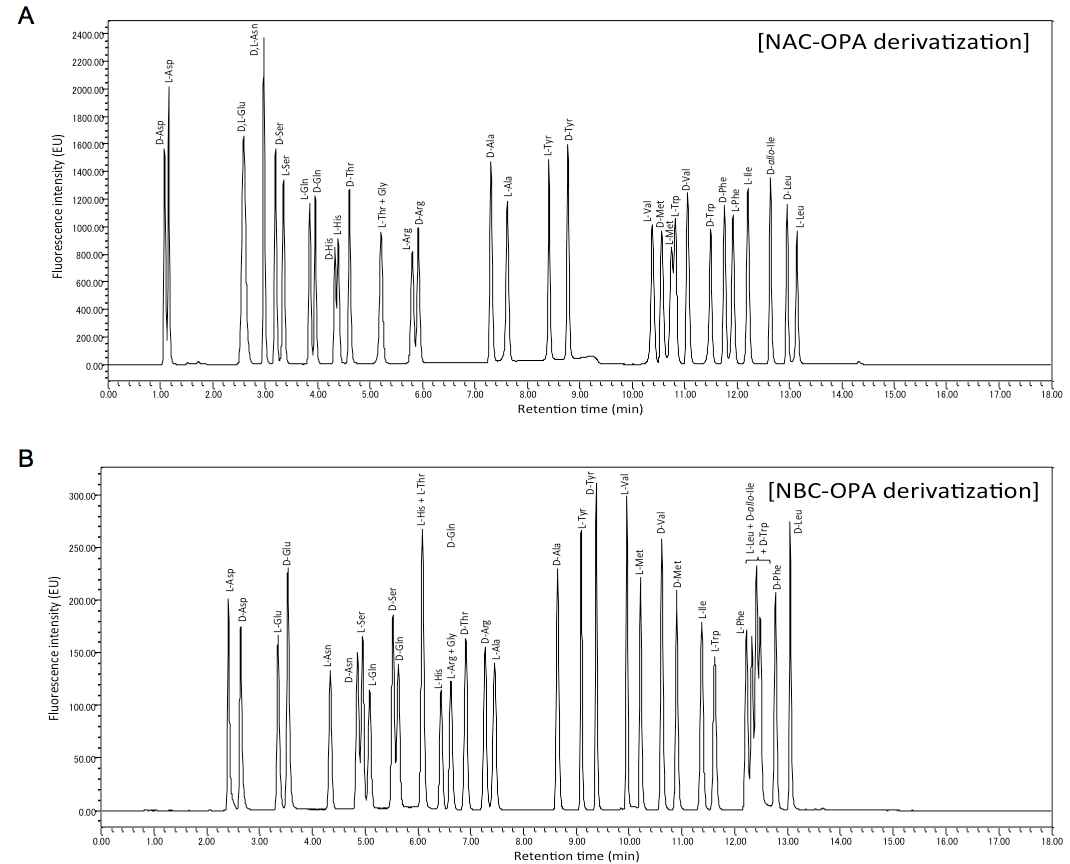
Figure 1. Chromatograms of NAC-OPA A and NBC-OPA B derivatives of free amino acids. A. Each peak corresponds to 100 pmol of amino acid. B. Each peak corresponds to 20 pmol of amino acid.
Recipes
- 0.4 M sodium borate buffer (pH 10.4, 100 ml)
Mix 2.47 g of boric acid with 80 ml of ultra pure water
pH to 10.4 with sodium hydroxide
Add ultra pure water to 100 ml
Filtration (0.22 μm)
Stored at room temperature
- 50 mM sodium acetate buffer (pH 5.9, 1 L)
Mix 4.1 g of sodium acetate with 900 ml of ultra pure water
pH to 5.9 with acetic acid.
Add ultra pure water to 1 L
Filtration (0.22 μm)
Stored at room temperature
- 50% TCA
Mix 25 g of TCA with 30 ml of ultra pure water
Add ultra pure water to 50 ml
Stored at 4 °C
- 1 M NaOH
Mix 4.0 g of sodium hydroxide with 80 ml of ultra pure water
Add ultra pure water to 100 ml
Stored at room temperature
Acknowledgments
In this method, we have modified Aswad’s and Hashimoto’s methods for the derivatization of amino acids using OPA and chiral thiols (NAC and NBC). In addition, this work was supported by a grant for Promotion of Basic Research Activities for Innovate Bioscience from the Bio-oriented Technology Research Advancement Institution (BRAIN) and JSPS KAKENHI Grant Number 2402734.
References
- Aswad, D. W. (1984). Determination of D- and L-aspartate in amino acid mixtures by high-performance liquid chromatography after derivatization with a chiral adduct of o-phthaldialdehyde. Anal Biochem 137(2): 405-409.
- Hashimoto, A., Nishikawa, T., Oka, T., Takahashi, K. and Hayashi, T. (1992). Determination of free amino acid enantiomers in rat brain and serum by high-performance liquid chromatography after derivatization with N-tert.-butyloxycarbonyl-L-cysteine and o-phthaldialdehyde. J Chromatogr 582(1-2): 41-48.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Mutaguchi, Y. and Ohshima, T. (2014). Analysis of L- and D-Amino Acids Using UPLC. Bio-protocol 4(17): e1231. DOI: 10.21769/BioProtoc.1231.
Category
Biochemistry > Protein > Fluorescence
Biochemistry > Protein > Structure
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








