- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Iodine Staining of Escherichia coli Expressing Genes Involved in the Synthesis of Bacterial Glycogen
Published: Vol 4, Iss 17, Sep 5, 2014 DOI: 10.21769/BioProtoc.1224 Views: 12769
Reviewed by: Kanika GeraFanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
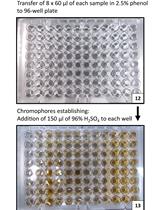

Determination of Storage (Starch/Glycogen) and Total Saccharides Content in Algae and Cyanobacteria by a Phenol-Sulfuric Acid Method
Tomáš Zavřel [...] Jan Červený
Aug 5, 2018 12620 Views
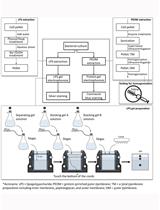
Extraction and Electrophoretic Analysis of Bacterial Lipopolysaccharides and Outer Membrane Proteins
Yue-Jia Lee and Thomas J. Inzana
Dec 20, 2021 5761 Views
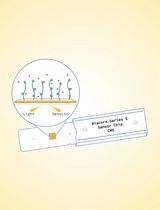
Surface Plasmon Resonance for the Interaction of Capsular Polysaccharide (CPS) With KpACE
Zhe Wang [...] Chao Cai
Jun 20, 2025 3554 Views
Abstract
The presence of intracellular glycogen can be detected by the following iodine staining technique. Cells with glycogen stain dark brown, whereas in its absence they remain with a pale yellowish color. It is hypothesized that iodine atoms fit into helical coils formed by the α-polyglucan to form a coloured glycogen-iodine complex. Here, we have studied the expression of Streptococcus mutans (S. mutans) genes that control the biosynthesis of this polysaccharide (Asencion Diez et al., 2013). Thus, we expressed glgC and glgD genes coding for both ADP-Glc pyrophosphorylase subunits in Escherichia coli (E. coli) AC70R1-504 cells to complement the deficient accumulation of glycogen by this strain (Iglesias et al., 1993). In control cells or in those where an inactive protein was expressed, the synthesis of the polysaccharide was undetectable by this iodine staining technique.
Keywords: Reserve polysaccharideMaterials and Reagents
- Cells: Non-transformed E. coli AC70R1-504 or harboring plasmids with the S. mutans glgC and glgD genes, separately or combined
Note: This strain has a deficient production of the ADP-glucose pyrophosphorylase enzyme in absence of complementary plasmids (Morán-Zorzano et al., 2007).
- Luria-Bertani (LB) liquid medium
- Antibiotics: Kanamycin (US Biological, catalog number: K0010 ) and spectinomycin (Sigma-Aldrich, catalog number: S4014 )
- Inducers: Isopropyl β-D-1-thiogalactopyranoside (IPTG) (Sigma-Aldrich, catalog number: I6758 ) and nalidixic acid (Sigma-Aldrich, catalog number: N4382 )
- Plasmids: pMAB6/glgC, expressing S. mutans glgC (induced by nalidixic acid) and pMAB5/glgD, expressing S. mutans glgD (induced by IPTG)
Note: These plasmids are compatible and bear resistance to spectinomycin and kanamycin, respectively (Asencion Diez et al., 2013).
- D(+) Glucose (Sigma-Aldrich, catalog number: G5767 )
- Iodine crystals (Biopack Medical, catalog number: 2000162300 )
Equipment
- 1.5 ml microcentrifuge tube (Deltalab, catalog number: 200400 )
- Microcentrifuge (Beckman Coulter, model: 22R )
- Shaker (at least 200 rpm) at 37 °C (Thermo Fisher Scientific)
Procedure
- Inoculate non-transformed and transformed E. coli AC70R1-504 cells onto 3 ml of LB medium with appropriate antibiotics to ensure plasmid permanence. Then, grow at 37 °C until an OD600 ~0.8 is reached. Usually the initial inoculate comes from a saturated culture.
Induce protein expression for 3 h at 200 rpm and 25 °C in presence of IPTG, nalidixic acid or both, depending on the genes being expressed.
Note: Temperature, inducer concentration and other conditions are specific for the system in order to assure a correct expression level of soluble and active protein.
- Afterwards, to stimulate glycogen production, add glucose to a final concentration of 0.2% (wt/vol) and further incubate (under the same conditions) for 1 h.
- Withdraw an aliquot of 0.1 ml and centrifuge it in a 1.5 ml microcentrifuge tube at 16,000 x g for 5 min. Aspirate supernatant carefully, leaving a compact cell pellet in the bottom of the tube.
Note: The pellet should be as dried and compact as possible to avoid drops sliding down the walls of the tube.
- Turn upside down the microcentrifuge tube, so an iodine crystal could be placed in the cap of the closed tube. After 5 min at room temperature, iodine vapors stain the cell pellet if glycogen has been previously produced.
Note: The specific amount of iodine in the crystal is not critical, as it only provides an excess of the compound present in vapors.
Representative data

Figure 1. Iodine staining of cells accumulating different amounts of glycogen. Iodine staining of cell pellets from E. coli AC70RI-504 (from left to right): Untransformed cells (lacking ADP-Glc PPase, negative control), transformed to express GlgC (catalytic subunit of ADP-Glc PPase forming homotetrameric enzyme, low positive sample), transformed to express GlgD (non-catalytic, inactive, subunit of ADP-Glc PPase, negative sample), or to express GlgC/GlgD (heterotetrameric ADP-Glc PPase, high positive sample). The staining is mainly qualitative and as much it could be established a semi-quantitative scale based on the visualization by eye (or scanner).
Notes
- The culture conditions and induction of recombinant protein synthesis (temperature, concentration of antibiotics and inducers) should be appropriate for other specific cases. If the growth of different cells varies, the procedure should be adjusted to make sure similar OD are reached to ensure similar metabolic conditions.
- After 5 min, remove iodine crystal to prevent darkening of the walls of the microtube to facilitate the acquisition of the image. The staining may fade always after few hours, but they could be re-stained.
- Before exposure to iodine vapors, the pellet should be as dried and compact as possible to avoid drops sliding down the walls of the tube, which will cause smearing. After aspiration, a small and long piece of paper filter could be used to remove tiny amounts of the remaining liquid.
- Traditionally, this type of staining has been done in plates (Greene et al., 1996). But, one of the advantages of this technique is that it could be used with any type of cells and expression systems, such as BL21 (DE3) E. coli and pET derivative vectors (Ballicora et al., 2007; Kuhn et al., 2010). In those cases, cells cannot be efficiently grown in plates in presence of IPTG.
Acknowledgments
This work was supported by grants to AAI from CONICET [PIP 2519 and CONICET-NSF 19537/28/06/12], UNL [CAI+D Orientado and Redes], and ANPCyT [PICT’08 1754]; and to MAB from the NSF [MCB 1024945]. MDAD is fellow from CONICET; SAG and AAI are investigators from the same institution.
References
- Asencion Diez, M. D., Demonte, A. M., Guerrero, S. A., Ballicora, M. A. and Iglesias, A. A. (2013). The ADP-glucose pyrophosphorylase from Streptococcus mutans provides evidence for the regulation of polysaccharide biosynthesis in Firmicutes. Mol Microbiol 90(5): 1011-1027.
- Ballicora, M. A., Erben, E. D., Yazaki, T., Bertolo, A. L., Demonte, A. M., Schmidt, J. R., Aleanzi, M., Bejar, C. M., Figueroa, C. M., Fusari, C. M., Iglesias, A. A. and Preiss, J. (2007). Identification of regions critically affecting kinetics and allosteric regulation of the Escherichia coli ADP-glucose pyrophosphorylase by modeling and pentapeptide-scanning mutagenesis. J Bacteriol 189(14): 5325-5333.
- Greene, T. W., Chantler, S. E., Kahn, M. L., Barry, G. F., Preiss, J. and Okita, T. W. (1996). Mutagenesis of the potato ADPglucose pyrophosphorylase and characterization of an allosteric mutant defective in 3-phosphoglycerate activation. Proc Natl Acad Sci U S A 93(4): 1509-1513.
- Iglesias, A. A., Barry, G. F., Meyer, C., Bloksberg, L., Nakata, P. A., Greene, T., Laughlin, M. J., Okita, T. W., Kishore, G. M. and Preiss, J. (1993). Expression of the potato tuber ADP-glucose pyrophosphorylase in Escherichia coli. J Biol Chem 268(2): 1081-1086.
- Kuhn, M. L., Figueroa, C. M., Aleanzi, M., Olsen, K. W., Iglesias, A. A. and Ballicora, M. A. (2010). Bi-national and interdisciplinary course in enzyme engineering. Biochem Mol Biol Educ 38(6): 370-379.
- Morán-Zorzano, M. T., Alonso-Casajús, N., Muñoz, F. J., Viale, A. M., Baroja-Fernández, E., Eydallin, G. and Pozueta-Romero, J. (2007). Occurrence of more than one important source of ADPglucose linked to glycogen biosynthesis in Escherichia coli and Salmonella. FEBS Lett 581(23): 4423-4429.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Demonte, A. M., Diez, M. D. A., Guerrero, S. A., Ballicora, M. A. and Iglesias, A. A. (2014). Iodine Staining of Escherichia coli Expressing Genes Involved in the Synthesis of Bacterial Glycogen. Bio-protocol 4(17): e1224. DOI: 10.21769/BioProtoc.1224.
Category
Microbiology > Microbial biochemistry > Carbohydrate
Biochemistry > Carbohydrate > Glycogen
Cell Biology > Cell staining > Carbohydrate
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









