- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Salinity Assay in Arabidopsis
Published: Vol 4, Iss 16, Aug 20, 2014 DOI: 10.21769/BioProtoc.1216 Views: 20756
Reviewed by: Arsalan DaudiAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
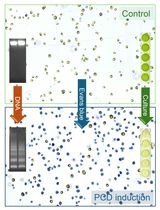
Detection and Quantification of Programmed Cell Death in Chlamydomonas reinhardtii: The Example of S-Nitrosoglutathione
Lou Lambert and Antoine Danon
Aug 5, 2024 1614 Views

Micrografting Technique of Hevea brasiliensis In Vitro Plantlets
Florence Dessailly [...] Julie Leclercq
Feb 20, 2025 1526 Views

Enzymatic Starch Quantification in Developing Flower Primordia of Sweet Cherry
Nestor Santolaria [...] Afif Hedhly
Apr 5, 2025 1934 Views
Abstract
Salinity is an important environmental constraint to crop productivity in arid and semi-arid regions of the world. The evaluation of the responses to salinity of different Arabidopsis ecotypes or transgenic lines is important to identify and investigate the role of different key genes. These new characterized genes involved in the response to salinity stress are of great interest to be incorporated in crops breeding programs. Here we provide a reproducible method to evaluate the performance of Arabidopsis lines to salinity stress by analysing primary and lateral root growth and fresh weight of plants grown under in vitro conditions in growth chambers. Even though NaCl is the most frequent used salinity tests, other salts (e.g. KCl, MgCl2) can be also evaluated by this method. Arabidopsis plants can be maintained for 15-20 days in these conditions, although effects on growth and biomass can be observed, depending on the used salt and concentration, within the first 10 days.
Keywords: Salinity toleranceMaterials and Reagents
- Arabidopsis thaliana ecotype Columbia (Col-0) and transgenic Arabidopsis lines 35S::SlCDF1 and 35S::SlCDF3 (background Col-0) seeds
- 70% ethanol
- Sodium hypochlorite (NaClO)
- Sodium dodecylsulphate (SDS)
- Sterile water
- Murashige & Skoog (MS) mediun including B5 vitamins (Duchefa Biochemie BV, catalog number: M0231 )
- MES (Duchefa Biochemie BV, catalog number: M1503 )
- Sucrose
- NaCl
- Agar (Duchefa Biochemie BV, catalog number: P1001 )
- Medium composition (see Recipes)
Equipment
- Square (120 x 120 mm) petri dishes polystyrene sterile
- Growth chamber
- Autoclave
- Shaker
- Image capturing device
- Precision balance (± 0.0001)
Software
- Image J software (http://rsb.info.nih.gov/ij) (Abrámoff et al., 2004)
Procedure
- Seed germination
- Seeds are surface sterilized by washing in 70% ethanol for 2 min, then in sterilization solution (1% sodium hypochlorite with 1% SDS in sterile water) for 12 min with shaking and finally washed five times with sterilized water.
- The seeds are stratified for 2 days at 4 °C in the dark.
- Seeds are surface sterilized by washing in 70% ethanol for 2 min, then in sterilization solution (1% sodium hypochlorite with 1% SDS in sterile water) for 12 min with shaking and finally washed five times with sterilized water.
- Growth conditions
- Sterilized seeds are placed in Petri dishes (approx. 70 seeds per Petri dish) on MS medium and grown for 6 days in vertical position, in a culture chamber at 22 °C under standard long-day conditions (16/8 h light/dark photoperiod).
- After six days of growth, seedlings exhibiting similar root length (7-10 mm) are transferred to square plates containing MS medium (control) or MS medium supplemented with 80 mM NaCl. About 20 seedlings are used per replicate and three replicates are made for each treatment.
- The initial position of the root tip is marked in the plate (Figure 1A).
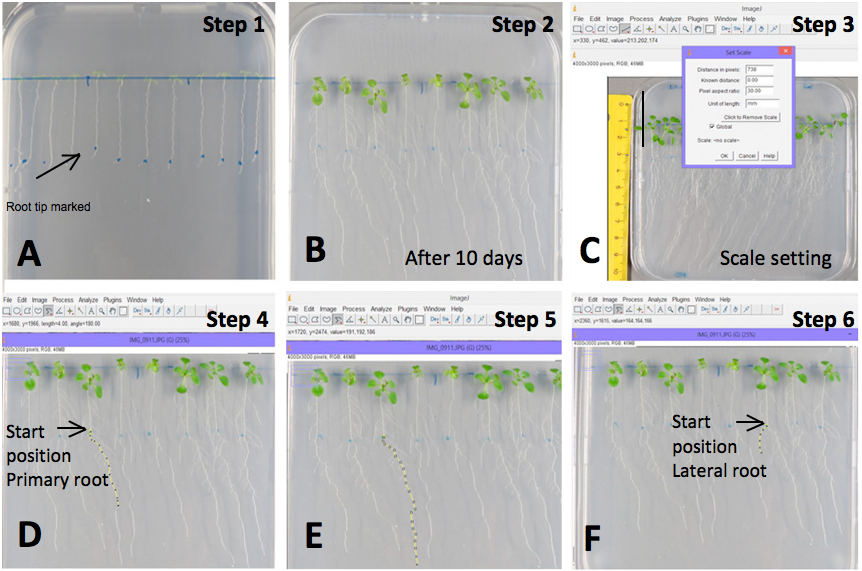

Figure 1. Example of primary and lateral root measurements with the Image J software. A) Root tip marked; B) Image of seedlings grown on MS medium supplemented with 80 mM NaCl after 10 days of treatment; C) Scale setting (e.g. 30 mm); D-E) Length measurement of the primary root from the marked point; and F) Length measurement of the lateral roots. - The plants are grown in vertical position under standard long day conditions (22 °C, 16/8 h light/dark) for 10 days, and then pictures are taken of each plate (Figure 1B). Primary and lateral root length and plant weight are then scored.
- Sterilized seeds are placed in Petri dishes (approx. 70 seeds per Petri dish) on MS medium and grown for 6 days in vertical position, in a culture chamber at 22 °C under standard long-day conditions (16/8 h light/dark photoperiod).
- Measurements root length
The measurement of primary and lateral root length is performed by analysis of the pictures with the Image J software.- First, set the scale using the ImageJ software: > draw a straight line of known distance > click on Analyze > set scale > known distance (e.g. 30 mm) > unit of length (mm) > global > ok (Figure 1C).
- Measure the length of primary root from the marked start position to the tip using the segmented line option of the software: Press Control + M to measure the length of the root (Figure 1D-E).
- Measure the length of each lateral root following the same procedure (Figure 1F).
- To evaluate growth differences between control and saline stress, measured lengths of the primary and lateral roots are represented as the percentage of root growth reduction relative to control conditions (MS medium) (for representative data see Figure 2).
- Statistical analyses are carried out by one-way analysis of variance (ANOVA) followed by a Student-Newman-Keuls test (P<0.01) (Figure 2).
- First, set the scale using the ImageJ software: > draw a straight line of known distance > click on Analyze > set scale > known distance (e.g. 30 mm) > unit of length (mm) > global > ok (Figure 1C).
- Measurement of plant fresh weight
Fresh weight measurements are estimated using a precision balance.- Whole 10 day-old-plants grown in MS medium or MS medium supplemented with 80 mM NaCl are weighed using a precision balance (Figure 3A-C).
- Statistical analyses are carried out by one-way analysis of variance (ANOVA) followed by a Student–Newman–Keuls test (P<0.01). The percentage of growth reduction by salinity relative to control conditions is compared between the studied genotypes.
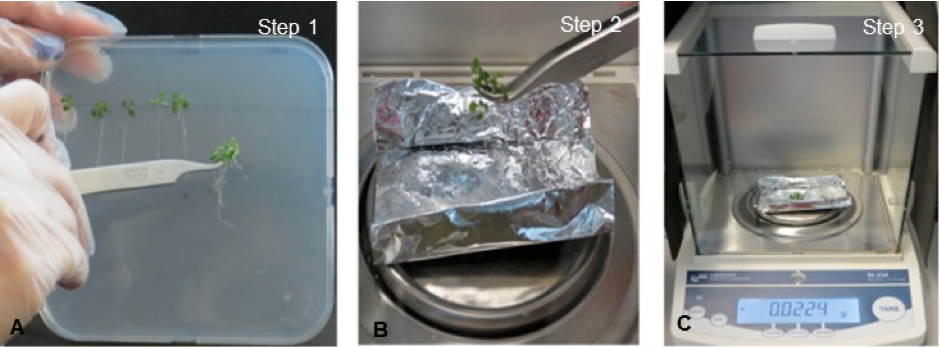

Figure 2. View of plant fresh weight measurements. A) Plantlets are carefully removed from the medium and B-C) weighed with a precision balance.
- Whole 10 day-old-plants grown in MS medium or MS medium supplemented with 80 mM NaCl are weighed using a precision balance (Figure 3A-C).
Representative data
- Example of results primary and lateral root elongation measurements under salinity stress after 10 days of treatment using ImageJ software. The measurement are performed as described above (item C: 4 and 5).
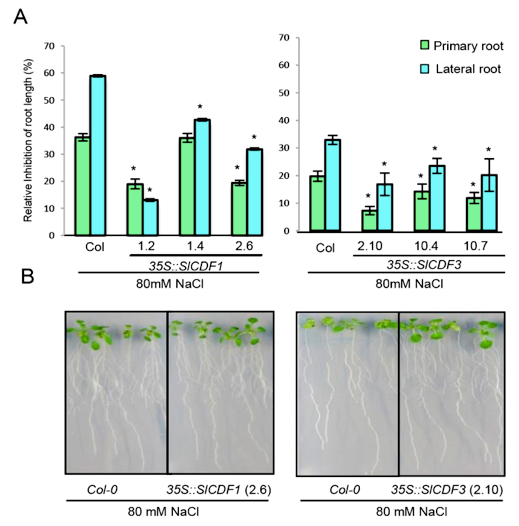
Figure 3. Example of measured effects of salt stress on Arabidopsis root growth in lines overexpressing SlCDF1 and SlCDF3 genes (Corrales et al., 2014). A) Salt stress tolerance estimated by determining the reduction of primary root and lateral root growth of 35S::SlCDF1 lines 1.2, 1.4 and 2.6, 35S::SlCDF3 lines 2.10, 10.4 and 10.7 and wild type Col-0 plants after 10 days in MS supplemented with 80 mM NaCl. The percentage of reduction relative to control conditions is represented. Data are expressed as means ± standard errors of three independent experiments with at least 20 plants each. Asterisks indicate significant differences between Col-0 and 35S::SlCDF1 or 35S::SlCDF3 overexpressing lines (P<0.05; ANOVA Student-Newman-Keuls tests. B) Representative images of WT, 35S::SlCDF1 line 2.6 and 35S::SlCDF3 line 2.10 after the treatments.
Recipes
- Medium composition
- For 1 L of MS control medium
2.2 g of Murashige & Skoog (MS) medium including B5 vitamins
5 g of sucrose
5 g MES
Adjust pH to 5.7 with KOH
Add 7 g agar
Sterilized for 20 min at 121 °C/1 atm using an autoclave - For 1 L the medium MS supplemented 80 mM with NaCl
4.67 g NaCl is added to the medium
- For 1 L of MS control medium
Acknowledgments
We gratefully acknowledge funding through grants from Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria (INIA; project numbers: 2009-0004-C01, 2012-0008-C01) and the Spanish Ministry of Science and Innovation (project number: BIO2010-14871 and ERA-NET GEN2006-27772-C2-2). Additionally, we are grateful for the protocol described by Lakhssassi et al. (2012).
References
- Abràmoff, M. D., Magalhães, P. J. and Ram, S. J. (2004). Image processing with ImageJ. Biophotonics Inter 11(7): 36-43.
- Corrales, A. R., Nebauer, S. G., Carrillo, L., Fernandez-Nohales, P., Marques, J., Renau-Morata, B., Granell, A., Pollmann, S., Vicente-Carbajosa, J., Molina, R. V. and Medina, J. (2014). Characterization of tomato Cycling Dof Factors reveals conserved and new functions in the control of flowering time and abiotic stress responses. J Exp Bot 65(4): 995-1012.
- Lakhssassi, N., Doblas, V. G., Rosado, A., del Valle, A. E., Posé, D., Jimenez, A. J., Castillo, A. G., Valpuesta, V., Borsani, O. and Botella, M. A. (2012). The Arabidopsis tetratricopeptide thioredoxin-like gene family is required for osmotic stress tolerance and male sporogenesis. Plant Physiol 158(3): 1252-1266.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Corrales, A. R., Carrillo, L., Nebauer, S. G., Renau-Morata, B., Sánchez-Perales, M., Fernández-Nohales, P., Marqués, J., Granell, A., Pollmann, S., Vicente-Carbajosa, J., Molina, R. V. and Medina, J. (2014). Salinity Assay in Arabidopsis. Bio-protocol 4(16): e1216. DOI: 10.21769/BioProtoc.1216.
Category
Plant Science > Plant physiology > Abiotic stress
Plant Science > Plant physiology > Plant growth
Plant Science > Plant developmental biology > General
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











