- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Signaling Assays for Detection of Human G-protein-coupled Receptors in Yeast
Published: Vol 4, Iss 16, Aug 20, 2014 DOI: 10.21769/BioProtoc.1206 Views: 10985
Reviewed by: Belen SanzMaureen WirschellAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
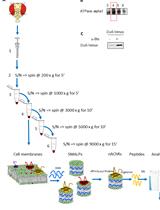
Enrichment of Membrane Proteins for Downstream Analysis Using Styrene Maleic Acid Lipid Particles (SMALPs) Extraction
Benedict Dirnberger [...] Kathryn S. Lilley
Aug 5, 2023 2978 Views
A Computational Workflow for Membrane Protein–Ligand Interaction Studies: Focus on α5-Containing GABA (A) Receptors
Syarifah Maisarah Sayed Mohamad [...] Ahmad Tarmizi Che Has
Nov 20, 2025 2123 Views
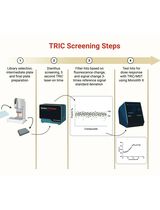
Orthogonal Temperature-Related Intensity Change and Time-Resolved Förster Resonance Energy Transfer High-Throughput Screening Platform for the Discovery of SLIT2 Binders
Moustafa T. Gabr [...] Somaya A. Abdel-Rahman
Feb 20, 2026 99 Views
Abstract
G-protein-coupled receptors (GPCRs) are the largest group of cell-surface proteins and are major molecular targets for drug development. The protocol described herein is for the detection of human GPCR signaling in the yeast Saccharomyces cerevisiae. Using Zoanthus sp. green fluorescent protein (ZsGreen) as the reporter, engineered yeast cells expressing human GPCRs emit strong fluorescence in response to stimuli leading to receptor signal activation. This assay method would allow screening for agonistic ligands and critical mutations required for human GPCR signaling.
Keywords: Signaling assayMaterials and Reagents
- Engineered Saccharomyces cerevisiae strain in which the ZsGreen reporter genes were integrated into the genome (IMFD-72ZsD: MATa his3Δ1 leu2Δ0 met15Δ0 ura3Δ0 sst2Δ::AUR1-C ste2Δ::LEU2 fig1Δ::ZsGreen his3Δ::PFIG1-ZsGreen far1Δ gpa1Δ::Gi3tp) (Nakamura et al., 2013)
Note: The expression of ZsGreen is controlled by the signal-responsive FIG1 promoter.
- Multi-copy expression plasmid (pGK421 containing the PGK1 promoter, 2 μ origin and MET15 marker) (Togawa et al., 2010) encoding the GPCR of interest [e.g., somatostatin receptor subtype-5 (SSTR5); somatostatin receptor subtype-2 (SSTR2); or neurotensin receptor type-1 (NTSR1)] (Ishii et al., 2012; Ishii et al., 2014)
- GPCR ligands [e.g., somatostatin (SST) (Merck KGaA, Calbiochem®, catalog number: 51110-01-1 ) and neurotensin (NTS) (Merck KGaA, Calbiochem®, catalog number: 39379-15-2)]
- 10 mg/ml carrier DNA (Takara Bio Company, Clontech, catalog number: 630440 )
- Tris (hydroxymethyl) aminomethane (Tris.HCl) (Nacalai Tesque, catalog number: 35409-45 )
- Ethylenediaminetetraacetic acid disodium salt dihydrate (EDTA 2Na∙2H2O) (Nacalai Tesque, catalog number: 15111-45 )
- 1 M Hydrochloric acid (HCl) (Nacalai Tesque, catalog number: 37314-15 )
- Lithium acetate dihydrate (Sigma-Aldrich, catalog number: L6883 )
- Acetic acid (Nacalai Tesque, catalog number: 00212-85 )
- Polyethylene glycol (PEG) #4000 (Nacalai Tesque, catalog number: 28221-05 )
- DMSO (Nacalai Tesque, catalog number: 13445-74 )
- BD FACSFlow sheath fluid (BD, catalog number: 342003 )
- Immersion oil (Olympus, catalog number: IMMOIL-F30CC )
- Distilled water (dH2O)
- Yeast extract (Nacalai Tesque, catalog number: 15838-45 )
- Peptone (BD, catalog number: 211677 )
- D-Glucose (Nacalai Tesque, catalog number: 16806-25 )
- Yeast nitrogen base without amino acids (YNB) (BD, catalog number: 291940 )
- L-Histidine (Nacalai Tesque, catalog number: 18116-92 )
- L-Leucine (Nacalai Tesque, catalog number: 20327-62 )
- Uracil (Nacalai Tesque, catalog number: 35824-82 )
- 3-(N-Morpholino)-2-hydroxypropanesulfonic acid (Mopso) (Nacalai Tesque, catalog number: 23421-64 )
- Sodium hydroxide (NaOH) (Nacalai Tesque, catalog number: 31511-05 )
- Agar (Nacalai Tesque, catalog number: 01028-85 )
- 10 x TE buffer (see Recipes)
- 10 x LiAc (see Recipes)
- 50% PEG (see Recipes)
- TE/LiAc solution (see Recipes)
- LiAc/PEG solution (see Recipes)
- YPD medium (see Recipes)
- Synthetic dextrose (SD) selective medium supplemented with histidine, leucine and uracil (see Recipes)
- SDM71 selective medium supplemented with histidine, leucine and uracil (see Recipes)
Equipment
- 16.5 x 105 mm test tubes (AGC Techno Glass, catalog number: 9820TST16.5-105NP )
- 10-ml conical flasks (AGC Techno Glass, catalog number: 4980FK10 )
- 2-ml microcentrifuge tubes (WATSON, catalog number: 332-720C )
- 1.5-ml microcentrifuge tubes (WATSON, catalog number: 131-815C )
- 96-well cell culture plate (Corning, catalog number: 3596 )
- Microplate sealing tape (AS ONE Corporation, catalog number: 1-6774-05 )
- 12.5 x 75 mm test tubes with 2-position caps (B & M Equipment, catalog number: 222-2036-050 )
- Microscope glass slides (Matsunami Glass, catalog number: S091150 )
- 18 x 18 mm coverslips (Matsunami Glass, catalog number: C218181 )
- Centrifuge (Eppendorf, model: MiniSpin plus )
- Block incubator (Astec Industries, model: BI-516C )
- Shaking incubator for test tubes and conical flasks (TAITEC, model: BR-43FL )
- Shaking incubator for a 96-well cell culture plate (TAITEC, model: M∙BR-022UP )
- Spectrophotometer (Shimadzu, model: UVmini-1240 )
- BD FACSCanto II flow cytometer (BD)
- Fluorescence microscope (Keyence Corporation, model: BZ-9000 )
- 100x objective lens (Nikon Corporation, model: CFI Plan Apo VC 100x H )
- BZ filter cube (excitation filter, absorption filter, dichroic mirror) (Keyence Corporation, model: OP-66836 GFP-BP )
Software
- BD FACSDiva software (v5.0) (BD)
Procedure
- Yeast transformation
Transformation was performed using the lithium acetate method (Gietz et al., 1992).- Inoculate 5 ml YPD medium in a 16.5 x 105 mm test tube with a single colony of S. cerevisiae strain IMFD-72ZsD.
- Incubate the culture overnight at 30 °C with shaking at 150 opm.
Note: Shaking incubator (TAITEC, model: BR-43FL).
- Transfer 2 ml of the cell culture to a 2-ml microcentrifuge tube and pellet the cells by centrifugation at 3,000 rpm for 5 min at room temperature.
- Remove the supernatant and wash the cells with 1 ml sterilized dH2O.
- Pellet the cells by centrifugation at 3,000 rpm for 5 min at room temperature.
- Remove the supernatant and resuspend the cells in 1.5 ml of TE/LiAc solution.
- Aliquot 100 μl of yeast suspension to a fresh 1.5-ml microcentrifuge tube.
- Add 500 ng of plasmid DNA (pGK421-SSTR5, -SSTR2 or -NTSR1) and 20 μg of carrier DNA to each tube.
Note: Before using the carrier DNA, denature it at 95 °C for 5 min, and then chill it quickly on ice.
- Add 600 μl of LiAc/PEG solution to each tube and vortex at high speed for 10 sec to mix.
- Incubate the cell suspension at 30 °C for 30 min.
- Add 70 μl of DMSO. Mix well by gentle inversion. Do not vortex.
- Heat shock at 42 °C for 15 min.
- Centrifuge the cell suspension at 14,000 rpm for 5 sec at room temperature. Remove the supernatant completely.
- Resuspend the cells in 0.5 ml of 1x TE buffer.
- Spread 100 μl of cell suspension on each SD selective agar plate.
Note: Since the yeast strain has his3Δ leu2Δ met15Δ ura3Δ alleles and the plasmid has MET15 as a selection marker, the selective medium requires supplementation with histidine, leucine and uracil.
- Incubate the plates at 30 °C until colonies appear (generally, 2-4 days).
- Inoculate 5 ml YPD medium in a 16.5 x 105 mm test tube with a single colony of S. cerevisiae strain IMFD-72ZsD.
- GPCR signaling assay
- Inoculate 5 ml SD selective medium in a 16.5 x 105 mm test tube with a single colony of a positive transformant.
- Incubate the culture overnight at 30 °C with shaking at 150 opm.
Note: Shaking incubator (TAITEC, model: BR-43FL).
- Measure the OD600 of the yeast cultures.
Note: Spectrophotometer.
- Transfer the cultured cells into 5 ml of fresh SD selective medium in a 10-ml conical flask to give an initial OD600 of 0.03.
- Further incubate the culture at 30 °C for 18 h with rotary shaking at 150 rpm.
Note: Shaking incubator (TAITEC, model: BR-43FL).
- Transfer 2 ml of the cell culture to a 2-ml microcentrifuge tube and pellet the cells by centrifugation at 3,000 rpm for 5 min at room temperature.
- Remove the supernatant and wash the cells with 1 ml sterilized dH2O.
- Pellet the cells by centrifugation at 3,000 rpm for 5 min at room temperature.
- Remove the supernatant and resuspend the cells in ~400 μl sterilized dH2O.
- Measure the OD600 of the cell suspension, and then dilute it to an OD600 of 10.
Note: Spectrophotometer.
- Add 10 μl of the resulting cell suspension (to give an OD600 of 1) to 80 μl fresh SDM71 selective medium per well of a 96-well cluster dish.
- Add 10 μl of 10-times concentrated ligands (SST or NTS) or sterilized dH2O (no ligand control) to each well.
Note: A final concentration of ~10 μM ligand is recommended (dissolved in and diluted with sterilized dH2O).
- Seal the 96-well cluster dish with sealing tape.
- Incubate the dish at 30 °C for 4 h with shaking at 150 rpm. After incubation, measure green fluorescence using a flow cytometer or fluorescence microscope (see below).
Note: Shaking incubator (TAITEC, model: M∙BR-022UP).
Flow cytometry analysis- Dilute the cell culture in 1 ml sheath fluid in test tubes with 2-position caps.
- Measure the green fluorescence signal emitted from 10,000 cells using a BD FACSCanto II flow cytometer equipped with a 530/30 nm band-pass filter.
- Analyze the data and assess the mean ZsGreen fluorescence of 10,000 cells using BD FACSDiva software.
Fluorescence microscopy imaging- Transfer the cell culture to a 1.5-ml microcentrifuge tube and pellet the cells by centrifugation at 3,000 rpm for 5 min at room temperature.
- Remove the supernatant and wash the cells with 100 μl sterilized dH2O.
- Pellet the cells by centrifugation at 3,000 rpm for 5 min at room temperature.
- Remove the supernatant and resuspend the cells in 10 μl sterilized dH2O.
- Spot 7 μl of the cell suspension on a glass slide and mount a coverslip over the spot.
- Observe the cells using a BZ-9000 fluorescence microscope equipped with a 100x objective lens or equivalent and acquire green fluorescence images with a 470/40 band-pass filter for excitation and a 535/50 band-pass filter for emission.
Note: The 100x objective lens should be immersed in oil.
- Dilute the cell culture in 1 ml sheath fluid in test tubes with 2-position caps.
- Inoculate 5 ml SD selective medium in a 16.5 x 105 mm test tube with a single colony of a positive transformant.
Representative data
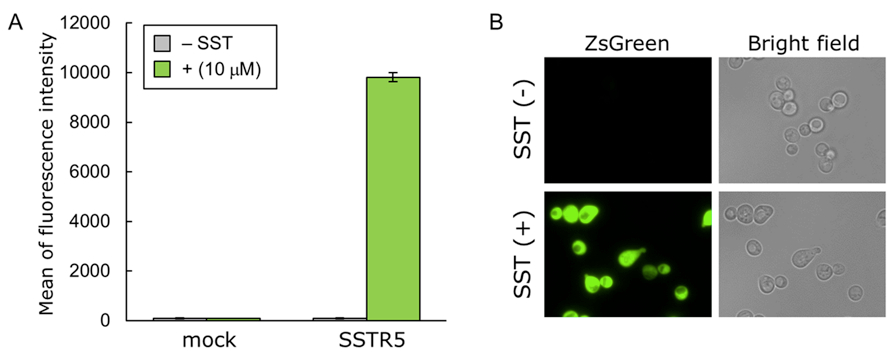
Figure 1. Activation of human SSTR5 produced in yeast following the exogenous addition of SST. Yeast strain IMFD-72ZsD was transformed with pGK421 (mock) or pGK421-SSTR5. All transformants were grown in SD medium for 18 h. The cells then were incubated for another 4 h in SDM71 medium with or without 10 μM SST. A. The ZsGreen fluorescence of 10,000 cells was measured by flow cytometry. Data are shown as means ± SDs of triplicate samples. B. Visualization of the ZsGreen fluorescence of IMFD-72ZsD/SSTR5 with or without 10 μM SST. Images were obtained using a 100x objective on a fluorescence microscope. Reprinted with permission from Ishii et al. (2012).
Notes
- Work on a clean bench.
Recipes
- 10x TE buffer (200 ml)
2.4 g Tris-HCl (0.1 M)
0.74 g EDTA 2Na∙2H2O (10 mM)
Dissolved in 175 ml dH2O
Adjust pH to 7.5 with 1 M HCl
Adjust volume to 200 ml with dH2O
Sterilized by autoclaving
- 10x LiAc (200 ml)
20.4 g lithium acetate dihydrate (1 M)
Dissolved in 175 ml dH2O
Adjust pH to 7.5 with dilute acetic acid
Adjust volume to 200 ml with dH2O
Sterilized by autoclaving
- 50% PEG (200 ml)
100 g PEG #4000
Add dH2O to 200 ml
Sterilized by autoclaving
- TE/LiAc solution (1.5 ml)
150 μl 10 x TE buffer
150 μl 10 x LiAc
1.2ml sterilized dH2O
Note: Prepare fresh just prior to use.
- LiAc/PEG solution (1 ml)
800 μl 50% PEG
100 μl 10 x TE buffer
100 μl 10 x LiAc
Note: Prepare fresh just prior to use.
- YPD medium (1 L)
10 g yeast extract
20 g peptone
20 g D-Glucose
Add dH2O to 1 L
Sterilized by autoclaving
- SD selective medium supplemented with histidine, leucine and uracil (1 L)
20 g D-Glucose
6.7 g yeast nitrogen base without amino acids (YNB)
20 mg L-Histidine
60 mg L-Leucine
20 mg Uracil
Add dH2O to 1 L
Sterilized by autoclaving
Note: For solid plates, add 2% agar to the media.
- SDM71 selective medium supplemented with histidine, leucine and uracil (1 L)
20 g D-Glucose
1.7g yeast nitrogen base without amino acids (YNB)
45 g 3-(N-morpholino)-2-hydroxypropanesulfonic acid (Mopso)
20 mg L-Histidine
60 mg L-Leucine
20 mg uracil
Dissolved in 800 ml dH2O
Adjust pH to 7.1 with 10 N NaOH
Adjust volume to 1 L with dH2O
Filter sterilized
Acknowledgments
This protocol was adapted from the previously published paper Nakamura et al. (2013). This work was supported by a Grant-in-Aid for JSPS Fellows (23∙2292) and for Young Scientists (B) (26820362) from the Japan Society for the Promotion of Science.
References
- Gietz, D., St Jean, A., Woods, R. A. and Schiestl, R. H. (1992). Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res 20(6): 1425.
- Ishii, J., Moriguchi, M., Hara, K. Y., Shibasaki, S., Fukuda, H. and Kondo, A. (2012). Improved identification of agonist-mediated Gαi-specific human G-protein-coupled receptor signaling in yeast cells by flow cytometry. Anal Biochem 426(2): 129-133.
- Ishii, J., Oda, A., Togawa, S., Fukao, A., Fujiwara, T., Ogino, C. and Kondo, A. (2014). Microbial fluorescence sensing for human neurotensin receptor type 1 using Gα-engineered yeast cells. Anal Biochem 446: 37-43.
- *Nakamura, Y., Ishii, J. and Kondo, A. (2013). Bright fluorescence monitoring system utilizing Zoanthus sp. green fluorescent protein (ZsGreen) for human G-protein-coupled receptor signaling in microbial yeast cells. PLos One 8(12): e82237.
- Togawa, S., Ishii, J., Ishikura, A., Tanaka, T., Ogino, C. and Kondo, A. (2010). Importance of asparagine residues at positions 13 and 26 on the amino-terminal domain of human somatostatin receptor subtype-5 in signalling. J Biochem 147(6): 867-873.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Nakamura, Y., Ishii, J. and Kondo, A. (2014). Signaling Assays for Detection of Human G-protein-coupled Receptors in Yeast. Bio-protocol 4(16): e1206. DOI: 10.21769/BioProtoc.1206.
Category
Immunology > Immune cell function > General
Biochemistry > Protein > Interaction > Protein-ligand interaction
Cell Biology > Cell signaling > Stress response
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








