- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Extraction and Measurement the Activities of Cytosolic Phosphoenolpyruvate Carboxykinase (PEPCK) and Plastidic NADP-dependent Malic Enzyme (ME) on Tomato (Solanum lycopersicum)
Published: Vol 4, Iss 9, May 5, 2014 DOI: 10.21769/BioProtoc.1122 Views: 11164
Reviewed by: Tie Liu

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
Immunofluorescence for Detection of TOR Kinase Activity In Situ in Photosynthetic Organisms
Ana P. Lando [...] Giselle M. A. Martínez-Noël
Dec 20, 2024 1824 Views

An Activity-Based Proteomics with Two-Dimensional Polyacrylamide Gel Electrophoresis (2D-PAGE) for Identifying Target Proteases in Arabidopsis Apoplastic Fluid
Sayaka Matsui and Yoshikatsu Matsubayashi
Mar 5, 2025 1989 Views
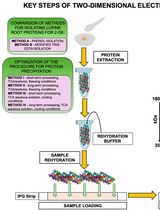
Advancing 2-DE Techniques: High-Efficiency Protein Extraction From Lupine Roots
Sebastian Burchardt [...] Emilia Wilmowicz
Oct 5, 2025 1772 Views
Abstract
A recent study demonstrated that cytosolic phosphoenolpyruvate carboxykinase (PEPCK) and NADP-malic enzyme (NADP-ME) have an important role in malate metabolism during fruit ripening (Osorio et al., 2013). PEPCK catalyze the ATP-dependent decarboxylation of oxaloacetate (OAA) to phosphoenolpyruvate (PEP) and NADP-ME, the reversible conversion of malate and pyruvate. Here, we present the detailed protocols to measure PEPCK activity in carboxylation direction by following oxidation of NADH and to measure NADP-ME activity based upon the reduction of NADP+.
Materials and Reagents
- PEPCK activity
- Un-harvested plant tissues (how to freeze the tissue is explained in the procedure section)
- Liquid N2
- Ice
- Bicine (Sigma-Aldrich, catalog number: B8660 )
- KOH
- EDTA (Sigma-Aldrich, catalog number: EDS-100G )
- Poly (ethylene glycol)-4000 (Sigma-Aldrich, catalog number: 81240 )
- Dithiothreitol (DTT) (Sigma-Aldrich, catalog number: 43815 )
- β-nicotinamide adenine dinucleotide reduced form (NADH) (Roche Diagnostics, catalog number: 10128015001 )
- 4-(2-Hydroxyethyl)piperazine-1-ethanesulfonic acid, N-(2-Hydroxyethyl)piperazine-N′-(2-ethanesulfonic acid) (HEPES) (Sigma-Aldrich, catalog number: H3375 )
- KCl
- MnCl2
- Phosphoelnolpyryvate (PEP) (Bio Vectra, catalog number: 2552)
- Adenosine 5′-diphosphate sodium salt (ADP) (Sigma-Aldrich, catalog number: A2754 )
- KHCO3
- L-Malate dehydrogenase (Roche Diagnostics, catalog number: 10127248001 )
- Bradford stock solution (Bio-Rad Laboratories, catalog number, 500-0006 )
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: A2058 )
- Extraction buffer 1 (see Recipes)
- Extraction buffer 2 (see Recipes)
- Buffer 3 (see Recipes)
- PEPCK assay mix 1 (see Recipes)
- Un-harvested plant tissues (how to freeze the tissue is explained in the procedure section)
- NADP-ME activity
- Un-harvested plant tissues (how to freeze the tissue is explained in the procedure section)
- Liquid N2
- Ice
- Tris-Base (United State Biological, catalog number: T8600 )
- MnCl2
- EDTA (Sigma-Aldrich, catalog number: EDS-100G)
- Glycerol (Sigma-Aldrich, catalog number: G5516 )
- 2-mercaptoethanol (Sigma-Aldrich, catalog number: M6250 )
- β-nicotinamide adenine dinucleotide phosphate (NADP+) (Roche Diagnostics, catalog number: 10128058001 )
- L-Malate (Sigma-Aldrich, catalog number: M1000 )
- Bradford stock solution (Bio-Rad Laboratories, catalog number, 500-0006)
- Bovine serum albumin (BSA) (Sigma-Aldrich, catalog number: A2058)
- Extraction buffer 4 (see Recipes)
- NADP-ME assay mix 1 (see Recipes)
- Un-harvested plant tissues (how to freeze the tissue is explained in the procedure section)
Equipment
- Small mortar and pestle
- 2 ml and 1.5 ml microfuge tubes
- Pipettes
- Balance
- 2 ml centrifuge (Hettich Mikro 22R)
- 96 well polystyrene microplate (flat bottom) (Corning, catalog number: 3300 )
- A computer supported microplate spectrophotometer for kinetic (time-course) measurement mode (Elisa microplate-spetrophotometer) (BioTek Instruments, model: EL808 )
Procedure
- Extraction for measuring PEPCK activity
This protocol applies to extraction of PEPCKs from plant tissues.
- Collect the sample and freeze immediately in liquid N2. Store at -80 °C until use.
- Add small amount of liquid N2 and frozen sample into a mortar. Grind the tissue until the sample is a fine powder. Carefully add more liquid N2 if needed to keep frozen.
- Weight 500 mg of tissue into microfuge tube and add 900 µl of extraction buffer 1 (EB1) (for fruit tissue the ratio of powered tissue/EB is 1:1.8, w/v. This ratio may need to be increased to 1:3 for leaf tissue).
- Keep on ice while preparing all samples and vortex for 30 sec.
- Centrifuge 15 min at 4 °C and 13,000 x g. Remove 500 µl supernatant (clarified extract) to a fresh microfuge tube and add 850 µl of extraction buffer 2 (EB2), to give a final concentration of 35% PEG (mix well).
- Incubate on ice for 10 min and centrifuge for 20 min at 13,000 x g. The supernatant is discarded.
- Re-suspend the pellet in 100 µl of buffer 3 (B3) and keep on ice. Measure immediately PEPCK activity or the aliquots of clarified extracts can be snap frozen in liquid N2 and stored at -80 °C for future use.
- Collect the sample and freeze immediately in liquid N2. Store at -80 °C until use.
- PEPCK activity
The activity of PEPCK is measured in carboxylation direction by following oxidation of NADH at 340 nm at 25 °C. Briefly, a continuous assay is used in which OAA produced by PEPCK is immediately reduced to malate, and this is achieved by the inclusion of malate dehydrogenase. The oxidation of NADH by malate dehydrogenase is measured at 340 nm using a spectrophotometer.
- Accurately pipette 2-10 µl of clarified extract into a microplate well.
- Use repeat pipetor to add 150 µl PEPCK assay mix 1 to each well and immediately place in microplate spectrophotometer. Continuously monitor NADH oxidation to NAD+ as a decline in absorbance at 340 nm (A340), taking readings every 5-10 sec for up to 5 min (the samples need to shake while reading).
- Correct for background NADH oxidation by omitting PEP from the reaction mixture. Ensure that the decline in A340 (amount of NADH being oxidized; ε340 = 6,220/M/cm) is proportional to assay time and concentration of enzyme assayed. Dilution of clarified extract in extraction buffer may be necessary for samples containing abundant PEPCK activity.
Note: One international unit (U) of enzyme activity is defined as the amount of enzyme resulting in the production of 1 µmol of product per min at 25 °C. PEPCK activity in (U/ml clarified extract) = (ΔA340/min x clarified extract dilution factor)/6.22. Thus, if 3.0 μl of clarified extract mixed with 150 μl of PEPCK reaction mixture yields a ΔA340/min of 0.2 at 340 nm, then the PEPCK activity = (0.2 x 50)/6.22 U/ml = 1.6 U/ml.
- Accurately pipette 2-10 µl of clarified extract into a microplate well.
- Extraction for measuring NADP-ME activity
This protocol applies to extraction of NADP-MEs from plant tissues.
- Collect the sample and freeze immediately in liquid N2. Store at -80 °C until use.
- Add small amount of liquid N2 and frozen sample into a mortar. Grind the tissue until the sample is a fine powder. Carefully add more liquid N2 if needed to keep frozen.
- Weight 100 mg of tissue into microfuge tube and add 200 µl of extraction buffer 4 (EB4) (for fruit tissue the ratio of powered tissue/EB is 1:2, w/v. This ratio may need to be increased to 1:3 for leaf tissue).
- Keep on ice while preparing all samples and vortex for 30 sec.
- Centrifuge 10 min at 4 °C and 13,000 x g. Transfer the supernatant to a fresh microfuge tube. Measure NADP-ME activity immediately or the aliquots of clarified extracts can be snap frozen in liquid N2 and stored at -80 °C for future use.
- Collect the sample and freeze immediately in liquid N2. Store at -80 °C until use.
- NADP-ME activity
The activity of NADP-ME is based upon the reduction of NADP+ at 340 nm at 25 °C. Briefly, a continuous assay is used in which malate is oxidized to pyruvate and CO2, and NADP+ is reduced to NADPH. The reduction of NADP+ by NADP-ME is measured at 340 nm using a spectrophotometer.
- Accurately pipette 5-10 µl of clarified extract into a microplate well.
- Use repeat pipetor to add 100 µl NADP-ME assay mix 1 to each well and immediately place in microplate spectrophotometer. Continuously monitor NADP+ reduction to NADPH as an increase in absorbance at 340 nm (A340), taking readings every 5-10 sec for up to 5 min (the samples need to shake while reading).
- Correct for background NADP+ reduction by omitting L-malate from the reaction mixture. Ensure that the increase in A340 is proportional to assay time and concentration of enzyme assayed. Dilution of clarified extract in extraction buffer may be necessary for samples containing abundant NADP-ME activity.
Note: One international unit (U) of enzyme activity is defined as the amount of enzyme that catalyzes the formation of 1 µmol of NADPH/min under the specified conditions. NADP-ME activity in (U/ml clarified extract) = (ΔA340/min x clarified extract dilution factor)/6.22. Thus, if 2.0 μl of clarified extract mixed with 100 μl of NADP-ME reaction mixture yields a ΔA340/min of 1.5 at 340 nm, then the NADP-ME activity = (1.5 x 50)/6.22 U/ml = 12.5 U/ml.
- Accurately pipette 5-10 µl of clarified extract into a microplate well.
- Bradford assay of protein concentration
- For preparing the BSA standard curve:
Make 1 ml stock solution of 10 mg BSA/200 ml phosphate buffered saline (PBS) (pH 7.4) and prepare a duplicate standard curve, using template below:
Final concentration (ng/µl)
Vol. (µl) BSA stock (0.05 mg/ml)
PBS (µl)
µg protein/well
0
0
200
0
6.25
25
175
1.25
12.5
50
150
2.5
18.7
75
125
3.75
25.0
100
100
5
- Pipette 50 µl of aliquot supernatants (or its dilutions) or BSA standards to each well.
- Add 130 µl Bradford stock solution (see Materials and Reagents section) previously dissolved in distillate water (dilution 1/5).
- Shake briefly and wait 5 min. Determinate the OD at 595 nm. Use slope and blank obtained from the calibration to determinate the protein concentration in each extracts.
- For preparing the BSA standard curve:
Recipes
- Extraction buffer 1 (EB1, kept on ice)
500 mM Bicine-KOH (pH 9.0)
200 mM KCl
3 mM EDTA
5% (w/v) PEG-4000
25 mM DTT (add freshly every time before using it)
0.4% BSA
- Extraction buffer 2 (EB 2, kept on ice)
500 mM Bicine-KOH (pH 9.0)
3 mM EDTA
55% (w/v) PEG-4000
25 mM DTT (add freshly every time before using it)
- Buffer 3 (EB3, kept on ice)
10 mM Bicine-KOH (pH 9.0)
25 mM DTT (add freshly every time before using it)
Note: EB1, EB2, and EB3 without DTT can be kept at 4 ºC for few weeks.
- PEPCK assay mix 1
100 mM Hepes-KOH (pH 6.8)
100 mM KCl
0.14 mM NADH
25 mM DTT (add freshly every time before using it)
6 mM MnCl2
6 mM PEP
1 mM ADP
90 mM KHCO3
6 U ml-1 malate dehydrogenase
- Extraction buffer 4 (EB4, kept on ice)
100 mM Tris-HCl (pH 8.0)
5 mM MgCl2
2 mM EDTA
10% (v/v) glycerol
10 mM 2-mercaptoethanol
- NADP-ME assay mix 1 (kept on ice)
50 mM Tris-HCl (pH 8.0)
10 mM MgCl2
0.5 mM NADP+
10 mM L-malate
Acknowledgments
We acknowledge the excellent care of the plants by Helga Kulka (Max- Planck-Institut für Molekulare Planzenphysiologie) and Hanna Levanony (Weizmann Institute of Science). This protocol was adapted from Osorio et al. (2013).
References
- Centeno, D. C., Osorio, S., Nunes-Nesi, A., Bertolo, A. L., Carneiro, R. T., Araujo, W. L., Steinhauser, M. C., Michalska, J., Rohrmann, J., Geigenberger, P., Oliver, S. N., Stitt, M., Carrari, F., Rose, J. K. and Fernie, A. R. (2011). Malate plays a crucial role in starch metabolism, ripening, and soluble solid content of tomato fruit and affects postharvest softening. Plant Cell 23(1): 162-184.
- Osorio, S., Vallarino, J. G., Szecowka, M., Ufaz, S., Tzin, V., Angelovici, R., Galili, G. and Fernie, A. R. (2013). Alteration of the interconversion of pyruvate and malate in the plastid or cytosol of ripening tomato fruit invokes diverse consequences on sugar but similar effects on cellular organic acid, metabolism, and transitory starch accumulation. Plant Physiol 161(2): 628-643.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Osorio, S., Vallarino, J. G., Szecowka, M., Ufaz, S., Tzin, V., Angelovici, R., Galili, G. and Fernie, A. R. (2014). Extraction and Measurement the Activities of Cytosolic Phosphoenolpyruvate Carboxykinase (PEPCK) and Plastidic NADP-dependent Malic Enzyme (ME) on Tomato (Solanum lycopersicum). Bio-protocol 4(9): e1122. DOI: 10.21769/BioProtoc.1122.
- Osorio, S., Vallarino, J. G., Szecowka, M., Ufaz, S., Tzin, V., Angelovici, R., Galili, G. and Fernie, A. R. (2013). Alteration of the interconversion of pyruvate and malate in the plastid or cytosol of ripening tomato fruit invokes diverse consequences on sugar but similar effects on cellular organic acid, metabolism, and transitory starch accumulation. Plant Physiol 161(2): 628-643.
Category
Plant Science > Plant biochemistry > Protein > Isolation and purification
Biochemistry > Protein > Isolation and purification
Plant Science > Plant biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









