- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
DNA Damage Sensitivity Assays with Arabidopsis Seedlings
Published: Vol 4, Iss 7, Apr 5, 2014 DOI: 10.21769/BioProtoc.1093 Views: 11332
Reviewed by: Masahiro MoritaRu ZhangAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Tensile Testing Assay for the Measurement of Tissue Stiffness in Arabidopsis Inflorescence Stem
Kouki Yoshida [...] Nobutaka Mitsuda
Aug 5, 2019 10171 Views
Analysis of Monosaccharides from Arabidopsis Seed Mucilage and Whole Seeds Using HPAEC-PAD
Gillian H. Dean [...] George W. Haughn
Dec 20, 2019 5755 Views
Targeting Ultrastructural Events at the Graft Interface of Arabidopsis thaliana by A Correlative Light Electron Microscopy Approach
Clément Chambaud [...] Lysiane Brocard
Jan 20, 2023 2981 Views
Abstract
We describe fast and reproducible sensitivity assays to quantify the response of Arabidopsis seedlings of different genotypes to a wide range of DNA damaging agents. We apply (1) γ-irradiation, which produces DNA breaks, (2) bleocin, a radiomimetic drug, (3) mitomycin C, a DNA intrastrand cross-linker, (4) hydroxyurea, an inhibitor of DNA synthesis and (5) UV-C, which causes mainly photoproducts. The “true leaf assay” and the “UV resistance assay” are based on easily determined phenotypes as readouts. Using a set of diverse damaging agents combined with different readouts allows establishing relative sensitivity/resistance compared to a reference line, e.g. wild type, determining the most effective type of induced damage and the potential repair pathway affected.
Keywords: DNA damageMaterials and Reagents
- Arabidopsis thaliana seeds [Wild type and T-DNA insertion lines of arp6-3, swc6-1 and sensitive ku70-2 (Rosa et al., 2013) are used as examples. All mutants are in the Columbia-0 background.]
- Sodium hypochlorite (Sigma-Aldrich, catalog number: 425044 )
- Tween-80 (Sigma-Aldrich, catalog number: 4780 )
- Sterile H2O
- Solid growth medium
- Liquid plant growth medium (same as solid growth media but without agar)
- Hydroxyurea (Sigma-Aldrich, catalog number: H8627 )
- 70% ethanol
- Seed sterilization solution (see Recipes)
- Bleocin (commercial name for bleomycin) (EMD Millipore, catalog number: 203408 ) (see Recipes)
- Mitomycin C (Duchefa Biochemie BV, catalog number: M0133 ) (see Recipes)
Equipment
- Sterile hood, preferably a biological safety cabinet to avoid exposure to the genotoxins
- Bench top block shaker (e.g. Eppendorf, Thermomixer®)
- UV crosslinker (254-nm UV light bulbs, 15 watts each) (Stratagene, model: Stratalinker 2400 )
- Gamma-irradiation source (Nordion, model: Co-60- Gamma-cell 220 )
- Forceps
- Box or aluminum foil
- Petri dishes for plant culture with solid growth medium (see Notes) (round 200 x 15 mm and 55 x 15 mm, and square 100 x 100 x 15 mm)
- 1.5 ml Eppendorf tubes
- Plant growth facilities with 16-h-light/8-h-dark cycles, at 21 °C
Procedure
- Seed sterilization
- Wash the seeds in a 1.5 ml Eppendorf tube with 1 ml of the sterilization solution, shake in a block shaker at 300 rpm for 6 min at room temperature. Allow seeds to set in the bottom of the Eppendorf and remove supernatant. Wash with 1 ml of sterile water, 5 times for 5 min in the shaker. After the final wash, remove supernatant and dry overnight in the closed sterile hood, leaving the tubes open.
Note: Sterilize approximately 350 seeds per genotype and treatment. All subsequent steps involving seed or plantlet manipulation should be carried in the sterile hood.
- Wash the seeds in a 1.5 ml Eppendorf tube with 1 ml of the sterilization solution, shake in a block shaker at 300 rpm for 6 min at room temperature. Allow seeds to set in the bottom of the Eppendorf and remove supernatant. Wash with 1 ml of sterile water, 5 times for 5 min in the shaker. After the final wash, remove supernatant and dry overnight in the closed sterile hood, leaving the tubes open.
- True leaf assay
- Plate seeds in solid growth medium (≈ 75 to 100 per plate) using flame-sterilized and cooled forceps. Stratify in the dark (in a box or wrapped in aluminum foil) at 4 °C for 2 to 4 days and transfer to the growth chambers with illumination.
Note: Any other sterile plating techniques are suitable. Each plate makes one biological replicate; for each data point at least 3 biological replicates should be analyzed.
- Treatments
- Gamma-irradiation: Treat 4-day-old-seedlings with a pulse of 100 Gray (Gy) with the Co-60- gamma source, with a dose of 27 to 34 Gy/min. Return plates to standard growth conditions to recover for 6 days until analysis.
Note: 4-day-old = 4 days after transfer to the growth chambers.
- Bleocin and mitomycin C: Using flame-sterilized and cooled forceps, transfer 4-day-old seedlings to small petri dishes containing 10 ml of liquid growth medium, either without (mock) or with a drug (1 μg/ml of bleocin or 10 μg/ml of mitomycin C). After 5 days of incubation (with seedlings floating in liquid, but no shaking) in the illuminated growth chamber (9-day-old seedlings), remove the medium, wash extensively by flooding the plate 5 times with 20 ml of liquid media. Transfer seedlings to solid plates using flame-sterilized and cooled forceps. Allow seedlings to recover for 24 h before analysis.
Note: Seedlings should be carefully picked and handled, avoiding damage by the forceps. For examples of results see Figure 1A-C.
- Hydroxyurea: Plate seeds directly on plates, without or with 1 mM of the chemical, and grow at standard conditions until analysis at day 10.
- Gamma-irradiation: Treat 4-day-old-seedlings with a pulse of 100 Gray (Gy) with the Co-60- gamma source, with a dose of 27 to 34 Gy/min. Return plates to standard growth conditions to recover for 6 days until analysis.
- Analysis
At day 10, photograph the plates for reference, and score the “percentage of plants with true leaves”.
Notes:- Upon undisturbed development, 10-day-old Arabidopsis seedlings possess one pair of true leaves.
- DNA damage causes the cells in the apical meristem to stop dividing, so that no true leaves are developed (Figure 1A).
- For the quantitative assay, calculate the percentage of plants with true leaves for each treated population in relation to the number of plants with true leaves after mock treatment. This calculation separates the effect of the DNA-damaging treatment from other factors, like vigor of the mutants or environmental factors.
- Only paired and normally shaped leaves should be scored; single, or small and narrow unexpanded leaves are not considered as true leaves.
- Upon undisturbed development, 10-day-old Arabidopsis seedlings possess one pair of true leaves.
- Plate seeds in solid growth medium (≈ 75 to 100 per plate) using flame-sterilized and cooled forceps. Stratify in the dark (in a box or wrapped in aluminum foil) at 4 °C for 2 to 4 days and transfer to the growth chambers with illumination.
- UV damage assay
- Plate seeds in solid growth medium (≈ 75 to 100 per plate) using flame-sterilized and cooled forceps. Stratify in the dark (in a box or wrapped in aluminum foil) at 4 °C for 2 to 4 days and transfer to the growth chambers with illumination.
Note: Each plate makes one biological replicate; for each data point at least 3 biological replicates should be analyzed.
- Treat seedlings at day 4 of growth, with 3 kJ/m2 UV-C in a Stratalinker. Grow under standard conditions until day 12 and score the percentage of resistant plants (Figure 1D). Sensitive plants have phenotypes such as reduced global size, chlorotic cotyledons and senescence, whilst resistant plants have a normal size range and remain green.
Note: Divide the value in treated populations by the same parameter in the mock treatment, and average the biological replicates.
Before the treatment sterilize the interior of the Stratalinker with 70% ethanol and run it for 3 min with the maximum dose of UV-C. Then place plates inside and remove the lid, keeping it inside the Stratalinker. Quickly close the door, to avoid contamination and run the desired dosage of UV-C. When the treatment is finished close the plates still inside of the Stratalinker and return them to normal growth conditions.
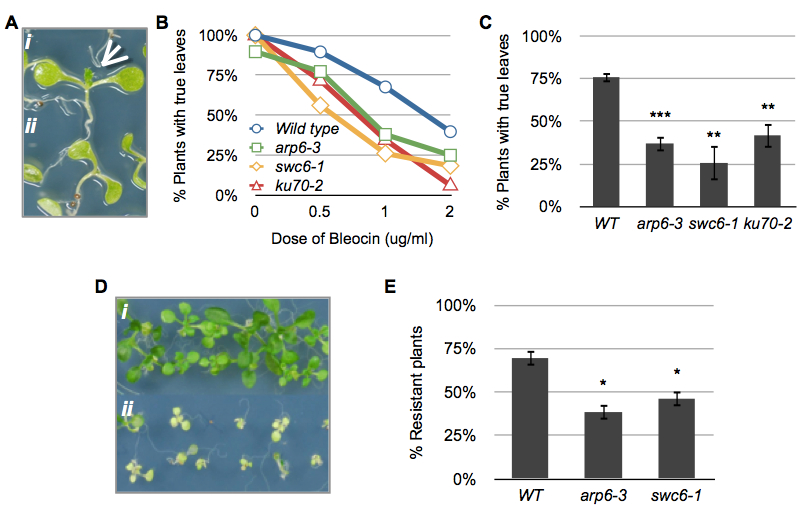
Figure 1. Examples of results. (A) Phenotypes of 10-day-old bleocin-treated seedlings with (i, arrow) and without (ii) true leaves. (B) True leaf assay showing the response of wild type, arp6-3, swc6-1 and ku70-2 to different doses of bleocin. Each plotted value corresponds to the scoring of one experiment with 60-80 seedlings. (C) True leaf assay with seedlings treated with 1 μg/ml of bleocin. The percentage of 10-d-old treated plants with true leaves was calculated in relation to mock samples. Bars depict the averages between 3 to 5 experiments with 75 to 100 seedlings per treatment. (D) Phenotypes of 12-day-old UV-C-treated resistant (i) and sensitive (ii) seedlings. (E) UV resistance assay of seedlings treated with 3 kJ/m2 of UV-C. Bars depict the averages between 4 experiments with 75 to 100 seedlings per treatment. Asterisks indicate the significance between treated and non-treated samples according to P values from unpaired t tests: ***P < 0.001 and **0.001 < P < 0.01.
- Plate seeds in solid growth medium (≈ 75 to 100 per plate) using flame-sterilized and cooled forceps. Stratify in the dark (in a box or wrapped in aluminum foil) at 4 °C for 2 to 4 days and transfer to the growth chambers with illumination.
Notes
- To better characterize the response of the different genotypes to the DNA damaging agents, generate response curves with at least 3 different concentrations of the genotoxins (example in Figure 1B): 50 to 200 Gy of gamma-irradiation, 0.5 to 2 mg/ml of bleocin, 5 to 20 of mitomycin C, 0.5 to 2 of hydroxyhurea and 1.5 to 6 kJ/m2 of UV-C.
- Always keep the plates belonging to one experiment (mock + treatment) growing side-by-side. For example, take plates for mock and gamma-irradiation treatment out of the growth chamber at the same time.
- In this experiment we used Germination Media (GM) described in Rosa et al. (2013) (detailed protocol available at http://www.gmi.oeaw.ac.at/download/gm_medium_protocol_oms.pdf). We have not tested the use of regular MS plates but have no reason to believe the media will be a determining factor as long as the plates used in each experiment come from the same batch of media.
Recipes
- Seed sterilization solution
5% sodium hypochlorite and 0.05% Tween-80
Mix 500 µl of sodium hypochlorite and 5 µl of Tween-80 in 10 ml of sterile H2O
- Bleocin (working solution: 1 μg /ml; stock: 10 mg/ml)
Dissolve 100 mg of Bleocin in 10 ml of H2O
Make 0.2 ml aliquots and keep at -20 °C until use
Avoid thawing/freezing cycles
Dilute to the desired concentration in liquid media, using disposable 50 ml Falcon tubes
- Mitomycin C (working solution: 10 μg/ml; stock: 0.5 mg/ml)
Dissolve 2 mg of Mitomycin C in 4 ml of H2O immediately before use
Dilute to the desired concentration in liquid media, using disposable 50 ml Falcon tubes
Acknowledgments
We thank the Department of Nutritional Sciences of the University of Vienna for help with the irradiation experiments and Gudrun Böhmdorfer for helpful discussions on DNA damaging agents. Work was supported by Grants GEN-AU GZ 200.140-VI/1/2006 from the Austrian Federal Ministry of Science and Research and FWF P18986-B17 from the Austrian Science Fund. This protocol was established in Rosa et al. (2013).
References
- Masson, J. and Paszkowski, J. (1992). The culture response of Arabidopsis thaliana protoplasts is determined by the growth conditions of donor plants. Plant J 2(5): 829-833.
- Rosa, M., Von Harder, M., Cigliano, R. A., Schlögelhofer, P. and Mittelsten Scheid, O. (2013). The Arabidopsis SWR1 chromatin-remodeling complex is important for DNA repair, somatic recombination, and meiosis. Plant Cell 25(6): 1990-2001.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Rosa, M. and Scheid, O. M. (2014). DNA Damage Sensitivity Assays with Arabidopsis Seedlings. Bio-protocol 4(7): e1093. DOI: 10.21769/BioProtoc.1093.
Category
Plant Science > Plant physiology > Tissue analysis
Molecular Biology > DNA > DNA damage and repair
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










