- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
In vivo Extravasation Assay
Published: Vol 4, Iss 4, Feb 20, 2014 DOI: 10.21769/BioProtoc.1051 Views: 15555
Reviewed by: Lin FangFanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Establishment of Patient-Derived Xenografts in Mice
Dongkyoo Park [...] Xingming Deng
Nov 20, 2016 15571 Views
A Murine Orthotopic Allograft to Model Prostate Cancer Growth and Metastasis
Robert M. Hughes [...] Paula J. Hurley
Feb 20, 2017 12189 Views
Qualitative in vivo Bioluminescence Imaging
Devbarna Sinha [...] Pritinder Kaur
Sep 20, 2018 10950 Views
Abstract
Tumor metastases develop when disseminated intravascular cancer cells acquire the ability to arrest by adhering to the capillary walls of distant organs, actively extravasate into their parenchyma, proliferate and establish secondary colonies. The extravasation assay described here is an in vivo technique aimed to analyze the ability of tumor cells to achieve early colonization of the lungs following tail vein injection in mice. Importantly, tumor cells need to be easily visible, therefore either they are fluorescent (e.g. expressing RFP or GFP) or they have to be pre-labelled with a fluorescent tracker prior to injection. Lungs are analyzed at different time points, experimentally determined by the researcher, depending on cell features and malignancy. Generally, an early time point is required to check equal lodging in the pulmonary vasculature for the various cells injected. At one or more later time points (from 6 to 48 h) extravasated cells dispersed in the lung parenchyma are quantitated. With our protocol extravasation is directly evaluated in the whole lungs ex vivo considering cell fluorescence. However, immunofluorescence stainings for endothelial markers and microscopic analyses of lung sections are recommended to evaluate positioning and status of tumor cells (i.e. inside, outside the vessels or associated to them; single cells or clusters). Since extravasation is not only influenced by tumor cell motility but also by their survival ability, the results obtained with this technique should be complemented with proliferation and apoptosis analyses.
Keywords: MetastasisMaterials and Reagents
- Tumor cells (in particular, this protocol was set up for A375P, MC-1, MA-2, WK-Mel and B16 melanoma cells and for MDA-MB-231 breast cancer cells)
- CD1 athymic nude female mice (6 to 9 week-old)
- Phosphate buffered saline (PBS)
- Trypsin-EDTA (Life Technologies, Gibco®, catalog number: 15400-054 )
- DMSO (Sigma-Aldrich, catalog number: 41648 )
- CellTrackerTM Orange CMRA (Life Technologies, Molecular Probes®, catalog number: C34551 ); alternatively tumor cells themselves are required to express red fluorescent protein (RFP)
- 4% Paraformaldehyde (PFA) in PBS
- 30% Sucrose in PBS
- Freezing resin (Killik, Bio-Optica Milano SpA, catalog number: 05-9801B )
- Isopentane
- Dry ice
- Anti-CD31 antibody (BD Biosciences, catalog number: 550274 ) or alternatively anti-Von Willebrandt Factor antibody (Dako, catalog number: A0082 ) to stain blood vessels
- Anti-rat or anti-rabbit Alexa-Fluor-488 secondary antibody (Life Technologies, Molecular Probes®, catalog numbers: A11006 and A11008 )
- FBS (Beromed, GmbH)
- Serum-free medium (see Recipes)
- Complete medium (Life Technologies, Gibco®) (see Recipes)
Equipment
- Cell culture set up, laminar flow hood, centrifuge, cell counting chamber
- 50 ml tubes
- P1000 and P200 tips
- Mouse housing and handling set up
- U-100 syringe with 29 G x 1/2 inch needle
- Scissor-handle straight Kelly hemostatic forceps
- Fluorescence stereomicroscope (e.g. Leica Microsystems, model: MZ16F )
- Cryostat
- Fluorescence microscope (e.g. ZEISS AxioObserver with ApoTome Module)
- Burker counting chamber (Marienfeld-superior)
Software
- Image analysis software (e.g. ImageJ, http://rsbweb.nih.gov/ij/)
Procedure
- Remove the medium from dishes or wells containing 80-90% confluent monolayers of tumor cells (either transfected/treated according to your purposes or not), wash the cells with PBS and detach them by incubating with 1 ml 0.05% trypsin-EDTA for 3-8 min; stop trypsinization with complete medium.
- (Only for non fluorescent cells) Spin down the cells at 800 rpm/100 rcf for 3 min and resuspend the pellet in 3-4 ml of serum-free medium in 50 ml tubes.
- (Only for non fluorescent cells) Resuspend the entire lyophilized CellTrackerTM Orange CMRA in 9.1 μl DMSO and add 3-4 μl CellTrackerTM Orange CMRA (1:1,000 dilution) to the cells in each tube. Incubate 35-45 min at 37 °C. During the incubation gently rock the tube every 10-15 min.
- (Only for non fluorescent cells) Spin down the cells at 800 rpm/100 rcf for 5 min and resuspend the pellet in 15-25 ml of PBS, depending on the initial confluency. The cells will appear pink/red due to the incorporation of the tracker.
- Count the cells with a Burker counting chamber.
- Prepare cell mixtures for injection. You will inject 1 to 1.7 x 106 cells in a volume of 200 μl PBS per mouse. The number of cells to be injected per mouse should be previously determined by the researcher in a pilot experiment and depends mainly on cell type, size, aggressiveness and on the propensity to form cell clusters. For example we inject 1 x 106 B16 cells, which are extremely metastatic and are prone to form big clusters and 1.6 x 106 A375P cells which are smaller and less metastatic.
- Spin down (at 800 rpm/100 rcf for 3 min) the proper number of cells needed for your experiment according to the number of mice you will inject. Resuspend the cells in PBS by pipetting the cell suspension up and down several times with P1000 and P200 tips and maintain the tube on ice until injection. To avoid cell cluster formation, which is potentially dangerous for the mice and could alter the cell behavior during extravasation and disturb your analyses, it is essential that the cells are in homogeneously single cell status.
- Warm up the CD1 nude mice under a red light lamp for some minutes to obtain vasodilation for easier processing. Inject 200 μl of cell mixture (1.4 to 1.7 x 106 cells) per mouse in the lateral tail vein with a U-100 syringe with 29 G x 1/2 inch needle.
- Analyze the mouse lungs (as indicated in steps 10-14) at an early time point after injection as a "loading control" (10 min to 2 h) and at later time points (12 h to 72 h) to detect extravasated cells. For melanoma (e.g. A375P, WK-Mel) and breast cancer (e.g. MDA-MB-231) cells we performed analyses at 2 and 48 h respectively (see Figure 1).
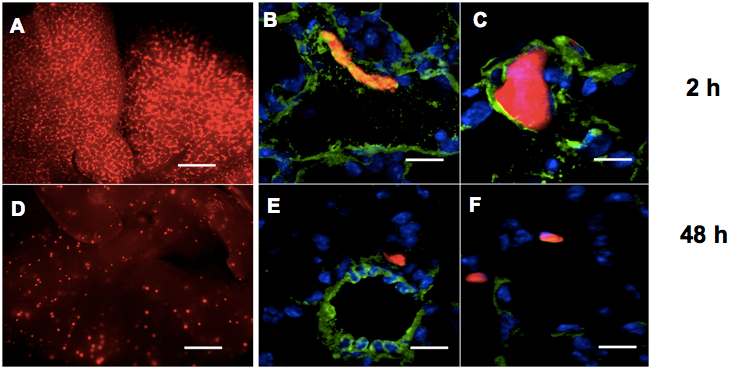
Figure 1. Lung extravasation macroscopic and microscopic analyses. A, D. Representative pictures of whole lungs 2 h (A) or 48 h (D) following tail vein injections of CMRA-labelled (red) melanoma cells in nude mice. Bar: 500 μm. B, C, E, F. Representative fields of murine lung sections 2 h (B, C) or 48 h (E, F) post-injections of CMRA-labelled (red) melanoma cells, stained for Von Willebrandt factor to highlight blood vessels (green) and counter-stained with DAPI (blue). Bar: 30 μm - Sacrifice the mice by cervical dislocation and dissect the animal chest to expose the trachea with sharp-edged microdissection tweezers and scissors, by eliminating all the connectival tissues surrounding the trachea, until the white cartilage rings are well visible. It is worth to proceed very gently to avoid trachea perforation. Make a longitudinal cut in the diaphragm (but without completely opening the chest) to allow the lungs to inflate. Clamp the trachea the most cranially you can with hemostatic forceps and gently inject 0.8-1 ml ice cold 4% PFA inside the trachea with a U-100 syringe with 29 G x 1/2 inch needle, with the needle parallel to the trachea. You will clearly see the lungs inflating. Complete the dissection and extract the lungs.
- Wash briefly the whole lungs in clean PBS to eliminate blood and fix them in 8-10 ml of 4% PFA inside 15 ml tubes at 4 °C for 2-3 h. Do not exceed 5 h otherwise a fluorescent background will disturb your analyses.
- Take photos (at least 3 fields/lung) of the whole lungs under a fluorescence stereomicroscope. In particular, rinse well the lungs in clean PBS and eliminate the excess liquid on a paper towel, then put the entire organ in a common cell culture petri dish (without the lid) under the microscope. Cells will appear as several red fluorescent clusters disposed along lung vasculature at early time points and as dispersed small dots at late time points, as you can observe in Figure 1A and 1D.
- Perform a quantitative and statistical analysis of the extravasated cells by counting the number of cells and/or calculating the area of red fluorescence in the photos, by means of an image analysis software (e.g. ImageJ, http://rsbweb.nih.gov/ij/).
- To verify the precise localisation of the tumor cells within the lungs, microscopic analyses should be performed.
- Wash extensively the fixed lungs in PBS and dehydrate them in a 30% sucrose solution for at least 24 h at 4 °C.
- Include the lungs in a freezing resin inside home-made tinfoil molds on dry-ice-cold (about -50 °C) isopentane and store them at -80 °C.
- Cryostat-cut the specimens in 6 μm thick sections on charged slides.
- For immunofluorescent stainings, thaw the slides, rinse them in PBS and draw a hydrophobic barrier around the specimen by using a PAP Pen.
Note: Optionally, you may want to post-fix the specimens in acetone for 5-8 min to reduce the risk of tissue loss, but avoid the use of formaldehyde-derived fixatives).- Samples are then blocked in 5% bovine serum albumin (BSA) for 1 h at room temperature (RT).
- Incubated with anti-CD31 or anti-Von Willebrandt Factor antibody (1:100 or 1:50 dilution, respectively) for 1 h at RT or overnight at 4 °C, rinsed 3 times in PBS.
- Incubated with anti-rat or anti-rabbit Alexa-Fluor-488 (green) secondary antibody (1:500 dilution) for 45 min at RT.
- Counterstained with a nuclear marker such as DAPI and finally rinsed well in PBS and protected by mounting a coverslip.
- Examine specimens using a standard fluorescence microscope. Representative images are shown in Figure 1B-F.
- Samples are then blocked in 5% bovine serum albumin (BSA) for 1 h at room temperature (RT).
- Wash extensively the fixed lungs in PBS and dehydrate them in a 30% sucrose solution for at least 24 h at 4 °C.
Recipes
- Serum-free medium
DMEM GlutamaxTM supplemented with 1 mM sodium pyruvate, 25 mM HEPES (pH 7.4) and 100 μg/ml gentamicin - Complete medium
DMEM GlutamaxTM supplemented with 10% heat-inactivated FBS, 1 mM sodium pyruvate, 25 mM HEPES (pH 7.4), 100 μg/ml gentamicin (for MDA-MB-231) and with 1x MEM Non-Essential Amino Acids (for WK-Mel and B16) and with 1x MEM Non-Essential Amino Acids and 1x MEM Vitamin Solution (for A375P, MC-1 and MA-2)
Acknowledgments
This method was adapted from Casazza et al. (2011). We acknowledge Andrea Casazza for suggestions during the development of the protocol. We also thank Francesca Orso for useful discussion. Our work is supported by grants from the Compagnia San Paolo (2008.1054/DT), PRIN 2008/DT, AIRC 2010 (IG10104/DT), and FIRB giovani 2008 (RBFR08F2FS-002/FO). Elisa Penna is a FIRC fellow (2012–2014).
References
- Casazza, A., Finisguerra, V., Capparuccia, L., Camperi, A., Swiercz, J. M., Rizzolio, S., Rolny, C., Christensen, C., Bertotti, A., Sarotto, I., Risio, M., Trusolino, L., Weitz, J., Schneider, M., Mazzone, M., Comoglio, P. M. and Tamagnone, L. (2011). Sema3E-Plexin D1 signaling drives human cancer cell invasiveness and metastatic spreading in mice. J Clin Invest 121(7):2945.
- Penna, E., Orso, F., Cimino, D., Vercellino, I., Grassi, E., Quaglino, E., Turco, E. and Taverna, D. (2013). miR-214 coordinates melanoma progression by upregulating ALCAM through TFAP2 and miR-148b downmodulation. Cancer Res 73(13): 4098-4111.
- Penna, E., Orso, F., Cimino, D., Tenaglia, E., Lembo, A., Quaglino, E., Poliseno, L., Haimovic, A., Osella-Abate, S., De Pitta, C., Pinatel, E., Stadler, M. B., Provero, P., Bernengo, M. G., Osman, I. and Taverna, D. (2011). microRNA-214 contributes to melanoma tumour progression through suppression of TFAP2C. EMBO J 30(10): 1990-2007.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Penna, E. and Taverna, D. (2014). In vivo Extravasation Assay. Bio-protocol 4(4): e1051. DOI: 10.21769/BioProtoc.1051.
Category
Cancer Biology > Invasion & metastasis > Animal models > Cell invasion
Cell Biology > Cell movement > Cell motility
Cell Biology > Cell imaging > Fixed-tissue imaging
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











