- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Detection of piggyBac-mediated Transposition by Splinkerette PCR in Transgenic Lines of Strongyloides ratti
Published: Vol 4, Iss 1, Jan 5, 2014 DOI: 10.21769/BioProtoc.1015 Views: 15208
Reviewed by: Fanglian He

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
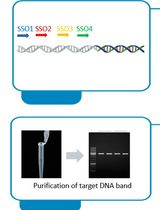
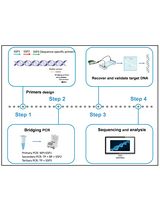
Implementation of Fusion Primer-Driven Racket PCR Protocol for Genome Walking
Yinwei Gu [...] Haixing Li
Dec 5, 2025 985 Views
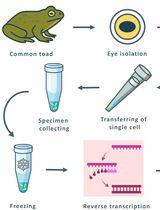
Combining Suction-Pipette Spectral Identification With Single-Cell RT-PCR to Make Differential Analyses of Amphibian Red and Green Rods
Olga V. Chernyshkova [...] Michael L. Firsov
Mar 5, 2026 83 Views
Abstract
Splinkerette PCR (spPCR) is a newly developed and efficient method to ascertain and characterize genomic insertion sites of transgenes. The method described in this protocol was successfully applied to confirm piggyBac transposon-mediated integration of transgenes into chromosomes of the parasitic nematode Strongyloides ratti. This work is described in detail in Shao et al. (2012) and presented here in a simplified diagram (Figure 1). Using this method, chromosomal loci of integration were determined based on target site and 5’- and 3’ flanking sequences. Therefore, spPCR can be a useful method to confirm integrative transgenesis in functional genomic studies of parasitic nematodes. Potter and Luo (2010) contains a protocol for use of spPCR to detect and map piggyBac transposon-mediated chromosomal integrations in Drosophila, and was the source of our method for Strongyloides. The splinkerette- and piggyBac-specific oligos described in that reference could be used without modification in Strongyloides. For interested readers, a general review of the biology of parasitic nematodes in the genus Strongyloides may be found in Viney and Lok (2007), and a methods-based article on S. stercoralis as an experimental model, with information on transgenesis, may be found in Lok (2007).
Keywords: Strongyloides
Figure 1. Diagrammatic representation of protocol for mapping transgene integrations in Strongyloides by splinkerette PCR (adapted from Potter and Luo, 2010)
Materials and Reagents
- Free-living adult worms
- Genomic DNA extraction
Gentra Puregene Tissue Kit (QIAGEN), including - Enzymes for digestion and treatment
Restriction enzymes include BstY I, BamH I and Bgl II (New England Biolabs)
Others include Shrimp Alkaline Phosphatase (United State Biological, catalog number: P4071-05 ) and Exonuclease I (New England Biolabs, catalog number: M0293S ) - Ligation reagents
10x Ligase buffer and T4 Ligase (New England Biolabs, catalog number: M0202S ) - PCR reagents
5x Phusion High-Fidelity Buffer, Phusion HF DNA Polymerase (New England Biolabs, catalog number: M0530S ) - Oligonucleotides and primers
The oligonucleotides detailed in Table 1 must be synthesized
All sequences are from Potter and Luo (2010), Table S13 - TE buffer (New England Biolabs, catalog number: E6293 )
- 10x NEB Buffer 2
- 1% agarose gel
Table 1. Oligonucleotides and primers for splinkerette PCR mapping of piggyBac transposon-mediated transgene integrationsOligo or Primer Sequence SPLNK-BOT 5’-CGAAGAGTAACCGTTGCTAGGAGAGACCGTGGCTGAATGAGACTGGTGTCGACACTAGTGG-3’ SPLNK-GATC-TOP 5’-GATCCCACTAGTGTCGACACCAGTCTCTAATTTTTTTTTTCAAAAAAA-3’ SPLNK#1 5’-CGAAGAGTAACCGTTGCTAGGAGAGACC-3’ SPLNK#2 5’-GTGGCTGAATGAGACTGGTGTCGAC-3’ 3’SPLNK-PB#1 5’-GTTTGTTGAATTTATTATTAGTATGTAAG-3’ 5’SPLNK-PB#1 5’-ACCGCATTGACAAGCACG-3’ 3’SPLNK-PB#2 5’-CGATAAAACACATGCGTC-3’ 5’SPLNK-PB#2 5’-CTCCAAGCGGCGACTGAG-3’ 3’SPLNK-PB-SEQ 5’-ACGCATGATTATCTTTAAC-3’ 5’SPLNK-PB-SEQ 5’-CGACTGAGATGTCCTAAATGC-3’
Equipment
- PCR Thermal Cyclers (Mastercycler Personal, Eppendorf)
- Gel electrophoresis
- Gel documentation (FOTODYNE, model: FOTO/Analyst FX )
Procedure
- Collect stably transformed free-living adult Strongyloides from charcoal cultures of fresh host feces incubated at 22 °C for 48 h (Lok, 2007). Extract genomic DNA (gDNA) from ~500 free-living adult worms using the Gentra Puregene Tissue Kit. Parasites do not need to be homogenized. Isolated worms are pelleted at 14,000 x g and then substituted for the “ground tissue” in step 2, followed by step 2b of the manufacturer’s protocol accompanying the Gentra Puregene Tissue Kit, using reagent volumes recommended for tissue samples of 5-10 mg. Resuspend extracted gDNA in 50 μl TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0) to yield ~ 25 ng/μl.
- Digest Genomic DNA (~100 ng) with BstYI, BamHI, and Bgl II for 2 h in a total volume of 15 μl for each reaction (Figure 1A). Follow manufacturer’s instructions for reaction conditions and inactivation steps for each restriction enzyme.
- Add 5 μl of 100 μM SPLINK-BOT oligonucleotide, 5 μl 100 μM SPLINK-GATC-TOP oligonucleotide and 10x NEB Buffer 2 in a total reaction volume of 100 μl. Heat to 95 °C for 3 min then cool on bench to room temperature to anneal splinkerette oligonucleotides. Ligate digested gDNA with GATC sticky ends to annealed splinkerette oligonucleotides using T4 ligase for 8 h at 16 °C in a total volume of 30 μl (Figure 1B).
- Two rounds of spPCR are conducted following ligation of genomic DNA restriction fragments to splinkerette oligo nucleotides.
- For the first round of spPCR, combine primer SPLINK#1, which targets the ligated splinkerette oligonucleotides, with either primer 3’SPLNK-PB#1 or primer 5’SPLNK-PB#1. 3’SPLNK-PB#1 and 5’SPLNK-PB#1 target the 3’ and 5’ ends of the transgene sequences, respectively. Carry out the amplification with Phusion Polymerase and 5-20 ng of splinkerette-ligated genomic DNA as template. Thermal cycling conditions are:
- 98 °C for 1 min
- 98 °C for 20 sec
- 55 °C for 15 sec
- 72 °C for 2 min
- 29 more cycles from ii to iv
- a final extension at 72 °C for 10 min.
- 98 °C for 1 min
- The second round of spPCR is nested PCR carried out with primer SPLNK#2 paired with either primer 3’SPLNK-PB#2 or primer 5’SPLNK-PB#2 and a 1:20 dilution of spPCR products from the first-round spPCR as template. Carry out the second-round amplification with Phusion High-Fidelity DNA Polymerase using the same thermal cycling conditions as specified for first-round spPCR. Analyze all PCR products by 1% agarose gel electrophoresis.
- Incubate the products of second-round spPCR with 1 μl Shrimp Alkaline Phosphatase (1 U/ml) and 1 μl Exonuclease I (20,000 U/ml) at 37 °C for 2 h, and then at 80 °C for 15 min (Figure 1C).
- Sequence the treated PCR products directly using primers 3’SPLNK-PB-SEQ and 5’SPLNK-PB-SEQ, targeting the 3’ and 5’ ends of the transgene sequence, respectively. Identify chromosomal locations of integrations based on generated sequences and matching fragments in the genome database (Figure 1D, 1E).
- For the first round of spPCR, combine primer SPLINK#1, which targets the ligated splinkerette oligonucleotides, with either primer 3’SPLNK-PB#1 or primer 5’SPLNK-PB#1. 3’SPLNK-PB#1 and 5’SPLNK-PB#1 target the 3’ and 5’ ends of the transgene sequences, respectively. Carry out the amplification with Phusion Polymerase and 5-20 ng of splinkerette-ligated genomic DNA as template. Thermal cycling conditions are:
Acknowledgments
This protocol details methods originally published in Shao et al. (2012). This work was funded by grant number AI050668 from the US National Institutes of Health.
References
- Lok, J. B. Strongyloides stercoralis: a model for translational research on parasitic nematode biology (February 17, 2007), WormBook, ed. The C. elegans Research Community, WormBook, doi/10.1895/wormbook.1.134.1, http://www.wormbook.org.
- Potter, C. J. and Luo, L. (2010). Splinkerette PCR for mapping transposable elements in Drosophila. PLoS One 5(4): e10168.
- Shao, H., Li, X., Nolan, T. J., Massey, H. C., Jr., Pearce, E. J. and Lok, J. B. (2012). Transposon-mediated chromosomal integration of transgenes in the parasitic nematode Strongyloides ratti and establishment of stable transgenic lines. PLoS Pathog 8(8): e1002871.
- Viney, M. E. and Lok, J. B. Strongyloides spp. (May 23, 2007), WormBook, ed. The C. elegans Research Community, WormBook, doi/10.1895/wormbook.1.141.1, http://www.wormbook.org.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Shao, H. and Lok, J. B. (2014). Detection of piggyBac-mediated Transposition by Splinkerette PCR in Transgenic Lines of Strongyloides ratti. Bio-protocol 4(1): e1015. DOI: 10.21769/BioProtoc.1015.
- Shao, H., Li, X., Nolan, T. J., Massey, H. C., Jr., Pearce, E. J. and Lok, J. B. (2012). Transposon-mediated chromosomal integration of transgenes in the parasitic nematode Strongyloides ratti and establishment of stable transgenic lines. PLoS Pathog 8(8): e1002871.
Category
Microbiology > Microbial genetics > DNA > Transposon
Molecular Biology > DNA > PCR
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link