- EN - English
- CN - 中文
Protocol for Microfluidic System to Automate the Preparation and Fractionation of the Nucleic Acids in the Cytoplasm Versus Nuclei of Single Cells
自动制备和分级分离单细胞细胞质与细胞核中核酸的微流体系统方法
(*contributed equally to this work) 发布: 2016年06月20日第6卷第12期 DOI: 10.21769/BioProtoc.1844 浏览次数: 9371
评审: Samik BhattacharyaAnonymous reviewer(s)

相关实验方案
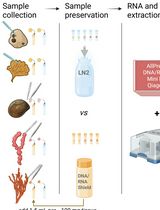
深海无脊椎动物组织保存的比较方案:DNA/RNA Shield 与液氮在高质量核酸双重提取中的应用对比
Ana S. Gomes [...] Olivier Laroche
2025年11月20日 1389 阅读
Abstract
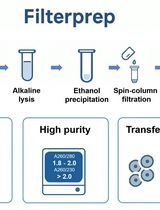
This protocol describes the extraction, fractionation, and recovery of cytoplasmic nucleic acids (e.g., cytoplasmic RNA) versus nucleic acids in the cell nucleus (including genomic DNA, gDNA) from single cells with a microfluidic system. The method enables independent, sequence-specific analyses of these critical markers (Kuriyama et al., 2015). The system uses a microfluidic chip with a simple geometry and four end-channel electrodes, and completes the entire process in less than 5 min, including lysis, purification, fractionation, and delivery to two output reservoirs: One for the nucleus (including gDNA and nuclear RNA) and one for cytoplasmic RNA. Each reservoir then contains high quality and purity aliquots with no measurable cross-contamination of cytoplasmic RNA versus nucleic acids in nucleus. As described here, our protocol focuses on the analysis of cytoplasmic RNA versus gDNA from the nucleus. We have tested this protocol with mouse and human cells but not with walled cells such as plant cells.
Keywords: Single cell analysis (单细胞分析)Materials and Reagents
- Microfluidic chip fabrication
Fabricate a microfluidic device (Figure 1) from polydimethylsiloxane (PDMS) (Dow Corning, Sylgard® 184 Silicone Elastomer Kit) and glass slides by using soft lithography. The nominal channel width and depth of the microfluidic device are 90 μm and 35 μm, respectively. The basic design is similar to the microchannel used by Shintaku et al. (2014) except for a downstream branch channel. We have made available the CAD file for the device in the supporting information.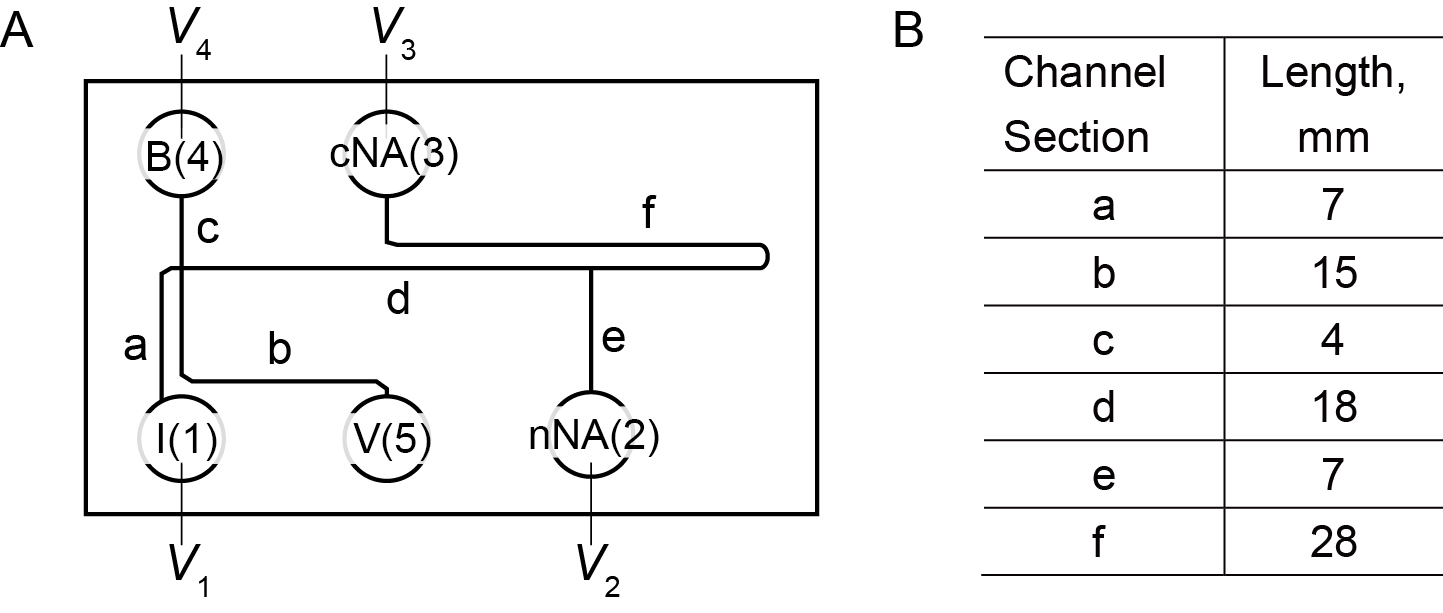
Figure 1. Schematic of channel geometry. A. The letters (a to f) in the figure identify the various channel sections. Vi (for I = 1 to 5) refer to the voltages applied to platinum electrodes placed in respective reservoirs. The numbers also refer to the reservoirs, e.g., 1 = I, 2 = nNA, 3 = cNA, 4 = B, and 5 = V. B. The lengths of channel sections in the chip. - Reagents for extraction
Prepare following solutions in UltraPure DNase-/RNase-free deionized (DI) water (Thermo Fisher Scientific, InvitrogenTM, catalog number: 10977 ). Prepare buffer solutions under DNA/RNA free and DNase/RNase free environment (e.g., clean room conditions)- Cells (A20, mouse B cell lymphoma)
- Sodium hydroxide (NaOH) (Sigma-Aldrich, catalog number: S8045 )
- TritonTM X-100 (Sigma-Aldrich, catalog number: X100 )
- Tris (Sigma-Aldrich, catalog number: 93362 )
- Hydrochloric acid (HCl) (Sigma-Aldrich, catalog number: 258148 )
- Polyvinylpyrrolidone (PVP) (Sigma-Aldrich, catalog number: 437190 )
- HEPES (Sigma-Aldrich, catalog number: PHG0001 )
- Sucrose (Sigma-Aldrich, catalog number: S0389 )
- Leading electrolyte (LE) (see Recipes)
- Trailing electrolyte (TE) (see Recipes)
- Cell suspension buffer (see Recipes)
- Cells (A20, mouse B cell lymphoma)
- Reagents for RT-qPCR and qPCR
The reagents described here are for an experiment similar to Kuriyama et al. (2015) wherein we performed quantitation of gDNA for the nucleus and cytoplasmic RNA for the cytoplasm. However, we note the general lysing, fractionation, and preparation protocol described here is well applicable to other studies (e.g., analysis of nuclear versus cytoplasmic fractions of RNA from single cells).- TaqMan® RT-PCR mix (2x)*
- TaqMan® gene expression assay
- TaqMan® RT enzyme mix (40x)*
- TaqMan® RNA-to-CTTM 1-step kit (Thermo Fisher Scientific, Applied BiosystemsTM, catalog number: 4392653 )
- Primer set (forward primer, reverse primer) (TaqMan® probe)
- DI water
- RT-qPCR master mix for cytoplasmic RNA analysis for two reactions (see Recipes)
- qPCR master mix for gDNA analysis for one reaction (see Recipes)
*Note: TaqMan® RNA-to-CTTM 1-step kit contains both reagents.
- TaqMan® RT-PCR mix (2x)*
Equipment
- Microscope
A phase contrast microscope fitted with a CCD camera enables monitoring the lysing and cytoplasmic nucleic acid fraction focusing via isotachophoresis (ITP) [see on-line movies by Garcia-Schwarz et al. (2012) for a typical on-chip ITP experiment.]. It also helps monitor migration of nucleus and its fractionation from the cytoplasmic fraction in the ITP zone. Avoid using fluorescence-based visualization, e.g., intercalation dye, which may interfere specificity and stringency of downstream assay. - High voltage sequencer
A voltage sequencer (e.g., LabSmith, Inc., model: HVS448-3000D) automates the lysis, extraction, and fractionation via ITP.
Note: A current measurement helps monitor the ITP zone migration and fractionation of the nucleus. - Send the voltage sequence to the voltage sequencer.
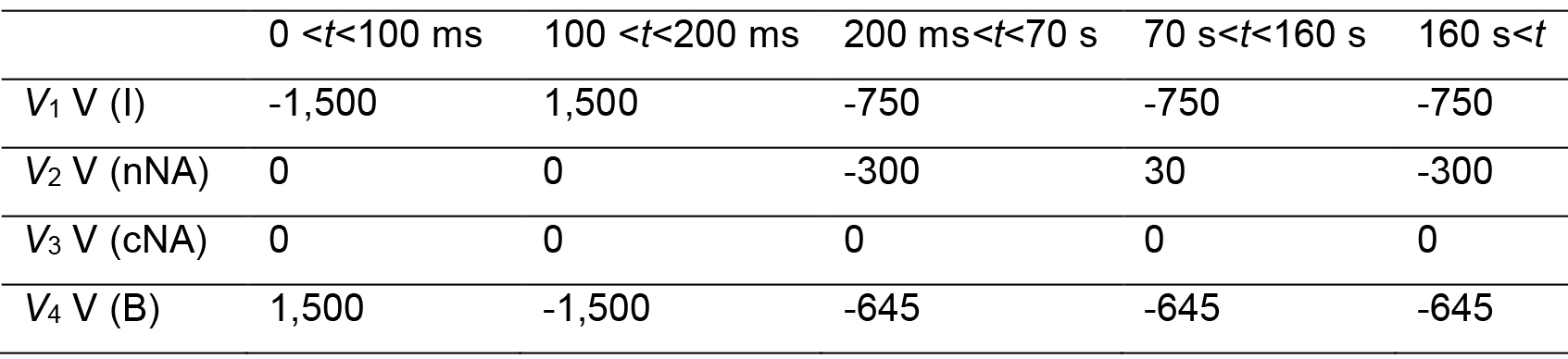
- Connect the voltage outputs to platinum wire electrodes that will be placed to the reservoirs.
Procedure
文章信息
版权信息
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
如何引用
Kuriyama, K., Shintaku, H. and Santiago, J. G. (2016). Protocol for Microfluidic System to Automate the Preparation and Fractionation of the Nucleic Acids in the Cytoplasm Versus Nuclei of Single Cells. Bio-protocol 6(12): e1844. DOI: 10.21769/BioProtoc.1844.
分类
细胞生物学 > 单细胞分析 > 微流体
分子生物学 > DNA > DNA 提取
分子生物学 > RNA > RNA 提取
您对这篇实验方法有问题吗?
在此处发布您的问题,我们将邀请本文作者来回答。同时,我们会将您的问题发布到Bio-protocol Exchange,以便寻求社区成员的帮助。
Share
Bluesky
X
Copy link