- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
59Fe Uptake Assays in Paracoccidioides Species
Published: Vol 6, Iss 18, Sep 20, 2016 DOI: 10.21769/BioProtoc.1930 Views: 8094
Reviewed by: Arsalan DaudiBelen SanzSoazig Le Guyon

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
I Plate-based Assay for Studying How Fungal Volatile Compounds (VCs) Affect Plant Growth and Development and the Identification of VCs via SPME-GC-MS
Wenzhao Wang [...] Seogchan Kang
Feb 20, 2019 8323 Views

Intracellular cAMP Measurements in Candida albicans Biofilms
Liuliu Jiang [...] Xin Wei
Dec 5, 2019 4686 Views
Determination of the Cellular Ion Concentration in Saccharomyces cerevisiae Using ICP-AES
Louise Thines [...] Pierre Morsomme
Aug 20, 2020 3450 Views
Abstract
Iron is an essential micronutrient required for virtually all organisms. This fact is related to the ability of the transition metal to exist in two oxidation states, the reduced ferrous (Fe2+) and the oxidized ferric (Fe3+). Given the relative availability of aqueous iron (the element which constitutes ~5% of the earth’s crust) one is not surprised that iron is the most common prosthetic element in biology. Usually, fungi can uptake iron through receptor-mediated internalization of a siderophore or heme, and/or reductive iron assimilation (RIA) (Kosman, 2013). In this way, the uptake of iron in the absence or presence of the reducing agent ascorbic acid can be investigated by 59Fe uptake assays, as previously described (Eide et al., 1992). In the presence of ascorbic acid, the reductive-independent 59Fe uptake route is investigated. On the other hand, in the absence of ascorbic acid, the reductive-dependent 59Fe uptake route is stimulated. Using this strategy for the human pathogenic fungus Paracoccidioides species, the results showed that iron uptake by Pb01 in the absence of ascorbic acid was low, unlike what was observed for Pb18. These results suggest that only in Pb18 the iron uptake pathway is coupled to a ferric reductase (Bailão et al., 2015). In this protocol, we describe how to perform 59Fe uptake assays in Paracoccidioides species.
Keywords: Reductive iron assimilation pathwayMaterials and Reagents
- Conical tube
- Wide-mouth screw-cap high-density polyethylene (scintillation) vials
- 13 x 100 mm test tubes
- Petri dish
- Filter type A/C 25 mm glass fiber filters from Pall Gelman
- 12 x 75 mm glass test tubes
- Wooden applicator (a thin wooden stick that is used in this protocol to put the filters containing radioactive cells inside the test tubes that will be read by the G counter)
- Disposable vacuum filtration system (TPP, catalog number: 99950 )
- Paracoccidioides spp. cells
- Culture media
- Liquid Brain Heart Infusion Broth (BHI) (Sigma-Aldrich, catalog number: 53286 ) with 4% glucose
- Liquid Synthetic Complete (SC medium) (see Recipes)
- Deionized water
- Yeast Nitrogen Base (YNB) without amino acids (BD, DifcoTM, catalog number: 291940 )
- Glucose (VWR, catalog number: 101175P )
- Amino acids (Sigma-Aldrich)
ADE, TRP, HIS, ARG, MET, TYR, ISOLEU, LYS, PHE, GLU. ACID, ASP. ACID, VAL, THR, SER, URA, LEU - Trypan blue solution, 0.4% for microscopy (Sigma-Aldrich, catalog number: 93595 )
- Ethylenediamine tetraacetic acid (EDTA) (Sigma-Aldrich, catalog number: E9884 )
- Succinic acid (Sigma-Aldrich, catalog number: 398055 )
- Tris
- NaOH
- Analar glucose
- MES buffer (Sigma-Aldrich, catalog number: M3671 )
- Sodium citrate
- 59Fe (Perkin-Elmer, catalog number: NEZ037001MC )
- 1 M ascorbic acid (0.176 g in 1 ml sterilized water)
- Nitric acid (HNO3)
- Bathophenanthrolinedisulfonic acid (BPS) (Sigma-Aldrich, catalog number: B1375 )
- Iron (III) chloride (Sigma-Aldrich, catalog number: 157740 )
- SC complete media (see Recipes)
- Amino acids solution (see Recipes)
- Amino acid mix stock (see Recipes)
- 59Fe stock solutions (see Recipes)
- 10x quench buffer (see Recipes)
- Citrate uptake buffer (see Recipes)
- Acid-wash glassware treatment (see Recipes)
Equipment
- Shaker
- Hemocytometer
- Microfine forceps
- Glassware (erlenmeyers, bottle with blue cap, beakers, cylinders)
- Magnetic stirrer
- pH meter
- Vortex
- G counter (LKB Wallac Compu Gamma) with plastic g counting tube carriers
- Vacuum filtration manifold (Merck Millipore, model: 1225 )
- Source of vacuum, either standard lab pump or water aspirator
- Autoclave
Procedure
- Grow the Paracoccidioides spp. cells (from a stock* grown in BHI with 4% glucose medium for at least 72 h at 36 °C) in 200 ml BHI with 4% glucose medium for 72 h at 36 °C with shaking (around 300 rpm). After this, collect 5 x 106 viable cells counted in a hemocytometer with Trypan blue solution** and transfer to 30 ml of SC medium with no supplementation or supplemented with 200 μM BPS or 10 μM FeCl3 for 24 h at 36 °C with shaking.
*Note: This stock remains viable until seven days. After this, a new culture has to be initiated in tubes containing fresh BHI with 4% glucose medium.
** Note: To count the viable cells, dilute the cells in 0.9% NaCl solution 100 times (it could be two 1:10 serial dilutions). After the dilution, get 10 μl of the cells and mix with 10 μl of Trypan blue solution. This corresponds to a 1:200 final dilution. Get 10 μl of this mixture and count the cells using a hemocytometer. The viable cells will not uptake the blue dye and will stay colorless. - Collect 20 ml of cells into a conical tube by centrifugation at 1,200 x g for 10 min at 4 °C and wash cells once with 1 mM EDTA in citrate uptake buffer.
- Wash cells twice with citrate uptake buffer (pH 6.0).
- Re-suspend cells to a final volume of 15 ml with citrate uptake buffer.
- Transfer 7 ml of cells into a scintillation vial. You’ll need more than 7 ml of cells if you have more than two time points.
- Incubate cells for 15 min at 36 °C with shaking prior to addition of 59Fe.
- While shaking cells, take an aliquot (100 μl) from each scintillation vial and count the viable cells by a hemocytometer with Trypan blue solution.
Note: This step is important to data normalization at the end of the experiment, since the data will be presented as pmol 59Fe/106 viable cells/time. - For reductase-independent 59Fe uptake, add 140 μl 1 M ascorbic acid in distilled H2O to each scintillation vial (20 mM final concentration).
Note: The ascorbic acid solution should be made just prior to use. - To begin uptake, add 70 μl of 20 μM 59Fe solution to each of the scintillation vials.
- At each time point (be sure to have a zero time point), stop shaker, carefully remove the scintillation vials, and pipet 1 ml of cells into 13 x 100 mm test tubes containing 3 ml of ice-cold 1x quench buffer on ice. Do this in triplicate for each time point (Figure 1).
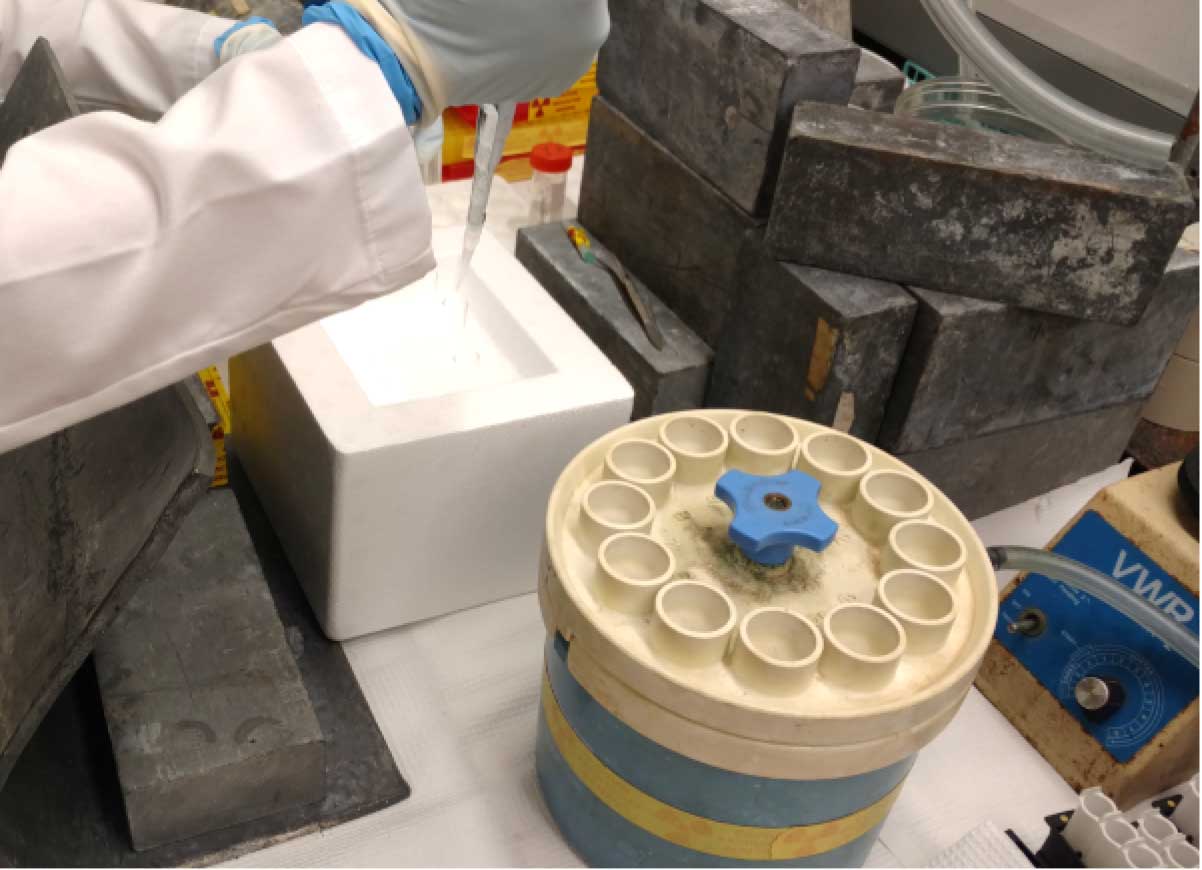

Figure 1. Quenching uptake samples. At each time point 1 ml samples are withdrawn from the uptake vials and quenched in 3 ml ice-cold quench buffer. It is important to add the quench buffer immediately after collecting 1 ml sample. - For filtration and washing of cells, pre-soak filters in 1x quench buffer in a Petri dish.
- Using the microfine forceps, carefully place 12 filters on the manifold (hold only the outer edges of the filters) (Figure 2). Place the top on the manifold and tighten it down with the blue plastic screw. Apply vacuum to the filter manifold using cold water to form a vacuum.
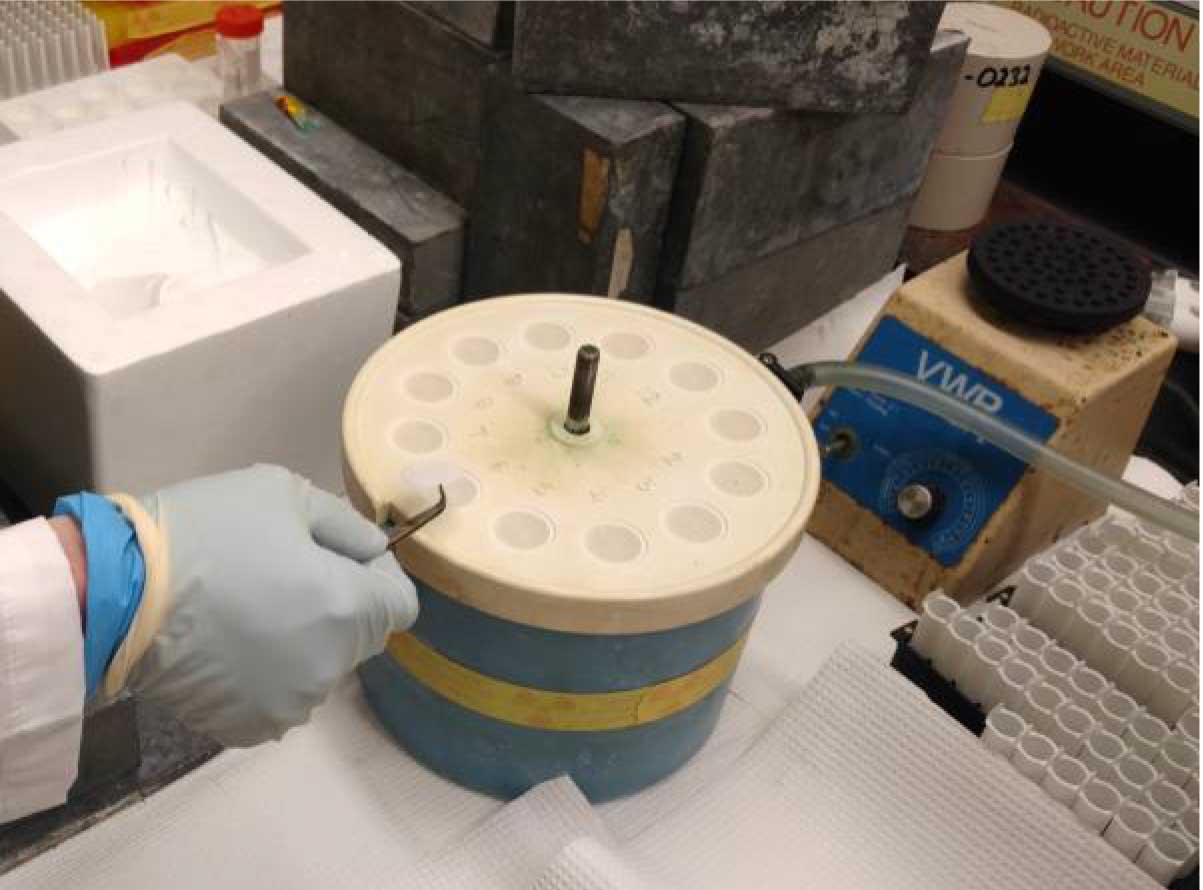

Figure 2. Placing filters on the Millipore manifold. Prior to initiating the 59Fe-uptake by your fungal samples, the filters are wetted in quench buffer and then placed on each of the manifold’s frits with the help of tweezers. The top on the manifold is then placed and tighten it down with the blue plastic screw. Apply vacuum to the filter manifold using cold water to form a vacuum. - When all the samples have been collected, wash quenched samples on the Millipore manifold (Figure 3).
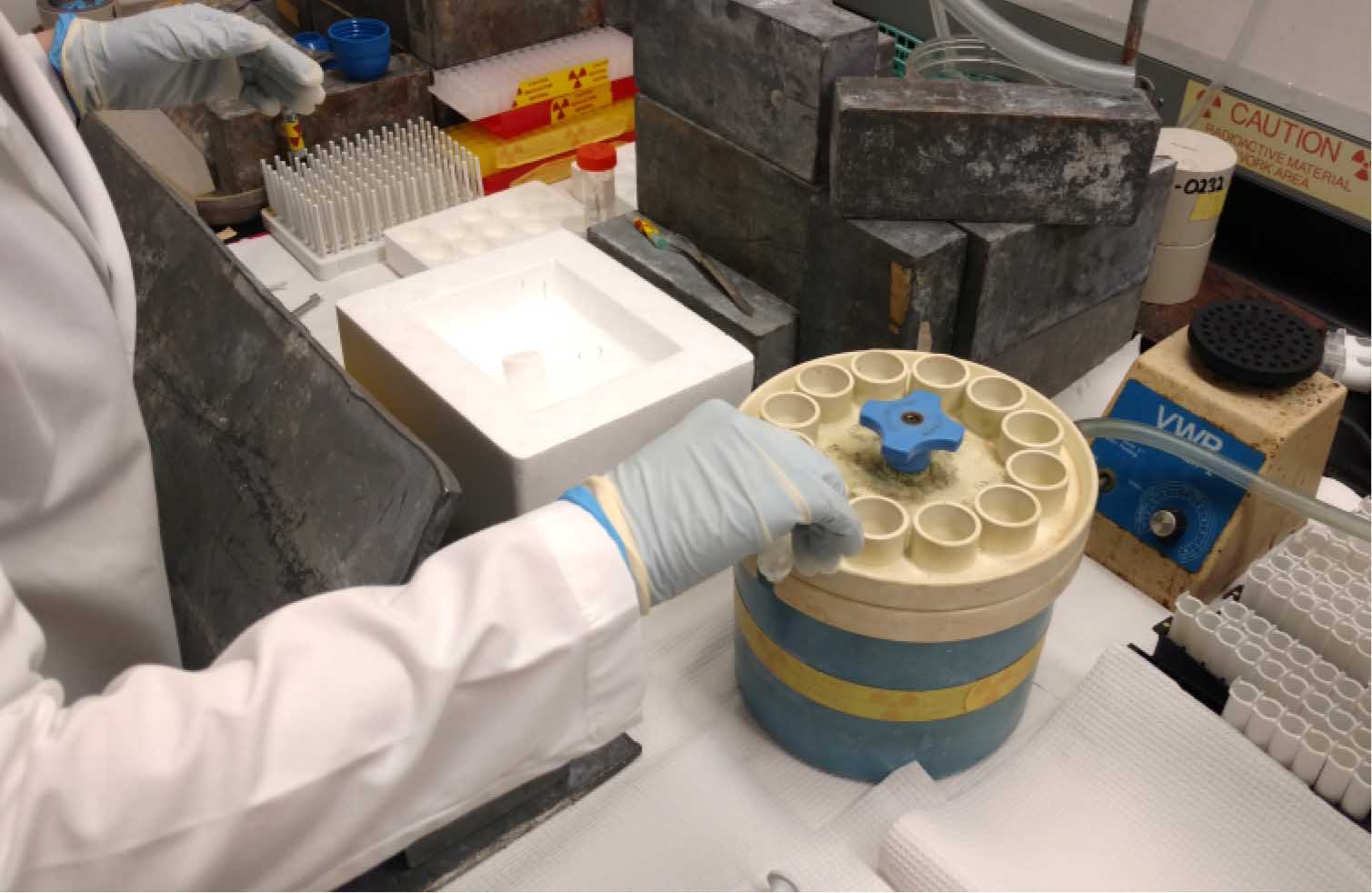

Figure 3. Filtering quenched cell samples. After collecting and quenching all of the uptake samples, they are filtered and washed on the manifold for three times. It is important to wash filters in a series of # 1-12. Do not wash filter # 1 three times and then proceed to wash # 2 three times. If collecting more than the number of wells on the manifold, the first 12 filter set is removed for counting and replaced with fresh filters to continue the washing of additional samples. - Pour samples over each filter. Rinse the test tube once with 3 ml ice-cold 1x quench buffer. Pour that over each corresponding filter. Rinse each filter three times with 3 ml ice-cold 1x quench buffer. Wash filters in a series of 1-12. For example, do not wash filter in position 1 three times and then proceed to wash the filter in position 2 three times. Leave manifold under vacuum until you have filtered and washed all samples.
- Leaving the manifold under vacuum, allow filters to dry (~30 sec) after the last wash.
- Using the forceps, remove each filter by folding it in on itself (NEVER touch the middle or top of the filter) and placing it in a 12 x 75 mm glass test tube.
- Continue with steps 11-15 until all of the samples have been processed.
- Using a wooden applicator, gently push the filters to the bottom of the test tube.
- Place the test tubes in the g counter carrier racks and count samples including 59Fe standards prepared in 12 x 75 mm test tubes. Use at least five points for standards, such as 20 μM, 10 μM, 1 μM, 0.1 μM and 0.05 μM (Figure 4).
- Dispose of radioactive waste as per institutional environmental health and safety regulations.
Representative data
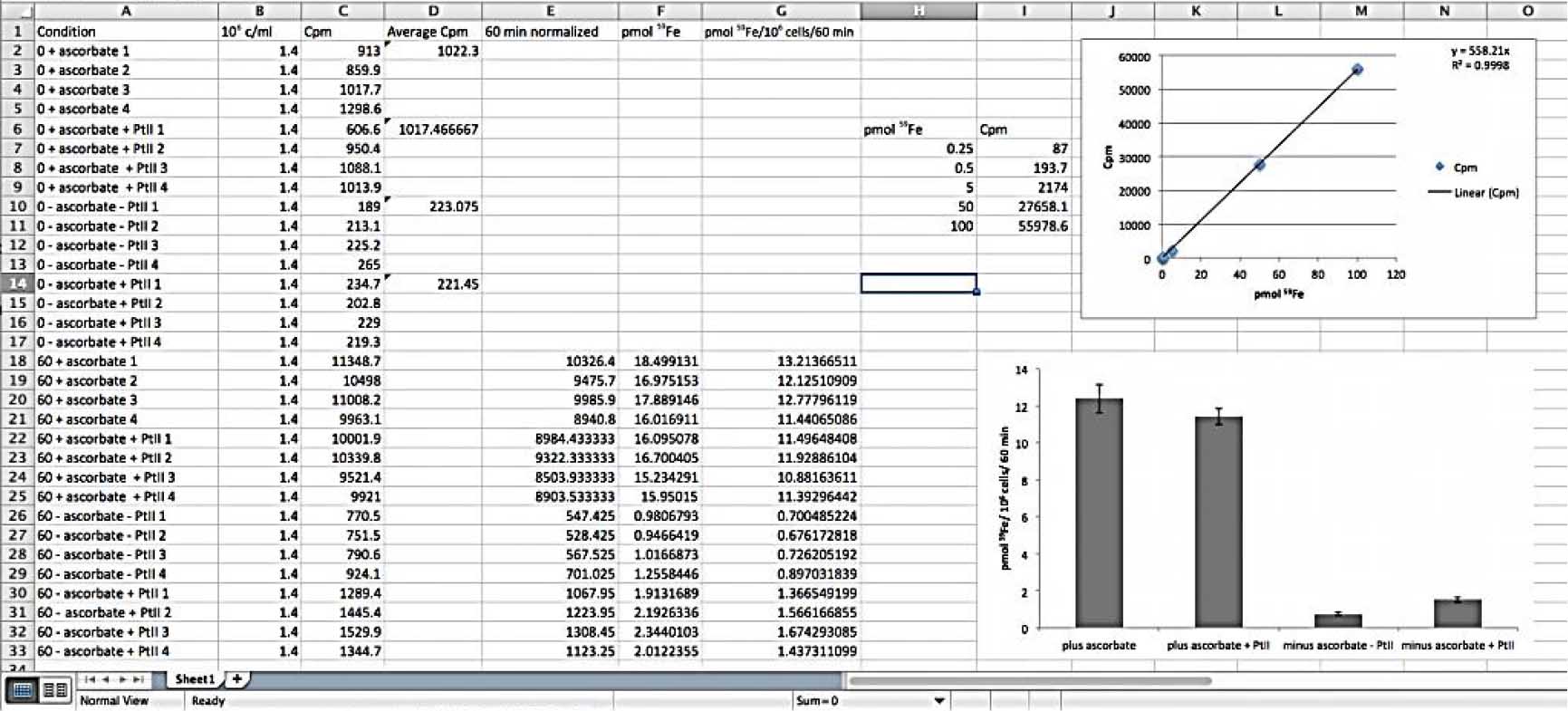
Figure 4. Representative data expected to obtain after the g counter reads. The standards are important to provide a calibration curve from which it will be possible to convert counts per minute (Cpm) to pmol 59Fe (top graphic). After normalization of the data with the 0 time point (fifth column), the pmol 59Fe is calculated based on the calibration curve and then it is possible to correlate the quantity of 59Fe (in pmol) 106 cells could internalize at a time, in this case 60 min (lower graphic).
Recipes
- SC complete media
6.7 g YNB to 750 deionized water
100 ml of 20% glucose
100 ml of amino acid solution
Sterile deionized water to 1,000 ml total volume
Filter sterilize media - Amino acid solution
2.26 g of amino acid mix stock to 100 ml of deionized water
Heat and stir into solution
Allow to cool - Amino acid mix stock
2 g ADE
10 g TRP
10 g HIS
7.5 g ARG
2.5 g MET
1.5 g TYR
2.5 g ISOLEU
2.5 g LYS
2.5 g PHE
5 g GLU. ACID
5 g ASP. ACID
7.5 g VAL
10 g THR
20 g SER
2 g URA
22.5 g LEU
Note: This is a powder stock. The amino acid solution is explained above. - 59Fe stock solutions
- To make a working stock of 20 μM 59Fe in ddH2O:
For an experiment with 2 time points and 6 samples (individual vials), a minimum of 420 μl of 20 μM 59Fe is needed since each sample (vial) needs 70 μl of 20 μM 59Fe. It is recommended that the amount of 59Fe solution to be used is calculated assuming an extra 1.5 samples. For this example, prepare 7.5 x 70 μl, or 525 μl of 59Fe stock. This precaution ensures that, even with pipetting errors, there will be enough to make standards.
Thus, for 525 μl of 20 μM 59Fe = 3.03 μl of 3.47 mM 59Fe stock (see Note below) plus 521.97 μl ddH2O. In each vial, therefore, 70 μl of 20 μM 59Fe stock in 7 ml uptake buffer = 0.2 μM 59Fe.
Make serial dilutions of the 20 μM 59Fe stock to make the 2 and 0.2 μM stocks used for the standards.
For example, 100 μl 20 μM 59Fe + 900 μl ddH2O = 2 μM 59Fe.
Notes: - Do not dilute the entire stock vial of radionuclide received from the vendor! Make up stock working solution only as needed.
- Calculate the [59Fe] in the radionuclide received from the vendor. An example is given for product number NEZ037001MC from Perkin Elmer Life Sciences.
- Divide the concentration by the specific activity. This gives the concentration in mg/ml [(mCi/ml)/(mCi/mg) = mg/ml].
For example, If Perkin Elmer ships 0.5 mCi 59Fe having a specific activity of 72.87 mCi/mg with a concentration of 49.36 mCi/ml, the [59Fe] is determined in the following way:
(49.36 mCi/ml)/(72.87 mCi/mg) = 0.67737 mg/ml
(0.67737 mg/ml)/(59 g) = 0.01148 mmoles of Fe/ml
(0.01148 mmol/ml) (1,000 ml/L) = 11.48 mM = [59Fe]
This concentration is as of the stock date (the date the batch of 59Fe was made). In order to determine the concentration as of the date of your experiment, count the number of days between the stock date and your experiment date. Using an 59Fe decay table, find the corresponding decay factor. Multiply the decay factor by the stock date calibration. This gives you the [59Fe] in the stock vial as of the date of your experiment.
To determine the volume of liquid in the 59Fe stock vial, divide the amount of iron you ordered (i.e., 500 μCi) by the concentration (mCi/ml) on the date one week after the iron was shipped. - 10x quench buffer (1,000 ml)
Note: Do not filter sterilize.
44.4 g succinic acid (Final concentration = 375 mM)
75.6 g Tris (Final concentration = 625 mM)
37.5 g EDTA (acid form) (Final concentration = 128 mM)
Everything will not go into solution until all three chemicals have been added and pH is adjusted.
Adjust pH to 6.0 using NaOH
Add ddH2O to final volume of 1,000 ml
Store at 4 °C in acid-washed glassware
Dilute to 1x for use in uptakes - Citrate uptake buffer (1,000 ml)
Note: Make and filter sterilize this buffer in acid-washed glassware.
20.0 g Analar glucose (Final concentration = 2%)
19.8 g MES buffer (Final concentration = 0.1 M)
5.9 g Na-Citrate (Final concentration = 20 mM)
Add NaOH to adjust pH to 6.0
Add ddH2O to final volume of 1,000 ml
Filter sterilize
Store at room temperature - Acid-wash glassware treatment
Soak glassware in HNO3 (3-6 h)
Note: This process could be performed overnight.
Rinse 3 times in deionized water
Fill 1/3 of the glassware with water and autoclave (cap with aluminum foil)
Pour out water and wash more 3 times with water
Refill glassware with water (1/3 full) and autoclave again (cap with aluminum foil)
Pour water out just prior to use
Acknowledgments
The authors would like to thank to Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Comissāo de Avaliação de Pessoal de Nível Superior (CAPES) and Fundação de Amparo a Pesquisa do Estado de Goiás (FAPEG). Development of this protocol was supported by a grant from the National Institutes of Health (US) DK053820 to DJK.
References
- Bailão, E. F. L. C., Lima, P. S., Silva-Bailão, M. G., Bailão, A. M., Fernandes. G. R., Kosman, D. J. and Soares, C. M. A. (2015). Paracoccidioides spp. ferrous and ferric iron assimilation pathways. Front Microbiol 6: 821.
- Eide, D., Davis-Kaplan, S., Jordan, I., Sipe, D. and Kaplan, J. (1992). Regulation of iron uptake in Saccharomyces cerevisiae. The ferrireductase and Fe(II) transporter are regulated independently. J Biol Chem 267(29): 20774-20781.
- Kosman, D. J. (2013). Iron metabolism in aerobes: managing ferric iron hydrolysis and ferrous iron autoxidation. Coord Chem Rev 257(1): 210-217.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kosman, D. J., Bailão, E. F. L. C., Silva-Bailão, M. G. and Soares, C. M. D. A. (2016). 59Fe Uptake Assays in Paracoccidioides Species. Bio-protocol 6(18): e1930. DOI: 10.21769/BioProtoc.1930.
Category
Microbiology > Microbial metabolism > Other compound
Microbiology > Microbial biochemistry > Other compound
Biochemistry > Other compound > Ion
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









