- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Determination of VPS34/PIK3C3 Activity in vitro Utilising 32P-γATP
Published: Vol 6, Iss 16, Aug 20, 2016 DOI: 10.21769/BioProtoc.1904 Views: 10025
Reviewed by: Ralph BottcherNeelanjan BoseMarc-Antoine Sani

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
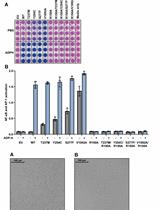

Measurement of the Activity of Wildtype and Disease-Causing ALPK1 Mutants in Transfected Cells With a 96-Well Format NF-κB/AP-1 Reporter Assay
Tom Snelling
Nov 20, 2024 1609 Views
Fluorescence Polarization-Based High-Throughput Screening Assay for Inhibitors Targeting Cathepsin L
Keyu Guo [...] Shuyi Si
Jul 20, 2025 2282 Views
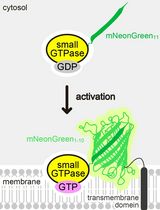
Detecting the Activation of Endogenous Small GTPases via Fluorescent Signals Utilizing a Split mNeonGreen: Small GTPase ActIvitY ANalyzing (SAIYAN) System
Miharu Maeda and Kota Saito
Jan 5, 2026 496 Views
Abstract
VPS34 is the only class III phosphatidylinositol-3-kinase (PI3K) in mammalian cells and produces the vast majority of cellular phosphatidylinositol-3-phosphate [PI(3)P]. PI(3)P is a key signalling lipid that plays many membrane trafficking roles in processes such as endocytosis and autophagy. VPS34 is a key cellular regulator, loss of function can have catastrophic effects and is embryonic lethal (Zhou et al., 2011). The levels of cellular PI(3)P can be determined by fluorescent staining techniques and can be used to monitor effects upon VPS34 activity, however it is important to verify that any changes are mediated by VPS34, particularly as alternate pathways of PI(3)P production are possible such as via class II PI3Ks (Devereaux et al., 2013). Assaying VPS34 activity directly in vitro can be a key stage in delineating the action of a particular stimulus.
Keywords: Phosphatidylinositol-3-phosphateMaterials and Reagents
- Hamilton syringe (500 μl) (Hamilton, catalog number: 80865 )
- Pyrex tubes (12 x 75 mm) (Corning, catalog number: 99445-12 )
- Eppendorf flex tubes (Eppendorf, catalog number: 022364111 )
- Spin-X columns (0.22 μm) (Corning, Costar®, catalog number: 8161 )
- Whatman paper (3 mm) (Sigma-Aldrich, catalog number: 3030-917 )
- Silica 60 TLC plates (Merck Millipore Corporation, catalog number: 1.05553.0001 )
- TLC spotting capillary tubes, 15.2 cm (VWR, catalog number: 80060-608 )
- Polycarbonate membranes, 100 nm pore size (Avanti Polar Lipids, catalog number: 610005 )
- Chloroform (VWR International, catalog number: 22707.320 )
- Crude liver phosphatidylinositol (PI) (Avanti, catalog number: 840042C )
- Methanol (VWR International, catalog number: 20847.307 )
- N2 gas tank (99.998% Pure)
- VPS34/VPS15 recombinant protein (MRCPPU Reagents, catalog number: DU8692 )
- 32P-γ-ATP (PerkinElmer, catalog number: NEG002A500UC )
- ATP (Sigma-Aldrich, catalog number: A2383 )
- Hydrochloric acid (37%) (VWR International, catalog number: 20252.335 )
- Glycerol (VWR International, catalog number: 24388.320 )
- Sodium dodecyl sulphate (SDS) (VWR International, catalog number: 444464T )
- Bromophenol blue (Sigma-Aldrich, catalog number: 114405 )
- β-mercaptoethanol
- Tris base (VWR International, catalog number: 103157P )
- Sodium chloride (VWR International, catalog number: 27810.364 )
- Manganese chloride (Sigma-Aldrich, catalog number: M3634 )
- CHAPS (Sigma-Aldrich, catalog number: C3023 )
- Dithiothreitol (DTT) (ForMedium, catalog number: DTT010 )
- Ammonium hydroxide (Sigma-Aldrich, catalog number: 320145 )
- ATP mix (5x) (see Recipes)
- Kinase assay buffer (see Recipes)
- Stop solution (see Recipes)
- Sample buffer (1x) (see Recipes)
- TLC solvent (see Recipes)
- Potassium oxalate (see Recipes)
Equipment
- Speedvac (Thermo Fisher Scientific, model: SPD131DDA )
- Thermomixer (Eppendorf)
- Sonicating Water Bath (Sigma-Aldrich, model: SONOREX Digital 10P )
- Mini-extruder set (Avanti, catalog number: 610000 )
- Hairdryer
- TLC chamber
- Fumehood
- Phosphoimager (optional) (Fujifilm, model: FLA-2000 )
Procedure
- Liposome preparation
- Lipids should be kept stored in a glass container in chloroform at -20 °C until required for use. Only prepare liposomes as required and use prepared liposomes within 24 h. Use a glass Hamilton syringe to transfer the required amount of phosphatidylinositol (PI) (10 mg/ml) in chloroform to a clean (methanol washed and dried) glass tube (Pyrex 12 x 75 mm). As 10 µg of extruded liposomes are used per reaction, it will require 1 µl per reaction to be carried out. It is advised to prepare slightly more liposomes than required as some volume is lost during the extrusion process (see step 1e). It is advisable to work with a minimum of 20 µl of chloroform (10 mg/ml) lipids; it is also advised to carry out each reaction in duplicate or triplicate (see Note 1 for further information).
- Evaporate the chloroform under a stream of nitrogen gas. Use a steady flow rate that is sufficient to disturb and push the lipid up against the side of the tube as it dries, this reduces the chances of residual chloroform becoming trapped under dried lipid. A subtle dried opaque coating should be visible on the tube.
- Dry the lipid further for 1 h in a speedvac with suitable attachment for the Pyrex tubes. Traces of chloroform can be detrimental to the assay, so it is important that all chloroform is evaporated.
- Re-suspend the PI in assay buffer (to give a 1 mg/ml solution) and allow 30 min to re-suspend with periodic vortexing and by using a sonicating water bath. The mixture should look chalky/cloudy.
- Prepare the lipid mini-extruder by assembling with a 100 nm pore filter, assay buffer (~1 ml) can be used to flush through the extruder and ensure the system is assembled correctly without leakage and this will minimise any loss of volume. Detailed assembly instructions can be found via the Avanti website (http://avantilipids.com/divisions/equipment/mini-extruder-assembly-instructions/). Collect the lipid solution in one glass Hamilton syringe and attach to the extruder system. Pass slowly through without excessive pressure from one side to the other (~10 times) until the solution becomes clear (as seen on Avanti website: http://avantilipids.com/divisions/equipment/mini-extruder-extrusion-technique/), indicating that liposomes (Large unilamellar vesicles - LUVs) have been formed and are now in solution. Always ensure that the lipid mixture is at the opposite side to where it began so that everything has passed through the filter.
- The liposomes are now prepared and ready for use in the in vitro assay.
- In vitro phosphorylation of PI
This assay is suitable for use with either recombinant (Bago et al., 2014) or immunoprecipitated VPS34 (Munson et al., 2015), although in the latter case it is important to ensure the antibody utilised does not interfere with activity. Follow a standard immunoprecipitation protocol to isolate VPS34 directly or co-immunoprecipitate it by targeting a binding partner (e.g., BECLIN1, ATG14L, UVRAG). As this assay involves the use of radioactive 32P-γ-ATP, ensure that all work is carried out in an area where radioactivity is allowed and meets local regulations and requirements for its use. When carrying out this assay, it is important to include appropriate controls such as a negative bead only/no kinase control, a positive kinase control [this could be recombinant protein of Class I or Class III PI3K as both will form PI(3)P in vitro from PI]. A PI3K inhibitor (such as Wortmannin or VPS34-IN1) can also be used to verify that PI(3)P formation is being observed (as these should block formation).- Suspend either recombinant protein or beads if immunoprecipitating in a final volume of 40 μl assay buffer inside Eppendorf tubes. Dilute the extruded liposome mix within the 40 µl volume to a total amount of 10 μg of extruded liposomes per reaction.
- Add 10 μl of the 5x ATP mix as a final addition to each sample and place directly into a thermomixer at 30 °C/1,100 rpm for 30 min. It is ideal to stagger the addition of ATP mix to each sample to allow time to process each sample at the end of the 30 min reaction. This time will vary dependent if the assay is being carried out on beads from an immunoprecipitation or with recombinant protein. It is advised to become familiar with the following steps and have a trial run to determine the optimal staggering time.
- Terminate the reaction by the addition of 500 μl Stop solution. If using beads, then first separate the beads by passing through a Spin-X column before addition of Stop solution to the eluate. Sample buffer (1x) can be added to the beads retained in the column and re-eluted into a fresh tube. These samples can then be used for western blot to verify and determine the levels of immunoprecipitated protein between samples and serve as a control for the assay.
- Once all samples are in Stop solution, the components are phase-split by the addition of 180 μl chloroform and subsequently 300 μl 0.1 M HCl per sample. Vortex samples gently before centrifugation at 0.5 x g/1 min. The samples will split into an upper aqueous (water-methanol) and lower organic (chloroform-methanol) lipid-containing layer.
- Prime a pipette tip by briefly pipetting chloroform up and down. Once primed, put the pipette tip through the methanol layer to aspirate the lower chloroform layer and remove to a separate tube. Expel the pipette whilst moving to collect the lower layer to minimise carry over.
- Dry chloroform samples in a speedvac (~5-10 min). Once dry, samples can either be retained at -20 °C to run at a later date or used immediately for separation of lipid products by TLC.
- Thin layer chromatography (TLC)
Lipid products can be separated by spotting and running on a Silica 60 TLC plate. A simple overview of the TLC process is shown in Figure 1.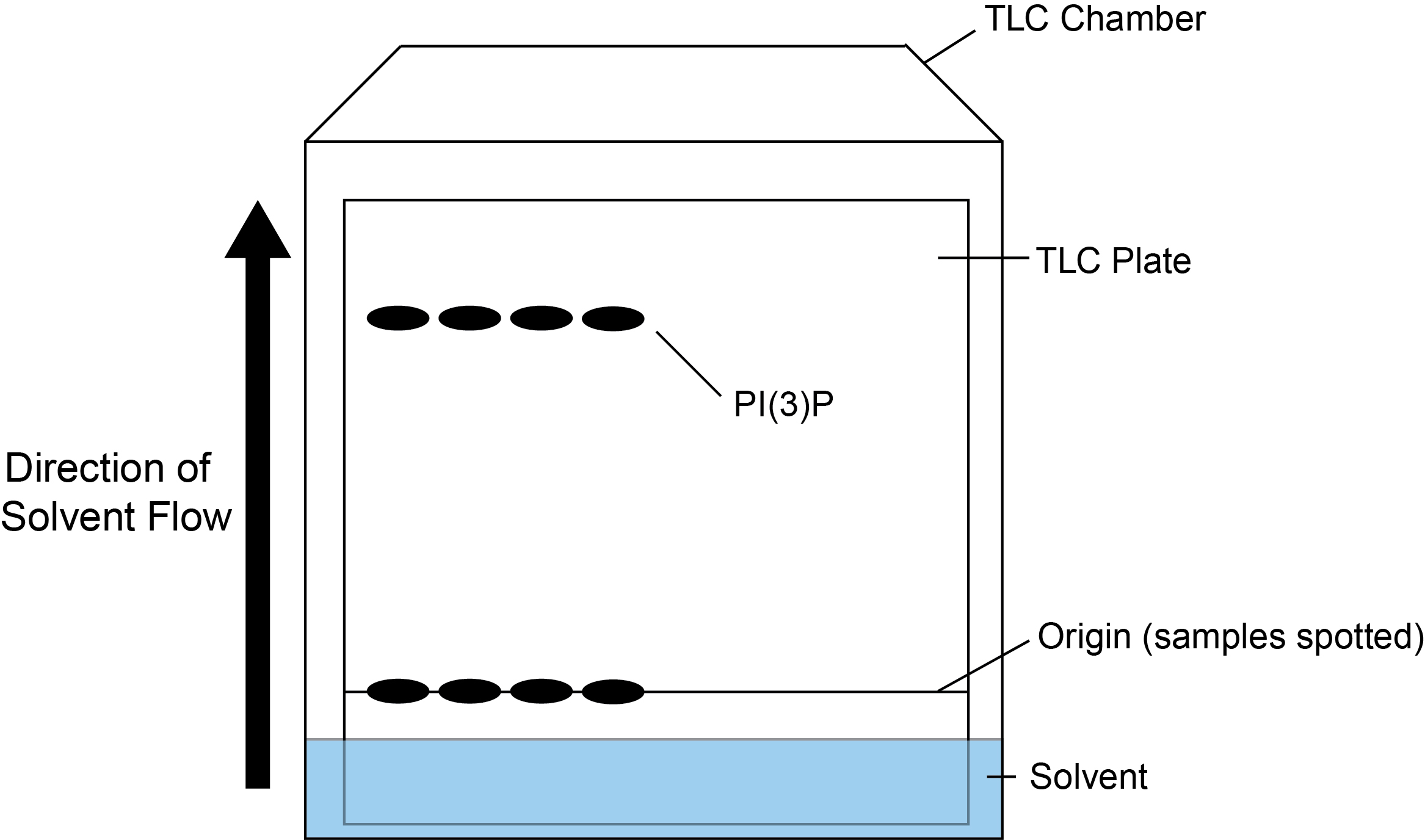
Figure 1. Overview of TLC procedure. Samples are spotted onto the origin (a point above the TLC solvent) and added to the TLC chamber containing solvent. As the solvent travels up the plate, the phosphorylated lipid products are separated. In this case, the lipid product PI(3)P cannot but seen by eye, but will be detected by the incorporation of 32P.- Pre-treat the TLC plate by wetting completely in potassium oxalate solution to improve the resolution of phospholipids (Gonzalez-Sastre and Folch-Pi, 1968). Allow to dry fully, this can be left at RT overnight (~16 h) or can be dried more rapidly using a hairdryer. The TLC chamber should also be prepared in advance by addition of the TLC solvent to saturate the chamber. Whatman paper can be added at the sides of the chamber to improve speed and maintain even resolution when running the TLC plates.
- Mark gently in pencil where each sample is to be loaded at least 2 cm from the bottom of the TLC plate. The loaded samples must not be lower than the solvent level in the tank.
- Dried samples should be resuspended in 50-100 μl of chloroform and spotted onto the TLC plate. A glass narrow-bore capillary can be utilised for spotting samples onto the TLC plate. The sample should be applied gradually, allowing the chloroform to dry before adding further. Application speed can be improved by utilising a hairdryer to evaporate the chloroform quickly between spotting.
- Once completely loaded, the TLC plate should be placed into the TLC chamber and left until the solvent front reaches just below the top of the plate. The plate should then be removed and allowed to dry in the fume hood.
The level of phosphorylation can then be determined by multiple means. The incorporation of 32P correlates to phosphorylated lipid and can be measured by exposure to x-ray film. Other methods that have a greater linear range of detection such as a Phosphorimager can be utilised if available for more accurate quantitation. Assay of recombinant VPS34/VPS15 is very clean and produces a single band that migrates up the plate. Signal observed at the origin that does not migrate is likely to be carryover 32P-γ-ATP that has not been incorporated into a lipid product.
In the case of an immunoprecipitation, the retained sample buffer from step 2c can be analysed by western blot for VPS34 levels. This can then serve as a loading control to normalise the values obtained via the TLC plate.
Representative data
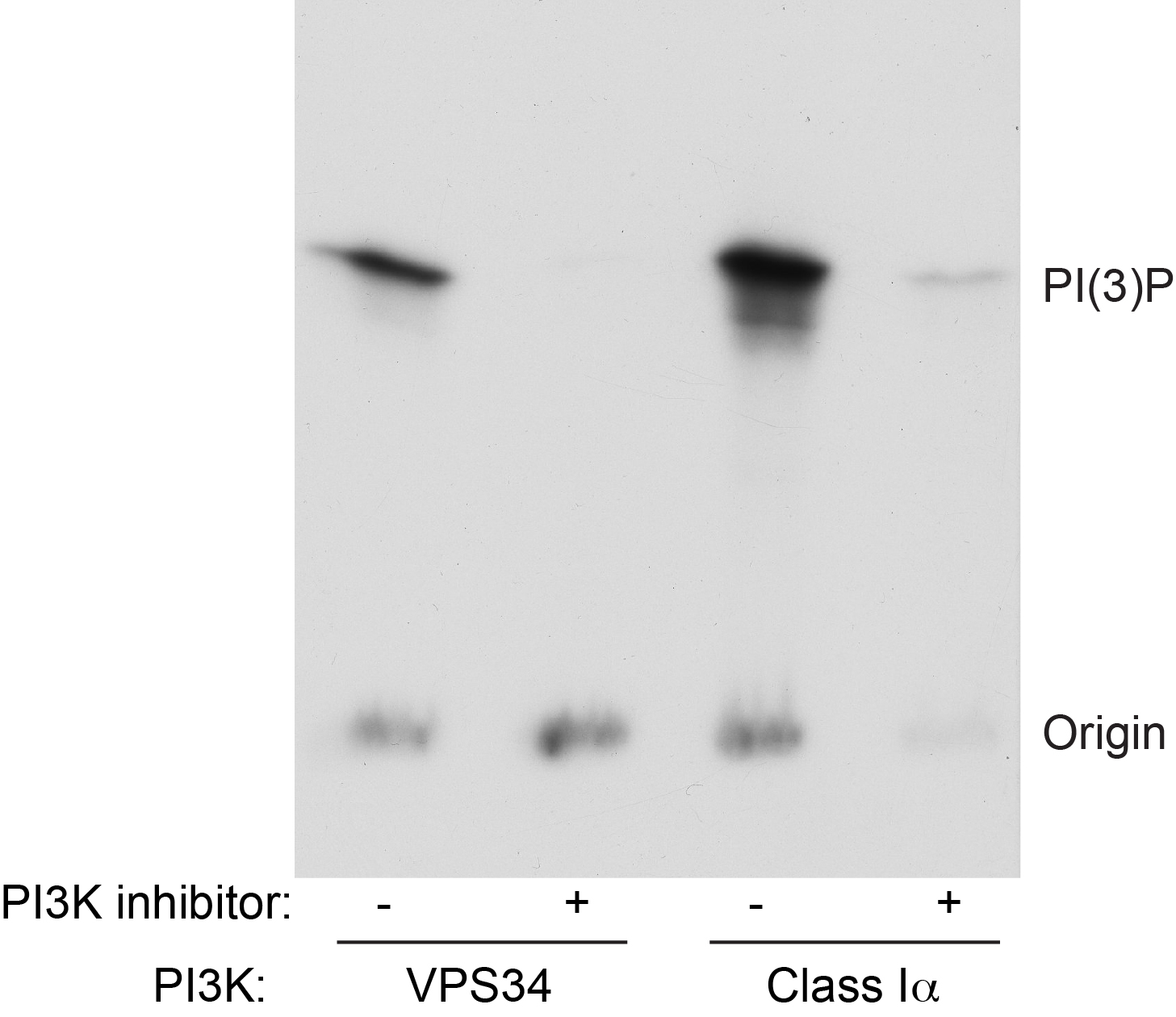
Figure 2. Example data from lipid kinase assay. Recombinant VPS34/VPS15 or Class I PI3Ka was assayed as described in the presence or absence of a pan PI3K inhibitor. Following loading of samples onto a TLC plate and running, the plate was exposed to x-ray film and developed.
Notes
- The data obtained via lipid kinase assay is inherently more variable than a protein kinase assay, largely due to the greater variation in the substrate itself. Extruding lipids to form liposomes of maximal size 100 nm helps to make the substrate more uniform and reduce variability, although some will remain. Extruding the solution a second time will help to increase homogeneity further. Carrying out each reaction in duplicate or triplicate and taking the average value can also help to increase reliability. The level of homogeneity can be determined by performing dynamic light scattering (DLS).
- The curvature of liposomes may also affect the ability to act as a substrate for VPS34, certain sub-complexes are reported to have different phosphorylation kinetics dependent upon membrane curvature (Rostislavleva et al., 2015). It may therefore be wise also to consider use of different filter pore sizes when extruding to test a range of liposome sizes.
Whilst assays of VPS34 activity can be very clean, it is prudent to include a suitable positive control to confirm that any bands observed via TLC are indeed PI(3)P. This could be either using recombinant VPS34/VPS15 or a class I PI3K [these will form PI(3)P from PI in vitro but not in cells]. Additionally, a specific VPS34 inhibitor such as VPS34-IN1 may be used to confirm any signal is VPS34 dependent (Bago et al., 2014).
Recipes
- ATP mix (5x)
5 μCi 32P-γ-ATP
5 μM ATP
Prepare in kinase assay buffer
Prepare fresh before use - Kinase assay buffer
20 mM Tris, pH 7
67 mM NaCl
10 mM MnCl2
0.02% (w/v) CHAPS
1 mM DTT
Prepare fresh before use - Stop solution
Chloroform:methanol:12 M HCl (100:200:3.5, v/v)
Store at RT in glass top bottles
Be wary of chloroform evaporation over time - Sample buffer (1x)
62.5 mM Tris, pH 6.8
10% (v/v) glycerol
2% (w/v) SDS
0.005% bromophenol blue
2.5% (v/v) β-mercaptoethanol
Can be stored at RT but add β-mercaptoethanol fresh - TLC solvent
Methanol:chloroform:water:14.5 M ammonium hydroxide (47:60:11.2:2, v/v)
Store at RT in glass top bottles
Solvent can be reused 2-3 times
Be wary of chloroform evaporation over time - Potassium oxalate
1% (w/v) potassium oxalate
5 mM EDTA
50% (v/v) methanol
Can be stored at RT
Acknowledgments
This work was supported by the Medical Research Council (MRC) and the pharmaceutical companies supporting the Division of Signal Transduction Therapy Unit (AstraZeneca, Boehringer-Ingelheim, GlaxoSmithKline, Merck KGaA, Janssen Pharmaceutica and Pfizer).
References
- Bago, R., Malik, N., Munson, M. J., Prescott, A. R., Davies, P., Sommer, E., Shpiro, N., Ward, R., Cross, D., Ganley, I. G. and Alessi, D. R. (2014). Characterization of VPS34-IN1, a selective inhibitor of Vps34, reveals that the phosphatidylinositol 3-phosphate-binding SGK3 protein kinase is a downstream target of class III phosphoinositide 3-kinase. Biochem J 463(3): 413-427.
- Devereaux, K., Dall'Armi, C., Alcazar-Roman, A., Ogasawara, Y., Zhou, X., Wang, F., Yamamoto, A., De Camilli, P. and Di Paolo, G. (2013). Regulation of mammalian autophagy by class II and III PI 3-kinases through PI3P synthesis. PLoS One 8(10): e76405.
- Gonzalez-Sastre, F. and Folch-Pi, J. (1968). Thin-layer chromatography of the phosphoinositides. J Lipid Res 9(4): 532-533.
- Munson, M. J., Allen, G. F., Toth, R., Campbell, D. G., Lucocq, J. M. and Ganley, I. G. (2015). mTOR activates the VPS34-UVRAG complex to regulate autolysosomal tubulation and cell survival. EMBO J 34(17): 2272-2290.
- Rostislavleva, K., Soler, N., Ohashi, Y., Zhang, L., Pardon, E., Burke, J. E., Masson, G. R., Johnson, C., Steyaert, J., Ktistakis, N. T. and Williams, R. L. (2015). Structure and flexibility of the endosomal Vps34 complex reveals the basis of its function on membranes. Science 350(6257): aac7365.
- Zhou, X., Takatoh, J. and Wang, F. (2011). The mammalian class 3 PI3K (PIK3C3) is required for early embryogenesis and cell proliferation. PLoS One 6(1): e16358.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Munson, M. J. and Ganley, I. G. (2016). Determination of VPS34/PIK3C3 Activity in vitro Utilising 32P-γATP. Bio-protocol 6(16): e1904. DOI: 10.21769/BioProtoc.1904.
- Munson, M. J., Allen, G. F., Toth, R., Campbell, D. G., Lucocq, J. M. and Ganley, I. G. (2015). mTOR activates the VPS34-UVRAG complex to regulate autolysosomal tubulation and cell survival. EMBO J 34(17): 2272-2290.
Category
Biochemistry > Protein > Activity
Biochemistry > Lipid > Lipid measurement
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link