- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Expression, Purification and Crystallization of the Herpesvirus Nuclear Egress Complex (NEC)
Published: Vol 6, Iss 14, Jul 20, 2016 DOI: 10.21769/BioProtoc.1872 Views: 8773
Reviewed by: Arsalan DaudiBalasubramanian VenkatakrishnanAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Optimized Secretome Sample Preparation From High Volume Cell Culture Media for LC–MS/MS Proteomic Analysis
Basil Baby Mattamana [...] Peter Allen Faull
Dec 20, 2025 1261 Views
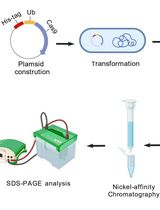
A One-Step Method for Efficient Purification of Functional Cas9 Protein
Xinzhi Duan [...] Aihua Mao
Feb 5, 2026 157 Views
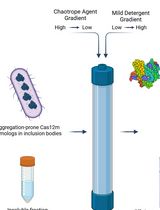
On-Column Dual-Gradient Refolding for Efficient Recovery of Insoluble Affinity-Tagged Recombinant Proteins
Anna Vlaskina [...] Maxim Patrushev
Feb 5, 2026 66 Views
Abstract
The protocol describes the production and crystallization of the soluble form of the nuclear egress complex (NEC) from Herpes simplex virus 1 and Pseudorabies virus. The NEC is a heterodimer that consists of conserved proteins UL31 and UL34. NEC oligomerization deforms the inner nuclear membrane around the capsid in infected cells, thereby mediating capsid budding into the perinuclear space during nuclear egress. We have successfully developed a protocol for large-scale preparation of highly pure NEC from two different viruses in a prokaryotic expression system, which enabled us to crystallize these viral protein complexes and determine their structures. This procedure may be adapted to purify and crystallize other soluble protein complexes.
Keywords: Nuclear egress complexMaterials and Reagents
- Talon HiTrap columns, 1 ml (GE Healthcare, catalog number: 28-9537-66 )
- 24-well crystallization plates, VDX Greased Plates (Hampton Research Corporation, catalog number: HR3-170 )
- Cover slips, 22 mm, siliconized (Hampton Research Corporation, catalog number: HR3-233 )
- Nylon hydrophilic membrane filters, 0.2 µm (Merck Millipore Corporation, catalog number: GNWP04700 )
- Ultrafree centrifugal filters, 0.1 µm (Merck Millipore Corporation, catalog number: UFC30VV00 )
- LB (Luria Bertani) agar plates
- Escherichia coli (E. coli) BL21 Rosetta (DE3) (Merck Millipore Corporation, Novagen, catalog number: 70954 )
- pGEX-6P1 vector (GE Healthcare, catalog number: 28-9546-48 )
- pET-24b vector (Merck Millipore Corporation, catalog number: 69750-3 ), modified to include a sequence encoding a His6-SUMO tag followed by a PreScission cleavage site
- Ampicillin (Fisher Scientific, catalog number: BP1760-25 )
- Kanamycin (Fisher Scientific, Durable, catalog number: BP906-5 )
- Chloramphenicol (Fisher Scientific, Bioreagents, catalog number: BP904-100 )
- Tryptone (Fisher Scientific, catalog number: BP1421-2 )
- Yeast extract (Fisher Scientific, catalog number: BP1422-2 )
- Glycerol (P212121, catalog number: RP-G22020-0.5 )
- KH2PO4 (Fisher Scientific, catalog number: P386-500 )
- K2HPO4 (Fisher Scientific, catalog number: LC200901 )
- Glucose (American Bio, catalog number: AB00715 )
- Lactose (Fisher Scientific, Durable, catalog number: L5-500 )
- MgSO4·7H2O (Sigma-Aldrich, Fluka, catalog number: 63140 )
- EDTA-free protease inhibitor cocktail tablets (Sigma-Aldrich, cOmpleteTM, catalog number: 5056489001 )
- Deoxyribonuclease I from bovine pancreas (Sigma-Aldrich, catalog number: DN25-1 g )
- HEPES (Fisher Scientific, Durable, catalog number: BP310500 )
- Sodium chloride (NaCl) (Thermo Fisher Scientific, Durable, catalog number: S271-10 )
- TCEP HCl (P212121, catalog number: SV-TCEP-25 g )
- Imidazole (Thermo Fisher Scientific, ACROS Organics, catalog number: 301870010 )
- Ni Sepharose 6 Fast Flow (GE Healthcare, catalog number: 17-5318-02 )
- Glutathione Sepharose 4B (GE Healthcare, catalog number: 17-0756-05 )
- PreScission protease (produced in house; for more information see Reference 3)
- 12% Mini Protean TGX precast gels (Bio-Rad Laboratories, catalog number: 456-1046 )
- Coomassie Brilliant Blue R-250 (Bio-Rad Laboratories, catalog number: 161-0400 )
- Acetic acid (Fisher Scientific, Durable, catalog number: A491-212 )
- Ethanol, 190 proof (Fisher Scientific, Durable, catalog number: 04-355-226 )
- Guanidine hydrochloride (Sigma-Aldrich, catalog number: 50950 )
- Trisodium citrate dihydrate (Alfa Aesar, catalog number: 36439-A3 )
- NiCl2 hexahydrate (Alfa Aesar, catalog number: 43185 )
- Polyethylene glycol 8000 (Sigma-Aldrich, catalog number: 89510 )
- NaSCN (VWR International, catalog number: 10118-142 )
- Polyethylene glycol 3350 (Sigma-Aldrich, catalog number: P3640 )
Note: Product P3640 has been discontinued. - Meso-erythritol (VWR International, catalog number: CAAAA15813-22 )
- TB (Terrific broth) medium (see Recipes)
- Lysis buffer (see Recipes)
- Gel filtration buffer (see Recipes)
Equipment
- Standard laboratory equipment
- Spectrophotometer device (Thermo Fisher Scientific, NanoDropTM, model: 1000 ) or comparable UV/VIS spectrophotometer
Note: This product has been discontinued by the manufacturer. - Microfluidizer cell disruptor (Microfluidics, model: 110S ) or comparable cell lysis equipment
Note: This product has been discontinued by the manufacturer. - Pumps (Bio-Rad Laboratories, model: EP-1 )
Note: This product has been discontinued by the manufacturer. - Chromatography system (Bio-Rad Laboratories, DuoFlowTM, model: Medium-pressure ) or comparable chromatography system
- Superdex 75 10/300 (GE Healthcare, catalog number: 17-5174-01 )
- Stereo microscope (ZEISS, model: SteREO Discovery V8 ) or comparable microscope
- Amicon Ultra-15 Centrifugal Filter Unit with Ultracel-50 membrane (Merck Millipore Corporation, catalog number: UFC905096 )
- Ultrasonic bath (Fisher Scientific, model: FS60) or comparable sonication device
Note: This product has been discontinued by the manufacturer.
Procedure
- Co-transform plasmids encoding HSV-1 UL31 (15-185) or PRV UL31 (1-176), in His6-SUMO-3C-pET24b, and HSV-1 UL34 (51-306) or PRV UL34 (18-271), in pGEX-6P1, into E. coli Rosetta cells using heat shock transformation. Plate on LB-agar plates containing 100 mg/L ampicillin, 50 mg/L kanamycin and 34 mg/L chloramphenicol.
- Pick a single colony to inoculate a starter culture of 50 ml TB medium containing 1% glucose, 2 mM MgSO4, 34 mg/L chloramphenicol, 100 mg/L ampicillin and 100 mg/L kanamycin. Incubate the culture in a shaker at 37 °C, 200 rpm, for 16 h.
- Inoculate 3 L of TB containing 0.2% lactose, 2 mM MgSO4, 100 mg/L ampicillin and 100 mg/L kanamycin with 10 ml of the starter culture and grow at 37 °C, 200 rpm, for 4 h. Reduce the temperature to 25 °C and grow the cultures for another 16 h. Typically a final OD600 of 5-6 is reached.
- Harvest cells by centrifugation at 5,000 x g for 30 min at 4 °C.
- Filter and degas all buffers used in purification.
- Throughout the purification, confirm the presence and quality of the purified proteins by SDS-PAGE analysis and Coomassie staining (Figure 1). Determine protein concentration by measuring the OD280 [extinction coefficients: 37,360 M-1 cm-1 (HSV-1 NEC) and 34,840 M-1 cm-1 (PRV NEC)] using a spectrophotometer.
- Wash cell pellets in lysis buffer and centrifuge in 50 ml conical tubes for 5 min at 5,000 x g at 4 °C. Discard the supernatant and process pellets immediately or flash freeze in liquid nitrogen and store at -80 °C.
- Resuspend cell pellets in lysis buffer supplemented with the cOmplete Protease Inhibitor mix (one tablet/25 ml lysate) and DNase I (one tip of a spatula/50 ml lysate).
- Lyse cells in a microfluidizer (3 runs, 80 psi). A comparable cell disruptor may be used instead.
- Clear the cell lysate by centrifugation at 20,000 x g at 4 °C for 30 min. All subsequent steps are to be carried out at 4 °C or on ice.
- Apply the supernatant onto a Ni-NTA sepharose column (8 ml), equilibrated with lysis buffer, with a flow rate of 2 ml/min and wash with lysis buffer containing 20-40 mM imidazole. Elute bound proteins with lysis buffer containing 250 mM imidazole.
- Pass the eluted proteins over a GSH sepharose column (8 ml), equilibrated with lysis buffer, to remove excess His6-SUMO-UL31. After washing with lysis buffer, use PreScission protease in a 1:30 molar ratio to cleave His6-SUMO and GST tags on the GSH column for 16 h in one column volume lysis buffer circulating at a flow rate of 0.3 ml/min.
- Apply lysis buffer to the GSH sepharose column and collect the flow-through fraction containing untagged NEC and His6-SUMO. Apply this sample onto 2 tandem-coupled 1 ml Talon HiTrap columns to separate His6-SUMO from the NEC. Collect the flow-through fraction, containing the untagged NEC, and concentrate it using a concentrator with a MWCO of 50 kDa.
- Between different protein preparations, wash the Ni-NTA and GSH columns with one column volume 6 M guanidine hydrochloride and 0.2 M acetic acid. Then wash with 10 column volumes dH2O.
- To obtain monodisperse NEC, as a final purification step, apply the concentrated sample from step 13 to a Superdex 75 10/300 column equilibrated with gel filtration buffer.
- Pool NEC-containing fractions from the monodisperse peak and concentrate up to ~10 mg/ml. Aliquot protein samples and flash freeze in liquid nitrogen for subsequent storage at -80 °C. The typical yield is 10 mg monodisperse NEC per L TB culture.
- Prior to crystallization, dilute proteins to 5 mg/ml and filter through a 0.1 µm membrane to remove aggregates.
- Grow HSV-1 NEC crystals by vapor diffusion in hanging drops with 0.5 µl protein and 0.5 µl reservoir solution in 10% PEG 8000, 0.1 M Na citrate, pH 5.6, 5 mM NiCl2 at 22 °C. HSV-1 NEC crystals resembling hexagonal plates appear after 2 days and reach their final size after 1 week (Figure 2A).
- Soak HSV-1 NEC crystals briefly in the reservoir solution containing 25% glycerol as a cryoprotectant and flash freeze in liquid nitrogen.
- Grow PRV NEC crystals by vapor diffusion in hanging drops with 0.5 µl protein and 0.5 µl reservoir solution in 18% PEG 3350, 0.3 M NaSCN, 0.3 M NaCl at 22 °C. Tetragonal crystals of PRV NEC appear after one day and reach their final size after 2 days (Figure 2B).
- Soak PRV NEC crystals briefly in the reservoir solution containing 15% meso-erythritol as a cryoprotectant and flash freeze in liquid nitrogen.
Representative data
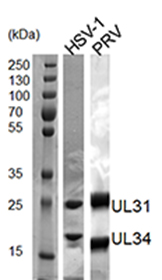
Figure 1. 12% SDS-PAGE analysis of pure HSV-1 and PRV NEC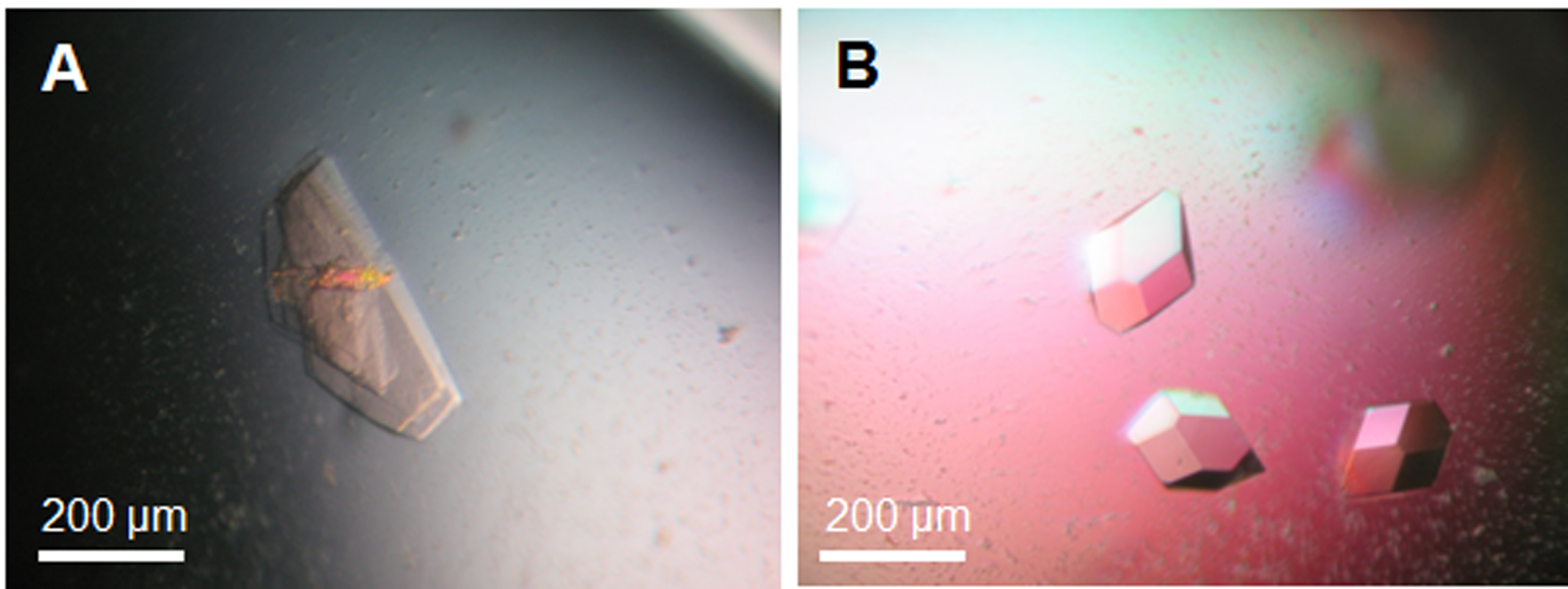
Figure 2. Crystals of HSV-1 NEC (A) and PRV NEC (B). Crystals were grown in hanging drops at 22 °C. The reservoir solution for growing HSV-1 NEC crystals contained 10% PEG 8000, 0.1 M Na citrate, pH 5.6, 5 mM NiCl2, and for PRV NEC crystals 18% PEG 3350, 0.3 M NaSCN, 0.3 M NaCl.
Notes
Crystallization of HSV-1 NEC depends on the formation of a disulfide bond involving Cys278UL31. Therefore, not more than 0.5 mM TCEP may be used during purification. Both protein complexes should be purified and frozen within 40 h to prevent degradation and aggregation.
Recipes
- TB medium (per liter), autoclaved or filter-sterilized
12 gtryptone
24 g yeast extract
4 ml glycerol
0.231 g KH2PO4
1.254 g K2HPO4 - Lysis buffer
50 mM HEPES, pH 7.0
500 mM NaCl
10% glycerol
0.5 mM TCEP - Gel filtration buffer
20 mM HEPES, pH 7.0
100 mM NaCl
0.5 mM TCEP
Acknowledgments
This work was funded by the NIH grants 1R21AI097573 and 1R01GM111795 (E.E.H.), the Burroughs Wellcome Fund (E.E.H.), and by the postdoctoral fellowship from the Deutsche Forschungsgemeinschaft GZ: BI 1658/1-1 (J.M.B.). Parts of this protocol were published previously in Bigalke et al., 2014 and Bigalke & Heldwein, 2015.
References
- Bigalke, J. M., Heuser, T., Nicastro, D. and Heldwein, E. E. (2014). Membrane deformation and scission by the HSV-1 nuclear egress complex. Nat Commun 5: 4131.
- Bigalke, J. M. and Heldwein, E. E. (2015). Structural basis of membrane budding by the nuclear egress complex of herpesviruses. EMBO J 34(23): 2921-2936.
- Pitts, J. D., Klabis, J., Richards, A. L., Smith, G. A. and Heldwein, E. E. (2014). Crystal structure of the herpesvirus inner tegument protein UL37 supports its essential role in control of viral trafficking. J Virol 88(10): 5462-5473.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Bigalke, J. M. and Heldwein, E. E. (2016). Expression, Purification and Crystallization of the Herpesvirus Nuclear Egress Complex (NEC) . Bio-protocol 6(14): e1872. DOI: 10.21769/BioProtoc.1872.
- Bigalke, J. M. and Heldwein, E. E. (2015). Structural basis of membrane budding by the nuclear egress complex of herpesviruses. EMBO J 34(23): 2921-2936.
Category
Biochemistry > Protein > Isolation and purification
Biochemistry > Protein > Expression
Biochemistry > Protein > Structure
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link











