- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Measurement of Mitochondrial DNA Release in Response to ER Stress
Published: Vol 6, Iss 12, Jun 20, 2016 DOI: 10.21769/BioProtoc.1839 Views: 17822
Reviewed by: Ivan ZanoniAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Flow-cytometric Detection of Low-level Reactive Oxygen Species in Cell Lines and Primary Immune Cells
Kevin Bode [...] Heiko Weyd
Sep 5, 2020 7243 Views
Sample Preparation and Integrative Data Analysis of a Droplet-based Single-Cell ATAC-sequencing Using Murine Thymic Epithelial Cells
Tatsuya Ishikawa [...] Taishin Akiyama
Jan 5, 2023 2415 Views
Proliferation Assay Using Cryopreserved Porcine Peripheral Mononuclear Cells Stimulated With Concanavalin A and Analyzed With FCS ExpressTM 7.18 Software
Marlene Bravo-Parra [...] Luis G. Giménez-Lirola
Jun 5, 2025 2620 Views
Abstract
Mitochondria house the metabolic machinery for cellular ATP production. The mitochondrial network is sensitive to perturbations (e.g., oxidative stress and pathogen invasion) that can alter membrane potential, thereby compromising function. Healthy mitochondria maintain high membrane potential due to oxidative phosphorylation (Ly et al., 2003). Changes in mitochondrial function or calcium levels can cause depolarization, or a sharp decrease in mitochondrial membrane potential (Bernardi, 2013). Mitochondrial depolarization induces opening of the mitochondrial permeability transition pore (MPTP), which allows release of mitochondrial components like reactive oxygen species (mtROS), mitochondrial DNA (mtDNA) or intermembrane space proteins into the cytosol (Martinou and Green, 2001; Tait and Green, 2010; Bronner and O'Riordan, 2014). These contents trigger inflammation, and can lead to cell death (West et al., 2011). Both mtROS and cytosolic mtDNA contribute to the activation of inflammasomes, multiprotein complexes that process the proinflammatory cytokines, IL-18 and IL-1β. Studies indicate that cytosolic mtDNA in particular can bind two different inflammasome sensors, AIM2 and NLRP3, leading to inflammasome activation (Burckstummer et al., 2009; Hornung and Latz, 2010). In this protocol, you will be able to specifically extract cytosolic mtDNA and quantify the amount using a qPCR assay.

Figure 1. Flowchart for extracting, purifying, and amplifying cytosolic mtDNA
Part I. Extraction and purification of cytosolic mtDNA
Materials and Reagents
- For extraction (see Figure 1)
- 1.5 ml microcentrifuge tubes (Denville Scientific Inc., catalog number: C2170 )
- Gloves
- 6 well-plates, tissue culture-treated (Corning, catalog number: 3506 )
- Cell lifter (Biologix Group Limited, catalog number: 70-2180 )
- Murine immortalized bone marrow derived macrophages (Bernardi, 2013)
- Thapsigargin, 97%, ACROS OrganicsTM (Thermo Fisher Scientific, Fisher ScientificTM, catalog number: AC328570010 ) (Bronner et al., 2015)
- 1% NP-40 (Igepal CA-630) (Sigma-Aldrich, catalog number: I8896 ) (Bronner and O'Riordan, 2014)
- DPBS (Thermo Fisher Scientific, GibcoTM, catalog number: 14040-133 ) (Burckstummer, 2009)
- DMEM (Thermo Fisher Scientific, GibcoTM, catalog number: 11965-092 )
- Fetal bovine serum (FBS) (Thermo Fisher Scientific, GibcoTM, catalog number: 10437-028 )
- Medium (see Recipes)
- NP-40 (Igepal CA-630) solution (see Recipes)
- Thapsigargin stock solution (see Recipes)
- 1.5 ml microcentrifuge tubes (Denville Scientific Inc., catalog number: C2170 )
- For purification
Equipment
- Centrifuge (Thermo Fisher Scientific, Fisher ScientificTM, model: accuSpinTM Micro 17 )
Procedure
- Seed 1 x 106 cells in 2 ml per well into 6-well plate in the appropriate medium.
- The following day (~16 h later), aspirate medium and add medium containing thapsigargin (10 µM in 2 ml medium for 4-6 h) (Hornung and Latz, 2010).
- At relevant times post-treatment (e.g., 2 h, 4 h and 8 h) wash cells with 1x DPBS once then aspirate 1x DPBS.
- Add 1% NP-40 (100 µl) to each well and scrape cells.
- Place lysates into prelabeled microcentrifuge tubes and incubate on ice for 15 min (Livak and Schmittgen, 2001).
- Spin lysates at 13,000 rpm (16,000 x g) for 15 min at 4 °C to pellet the insoluble fraction.
- Transfer supernatant (the cytosolic fraction) to a new tube and discard the pellet.
- Use supernatant in the next step to extract cytosolic mitochondrial DNA.
- Use DNeasy Blood & Tissue Kit to purify mitochondrial DNA from the cytosolic fraction according to the manufacturer’s instructions (Ly et al., 2003).
- Add 100 µl ethanol (96-100%) to the cytosolic fraction and continue to step 4 in the DNeasy Blood & Tissue Kit protocol.
Notes
- This protocol is optimized for using immortalized bone marrow derived macrophages (iBMDM) (Bronner et al., 2015). Protocol can be optimized for chosen experimental cell type.
- Thapsigargin serves as a positive control for triggering mitochondrial DNA release. Thapsigargin prevents the uptake of calcium into the endoplasmic reticulum (ER) by blocking SERCA channels. Under these conditions, the ER leaks calcium without being able to replenish its calcium stores, and consequently calcium accumulates in the cytosol or mitochondria. Mitochondrial calcium overload triggers depolarization, leading to the release mitochondrial contents into the cytosol.
- Make a 10% NP-40 stock solution and dilute to 1% NP-40.
- Use DPBS that contains calcium and magnesium to ensure that cells will remain attached during the wash step.
- Prelabeled microcentrifuge tubes do not need to be prechilled.
- 1 x 106 to 1 x 107 (seeded in 100 mm tissue culture dishes) cells will yield 1-20 µg of mtDNA.
Recipes
- Medium
DMEM
10% Fetal Bovine Serum (FBS)
Add 50 ml heat-inactivated FBS to 450 ml of DMEM.
Notes: - FBS is heat inactivated for 30 min at 55 °C.
- This medium has been optimized for iBMDM. Use medium optimized for experimental cell type.
- NP-40 (Igepal CA-630) solution
- For 10% NP-40, add 1 ml of NP-40 to 9 ml of dH2O.
- For 1% NP-40, add 1 ml of 10% NP-40 to 9 ml of dH2O.
- For 10% NP-40, add 1 ml of NP-40 to 9 ml of dH2O.
- Thapsigargin stock solution
- The concentration stated above has been optimized for iBMDM. The concentration used on different cell types must be optimized.
- Stock solution of thapsigargin remains usable up to one year after reconstitution and can be aliquoted and stored at -20 °C
- Stock solution of thapsigargin is 5 mM (1 mg in 307 µl DMSO). Dilute thapsigargin (4 µl into 2 ml of medium) into medium that will be added to the wells.
Part II. Amplificationof mtDNA via qPCR
After extracting DNA from the cytosolic fraction, quantitative PCR is employed to measure cytosolic mitochondrial DNA.
Materials and Reagents
- 96 well qPCR plate (Denville Scientific Inc., catalog number: C18096) (Bernardi, 2013)
- 1.5 ml microcentrifuge tubes (Denville Scientific Inc., catalog number: C2170 )
- Gloves
- Brilliant II SYBR® green with low ROX (Agilent Technologies, catalog number: 600830 )
- Sterilized double distilled water
- Primers (see sequences below)
Equipment
- qPCR machine (Stratagene MX3000P)
Procedure
- Prepare the SYBR qPCR master reaction mix as follows in Table 1 for mitochondrial genes of interest and internal control (housekeeping gene for used here is 18S rDNA) in 1.5 ml microcentrifuge tubes:
Table 1. Master Mix recipe for quantifying mtDNA release into cytosol via qPCR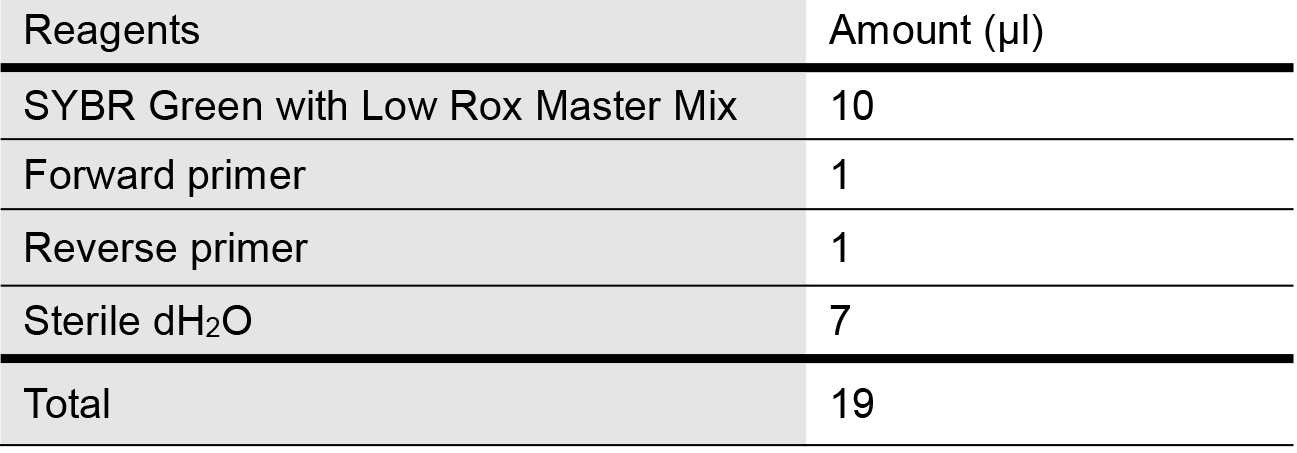
The following primers were used:
Cytochrome c oxidase I (mt gene)
Forward: 5’-GCCCCAGATATAGCATTCCC-3’
Reverse: 5’-GTTCATCCTGTTCCTGCTCC-3’
18S rDNA (internal control)
Forward: 5’-TAGAGGGACAAGTGGCGTTC-3’
Reverse: 5’-CGCTGAGCCAGTCAGTGT-3’ - Mix gently but thoroughly by pipetting.
- Incubate reaction mix on ice until ready to use.
- Aliquot DNA samples (1 µl) into wells (run in triplicates) (Bronner et al., 2015).
- Aliquot corresponding Master Mix (19 µl) into the corresponding wells (Bronner and O'Riordan, 2014).
- Use the thermal cycler program as seen below in Table 2:
Table 2. Thermal cycler program for mtDNA release qPCR
- Once qPCR is complete, calculate relative fold change in cytochrome c oxidase I from the Ct values.
Representative data
Adapted method for calculating relative fold change (Livak and Schmittgen, 2001) between thapsigargin-treated and untreated Ct values that are represented in Table 3:
Table 3. Representative values from mtDNA release qPCR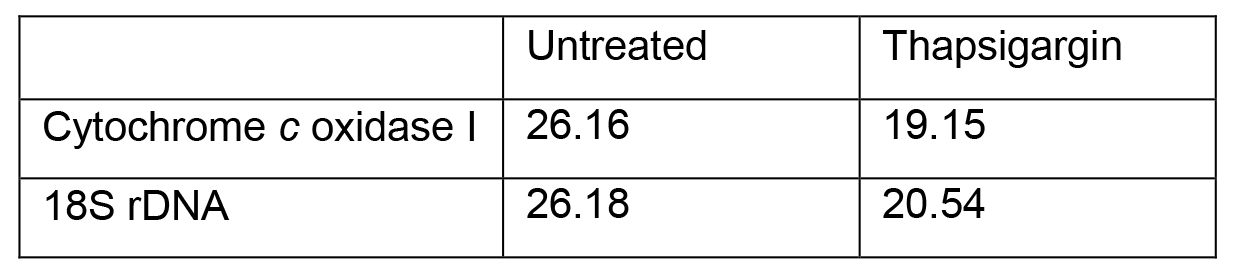
- ΔCt Treatment = Target gene - Reference gene
ΔCt Treatment (thapsigargin) = 19.15 - 20.54
ΔCt Control = Target gene - Reference gene
ΔCt Control (untreated) = 26.16 - 26.18 - ΔΔCt = ΔCt Treatment - ΔCt Control
ΔΔCt = -1.39 - (-0.02)
ΔΔCt = -1.37 - Relative Fold Change = 2-(ΔΔCt)
Relative Fold Change = 2-(-1.37)
Relative Fold Change = 2.6
This result indicates that thapsigargin treatment results in a 2.6 fold increase in cytosolic mtDNA compared to untreated (Figure 2).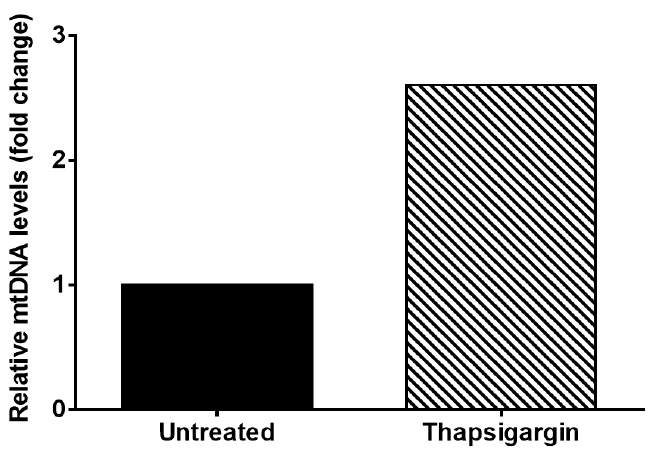
Figure 2. Graph of representative data from Table 3. Graph depicts cytosolic mtDNA relative fold change seen in Thapsigargin treated iBMDM when compared to untreated iBMDM.
Notes
- Use plates that are optimized for qPCR machine.
- The total reaction volume is 20 µl. When adding DNA ensure the amount is consistent between samples.
- 10 ng and 2 µg represent the minimum and maximum amount of DNA to use for qRT-PCR analysis
- Pipette as accurately as possible during steps 4 and 5, since small variations in volumes can increase variability in the qPCR results.
Acknowledgments
We acknowledge financial support from the University of Michigan Rackham Graduate School (D. N. B.), the UM Genetics Training Program (D. N. B., GM007544). This research was supported by funding from the NIH to M. X. D. O. (AI101777). The funders had no role in study design, data collection, and analysis, decision to publish, or preparation of the manuscript.
We acknowledge that this protocol was adapted (Nakahira et al., 2011) and modified for use with immortalized bone marrow macrophages infected by a bacterial pathogen.
References
- Bernardi, P. (2013). The mitochondrial permeability transition pore: a mystery solved? Front Physiol 4: 95.
- Bronner, D. N., Abuaita, B. H., Chen, X., Fitzgerald, K. A., Nunez, G., He, Y., Yin, X. M. and O'Riordan, M. X. (2015). Endoplasmic reticulum stress activates the inflammasome via NLRP3- and Caspase-2-driven mitochondrial damage. Immunity 43(3): 451-462.
- Bronner, D. N. and O'Riordan, M. X. (2014). A near death experience: Shigella manipulates host death machinery to silence innate immunity. EMBO J 33(19): 2137-2139.
- Burckstummer, T., Baumann, C., Bluml, S., Dixit, E., Durnberger, G., Jahn, H., Planyavsky, M., Bilban, M., Colinge, J., Bennett, K. L. and Superti-Furga, G. (2009). An orthogonal proteomic-genomic screen identifies AIM2 as a cytoplasmic DNA sensor for the inflammasome. Nat Immunol 10(3): 266-272.
- Hornung, V. and Latz, E. (2010). Intracellular DNA recognition. Nat Rev Immunol 10(2): 123-130.
- Livak, K. J. and Schmittgen, T. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25(4): 402-408.
- Ly, J. D., Grubb, D. R. and Lawen, A. (2003). The mitochondrial membrane potential (deltapsi(m)) in apoptosis; an update. Apoptosis 8(2): 115-128.
- Martinou, J. C. and Green, D. R. (2001). Breaking the mitochondrial barrier. Nat Rev Mol Cell Biol 2(1): 63-67.
- Nakahira, K., Haspel, J. A., Rathinam, V. A., Lee, S. J., Dolinay, T., Lam, H. C., Englert, J. A., Rabinovitch, M., Cernadas, M., Kim, H. P., Fitzgerald, K. A., Ryter, S. W. and Choi, A. M. (2011). Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat Immunol 12(3): 222-230.
- Tait, S. W. and Green, D. R. (2010). Mitochondria and cell death: outer membrane permeabilization and beyond. Nat Rev Mol Cell Biol 11(9): 621-632.
- West, A. P., Shadel, G. S. and Ghosh, S. (2011). Mitochondria in innate immune responses. Nat Rev Immunol 11(6): 389-402.
Article Information
Copyright
© 2016 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Bronner, D. N. and O’Riordan, M. X. (2016). Measurement of Mitochondrial DNA Release in Response to ER Stress. Bio-protocol 6(12): e1839. DOI: 10.21769/BioProtoc.1839.
Category
Immunology > Immune cell function > General
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









