- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Loading of Cells with Fluorescent Probe to Study Intracellular Acid-base Homeostasis in Lactic Acid Bacteria
Published: Vol 5, Iss 2, Jan 20, 2015 DOI: 10.21769/BioProtoc.1380 Views: 13062
Reviewed by: Fanglian HeKanika Gera

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Quantification of Bacterial Polyhydroxybutyrate Content by Flow Cytometry
Antonio Lagares (Jr.) and Claudio Valverde
Dec 5, 2017 8145 Views
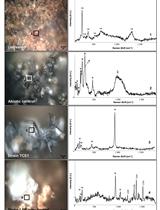
Soluble and Solid Iron Reduction Assays with Desulfitobacterium hafniense
Lucrezia Comensoli [...] Edith Joseph
Sep 5, 2018 6717 Views
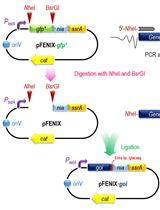
A SsrA/NIa-based Strategy for Post-Translational Regulation of Protein Levels in Gram-negative Bacteria
Gonzalo Durante-Rodríguez [...] Pablo I. Nikel
Jul 20, 2020 4380 Views
Abstract
Here we describe a protocol which we have used to study the homeostasis intracellular in vivo in lactic acid bacteria (LAB) using a fluorescent probe. This type of probes can be used for determining changes in the pH of cytoplasm with high sensitivity, temporal resolution and technical simplicity as well as accessing the rate of change of intracellular pH in response to a stimulus from kinetic measurements on short time scales (Breeuwer et al., 1996; Molenaar et al., 1991). This protocol has been designed to measure the intracellular pH using the pH-sensitive fluorescent probe 2´,7´-bis-(2-carboxyethyl)-5(and-6)-carboxyfluorescein (BCECF) in LAB, Enterococcus faecalis (E. faecalis), Lactococcus lactis (L. lactis) and Lactobacillus casei (L. casei).
Keywords: Fluorescent probeMaterials and Reagents
- Lactococcus lactis IL1403
- Enterococcus faecalis JH2-2
- Lactobacillus casei (ATCC, catalog number: 334 )
- 2´,7´-bis-(2-carboxyethyl)-5(and-6)-carboxyfluorescein (BCECF) (acid form) (Life Technologies, Molecular Probes®)
- Luria-Bertani broth (Sigma-Aldrich, catalog number: L3147 )
- MRS broth (Sigma-Aldrich, catalog number: 69966 )
- Citrate (Sigma-Aldrich, catalog number: C7254 )
- Malic acid (Supelco, catalog number: 46940U )
- Pyruvate (Sigma-Aldrich, catalog number: P2256 )
- D-(+)-glucose (Sigma-Aldrich, catalog number: G8270 )
- D-(+)-galactose (Sigma-Aldrich, catalog number: G0750 )
- Electrode filling solution (Orion, catalog number: 900011 )
- Na2HPO4 (Merck KGaA, catalog number: 1006559 )
- NaH2PO4 (Merck KGaA, catalog number: 106349 )
- HCl (Merck KGaA, catalog number: 100317 )
- NaOH (Merck KGaA, catalog number: 106462 )
- Triton X100 detergent (Merck KGaA, catalog number: 648466 )
- Valinomycin (Sigma-Aldrich, catalog number: V0627 )
- Nigericin (Sigma-Aldrich, catalog number: N7143 )
- BCECF stock solution (see Recipes)
- 50 mM potassium phosphate (KPi) buffer (see Recipes)
Equipment
- Stove Lab Tech (LIB-080M)
- Centrifuge for Eppendorf tubes (Eppendorf, model: 5418 )
- Centrifuge Sorvall ST 16R for centrifugation of Falcon tube under refrigerated conditions (Thermo Fisher Scientific)
- Fluorescence spectrophotometer (PerkinElmer, model: LS55 )
- Thermostatic bath (Lauda Alpha, model: RA8 )
- pH/MV meters (Orion, model: 420A )
- 4 M KCl saturated with AgCl for combination Ag/AgCl pH electrode (Orion, catalog number: 900011)
- 5 ml Quartz cuvette (optical path 1 cm)
- pH microelectrode (Horiba, model: 9669-10D )
Procedure
- Growth conditions
- Enterococcus faecalis cultures were grown at 37 °C without agitation in 100 ml stoppered bottles that contained 20 to 50 ml of LB medium (LB), initial pH 7.0 (pH adjusted by adding 5 M HCl or 5 M NaOH as correspond employing a pH meter) supplemented with organic acid (citrate, malate or pyruvate) and/or different carbon source.
- Lactococcus lactis cultures were routinely grown at 30 °C without shaking in 100 ml sealed bottles containing 20 ml of lactose free M17 medium, supplemented with 25 mM glucose (M17G). Overnight cultures prepared in this way were used to inoculate fresh M17 adjusted to different initial pH values as indicated. The medium was supplemented with organic acid and/or different carbon source.
- Lactobacillus casei was grown overnight at 37 °C in modified MRS broth adjusted to pH 6.0. Cells were grown in screw-capped tubes of 50 ml without shaking at 37 °C for about 15 hours inoculated with 15 μl of a preculture in stationary phase mMRS supplemented with 25 mM glucose (mMRSG) stored at -80 °C.
- Growth of the microbial cultures was followed by measurement of the optical density at a wavelength of 660 nm (OD660).
- Cells were harvested in mid-exponential growth phase when the optical density was 0.6-0.8, centrifugate for 10 min at 1,600 x g at 4 °C.
- The cells were washed twice with 50 mM potassium phosphate (KPi) (pH 5.5) at 4 °C, and finally resuspended in 1 ml the same buffer. For each wash, we used the same volume of solution that the initial culture volume.
The same KPi buffer condition (50 mM KPi, pH 5.5, kept at 4 °C) is used in the rest of steps in loading of cells with the BCECF probe.
- Enterococcus faecalis cultures were grown at 37 °C without agitation in 100 ml stoppered bottles that contained 20 to 50 ml of LB medium (LB), initial pH 7.0 (pH adjusted by adding 5 M HCl or 5 M NaOH as correspond employing a pH meter) supplemented with organic acid (citrate, malate or pyruvate) and/or different carbon source.
- Loading of cells with the BCECF probe
- The different OD-cultures were matched in such way to start with the loading-probe protocol with the same amount of cells.
- Thus, the cells were centrifugated at maximum speed 24,900 x g for 1 min at room temperature.
- Finally the cells were resuspended in 20 µl of KPi solution. To be loaded with BCECF, 1 μl of 10 mM BCECF was added to said resuspension cells. Then 2.5 µl of 0.5 M HCl was added and incubated for 10 min in the dark at room temperature.
- It was then neutralized with the addition of 1 ml of KPi ice solutions and washed twice with 1 ml of KPi. The centrifugation between wash steps was at maximum speed 24,900 x g for 1 min at room temperature.
- Centrifugated between each wash step at 24,900 x g for 1 min. Finally the cells were resuspended in 100-150 μl of KPi solutions and stored on ice until use.
- The different OD-cultures were matched in such way to start with the loading-probe protocol with the same amount of cells.
- Fluorescent determination
- For each experiment, 10 µl of charged BCECF cells were resuspended in 3 ml of 50 mM KPi solution at indicated pH in a 5 ml quartz cuvette (optical path 1 cm) and equilibrated at 37 °C for 2 min.
- The sample was stirred with a suitable magnetic strip and the fluorescence signal was monitored every 1 sec in a fluorometer.
- Fluorescence emissions were recorded at 525 nm with excitation wavelength at 503 nm (slit widths were 16 and 4 nm, respectively). The opening of the sample chamber causes data loss during the first 6-7-8 sec after addition of each substrate to be tested that was made in the cuvette.
- For each experiment, 10 µl of charged BCECF cells were resuspended in 3 ml of 50 mM KPi solution at indicated pH in a 5 ml quartz cuvette (optical path 1 cm) and equilibrated at 37 °C for 2 min.
- Conversion of the fluorescence signal in pH
- The conversion of the fluorescence signal to cytoplasm pH was performed by titration curves as shown in the Note 1. This procedure was performed for each batch of cells loaded. The loaded cells were resuspended in buffer 50 mM KPi at pH 3.0.
- In the same way, the determinations were performed with addition of 2% (v/v) Triton X-100, 75 μM valinomycin and nigericin for permeabilizing the membranes to reach the ionic balance between the intracellular and the external solution. This cell mixture was incubated for 2 min, at 37 °C (work temperature) with agitation until the fluorescent signal was stabilized. Thus, the measurement of fluorescence and pH (with a pH microelectrode) is performed per each addition of 5 of 0.1 N NaOH solutions. To each pH value we get a fluorescent signal value, measured in separate steps. Each determination is a point of the titration curves and covers the pH range from 3.0 to 11.0.
Figure 1 shows titration curves for fluorometric probe BCECF in L. lactis and E. faecalis. The cells loaded were titrated prior permeabilization of the cells with 2% (v/v) Triton X-100, 75 mM valinomycin and 75 mM nigericin. The pKa, Ifmín and Ifmax parameters were determined from nonlinear fit of experimental data.
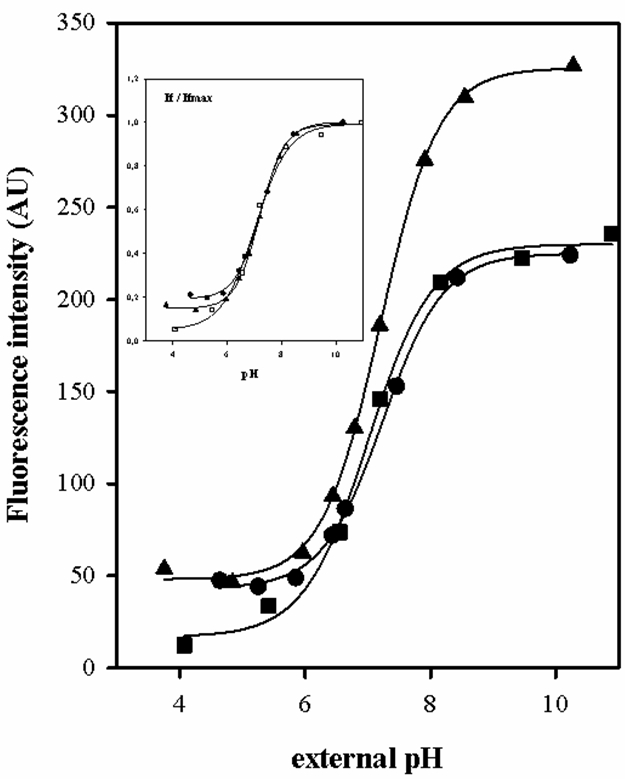
Figure 1. Fluorometric titration curves of the BCECF probe, fluorescence intensity as a function of external pH of 50 mM KPi buffer where the cells were resuspended. Wavelength excitation: 503 nm, wavelength emission: 525 nm. The pKa of BCECF was determined in cells of wild type L. lactis IL1403 (- ▲ -), L. lactis ILGR1 (IL1403 isogenic derivative als -α acetolactate sinthase- mutant) (- ● -) and E. faecalis JH2-2 (- ■ -), properly loaded with BCECF.
- This permeabilization allowed the electrochemical decoupling of the cell so that the pHint and pHext are equilibrated. Table 1 shows the four parameters of the non-linear fit curve (If vs pH) obtained for BCECF in each microorganism.
- Parameters a, d (Ifmin and Ifmax) and c (pKa) were then used in the transformation of the of fluorescence intensity to pH values.
Table 1. Parameter values of nonlinear adjustment from titration curves of BCECF probe
aAUif: Arbitrary units of fluorescence intensity
L. lactis
IL1403
L. lactis
ILGR1
E. faecalis
JH2-2
a (Ifmax) (AUifa)
277 ± 4
181 ± 4
213 ± 10
b (if.pH-1)
0.46 ± 0.02
0.48 ± 0.03
0.51 ± 0.08
c (pKa)
7.20 ± 0.02
7.24 ± 0.04
7.00 ± 0.08
d (Ifmin) (AUifa)
48 ± 2
43 ± 2
17 ± 7
- The conversion of the fluorescence signal to cytoplasm pH was performed by titration curves as shown in the Note 1. This procedure was performed for each batch of cells loaded. The loaded cells were resuspended in buffer 50 mM KPi at pH 3.0.
- Intracellular pH measurement
The following are examples of using BCECF for determination of intracellular pH in LAB as result the metabolism of organic acids and/or sugars. Figure 2 shows the intracellular pH variation during the metabolism of different carbon sources in E. faecalis JH2-2 (Espariz et al., 2011; Repizo et al., 2013).
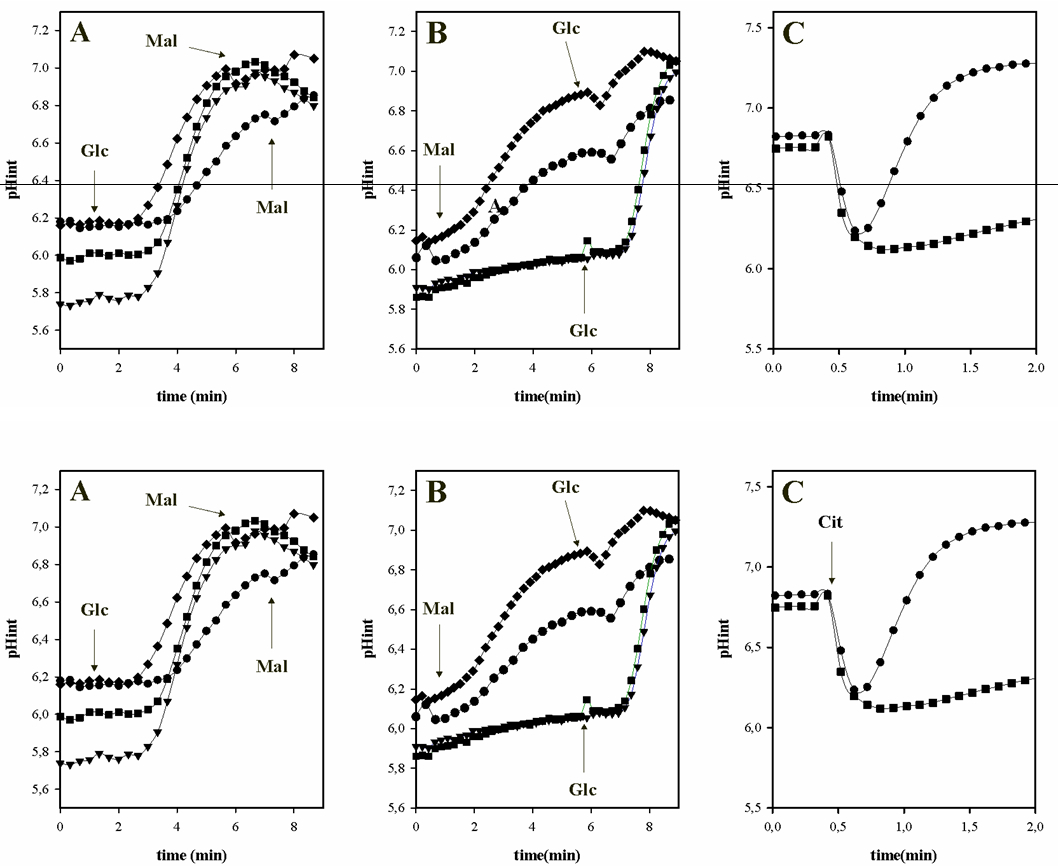
Figure 2. Effect of citrate and malate metabolism on the intracellular pH of E. faecalis JH2-2. A and B, cells were grown in LB supplemented with: 28 mM malate (- ♦ -); 28 mM malate and 10 mM arginine (- ● -); 28 mM glucose (- ▼ -); and 28 mM glucose and malate (- ■ -). C, cells were grown in LB: basal (- ■ -); and supplemented with 30 mM citrate (- ● -). Arrows indicated the substrates additions: 10 mM malate (Mal); 10 mM glucose (Glc) or 10 mM citrate (Cit) in loaded cell with BCECF and resuspended in 50 mM KPi solution (pH 6.5).
Figure 3 shows the intracellular pH variation during the metabolism of different carbon sources in L. casei ATCC 334 (Mortera et al., 2013).
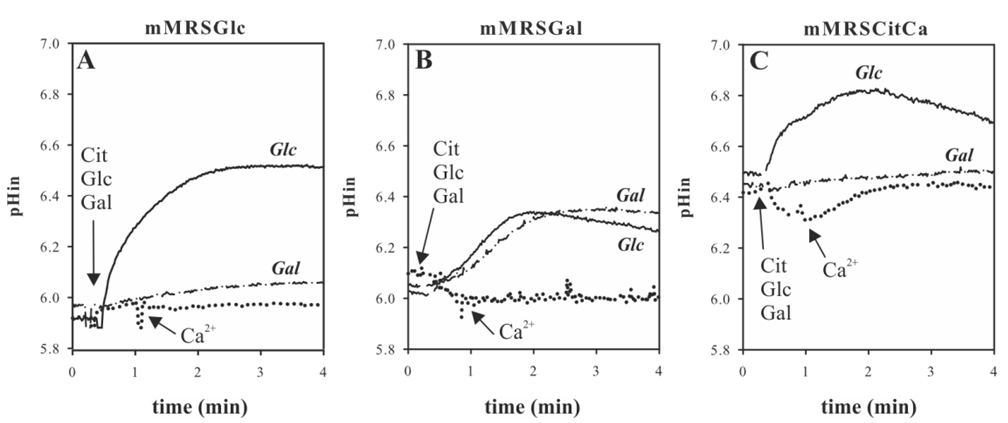
Figure 3. Energetics of carbohydrate and Ca2+-citrate metabolism in Lb. casei. Lb. casei ATCC 334 was grown in mMRSGlc A, mMRSGal B and mMRSCitCa C. The cells were harvested at an OD660 of 0.6 and loaded with BCECF for intracellular pH (pHin) measurements (A, B, and C). For intracellular pH measurements, 5 mM glucose (solid line), 5 mM galactose (dashed line), or 2 mM citrate and then 2 mM Ca2+ (dotted line) was added at the time point indicated by the arrow.
Figure 4 shows the intracellular pH variation in L. lactis IL1403 and the mutant ILGR1 product the metabolism of different carbon source (Zuljan et al., 2014).
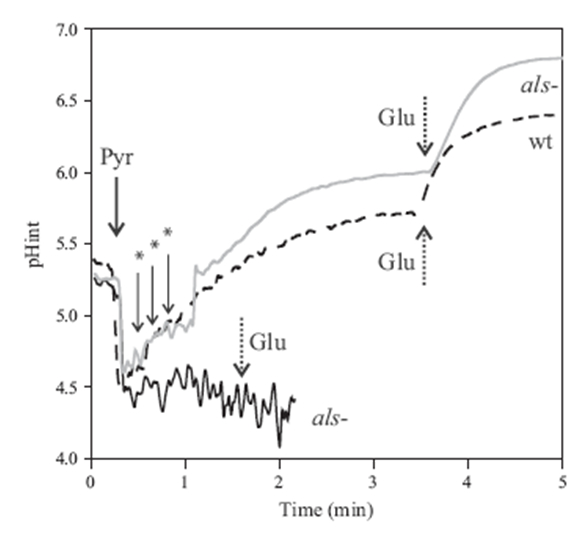
Figure 4. L. lactis intracellular pH variations in response to pyruvate. L. lactis IL1403 (dotted line, wt) and L. lactis ILGR1 (continuous line) strains were grown in M17G up to exponential phase. Then, cells were loaded with the BCECF fluorescent probe and resuspended in phosphate buffer at 4.5. Pulses of 50 mM pyruvate (Pyr), 3 mM glucose (Glu) or 0.1 N NaOH (*) were added at the times indicated by the arrows (gray line).
Notes
- We can represent the fluorescent activity of the probe from its acid-base equilibrium; Where SF is the fluorescent specie, which is the one that allows us follow the pH changes in the environment:
HSF ↔ SF- + H+ Ka = [SF-]x [H+]x/[HSF]x
Figure 5A shows a titration curve of a generic probe where fluorescence intensity versus pH is plotted. For a given value Ifx fluorescence intensity corresponds to a pHx value within the range in the titration curve (pHx; Ifx). In this case the concentrations of both species will be given by:
[SF-]x = Ifx – Ifmin
[HSF]x = Ifmax – Ifx
Replacing in the expression of the Ka, have:
pHx = pKa + log {(Ifx – Ifmin)/(Ifmax – Ifx)}
This is an expression which allows us to transform each Ifx at pHint scale at a certain time x (tx; Ifx) in a kinetic experiment, as shown in Figure 5B.
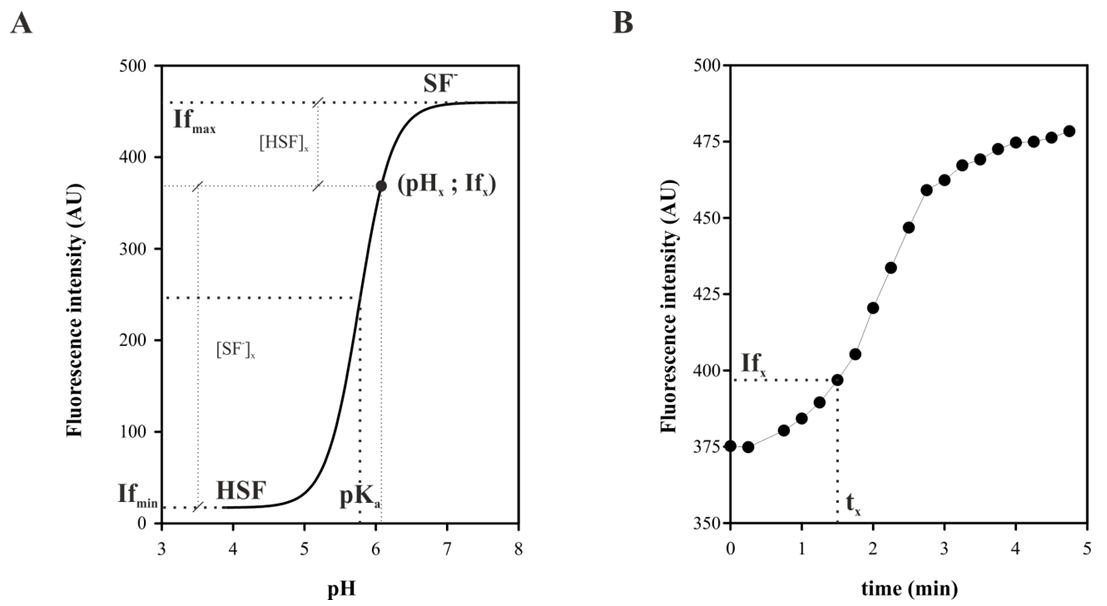
Figure 5. If conversion to pH scale. Schematic representation of the correlation between of a titration curve A and fluorescence emission kinetics, for a generic fluorescent probe in BAL strain. A point (pHx; Ifx) indicated in A that is linked to the point (tx; Ifx) to B as explained in the text.
The pKa, Ifmax and Ifmín parameters of (1) associated with the titration curve are obtained from nonlinear four parameter fit using the following expression:
f (x) = {(a − d) / [1 + (x/c)b]} + d
where a and d are the asymptotic maximum and minimum values corresponding to Ifmax and Ifmin for the probe, c the inflection point corresponding to the pKa of the probe and b the slope that gives us an idea the sensitivity of the system. The pH range for the use of the probe will be set to the value of pKa and system sensitivity.
- Measurement of the pH intracellular in E. faecalis loaded with BCECF when the extracellular pH is below to 5.5. As shown in Figure 6, the response of JH2-2/BCECF cells was to be maximal at pHext 7.5 while is to virtually unable to be detected at pHext under 6.0. Thus, it is impossible to study organic acid metabolism in E. faecalis with BCECF probe at pHext less than 6.0 (data not shown).
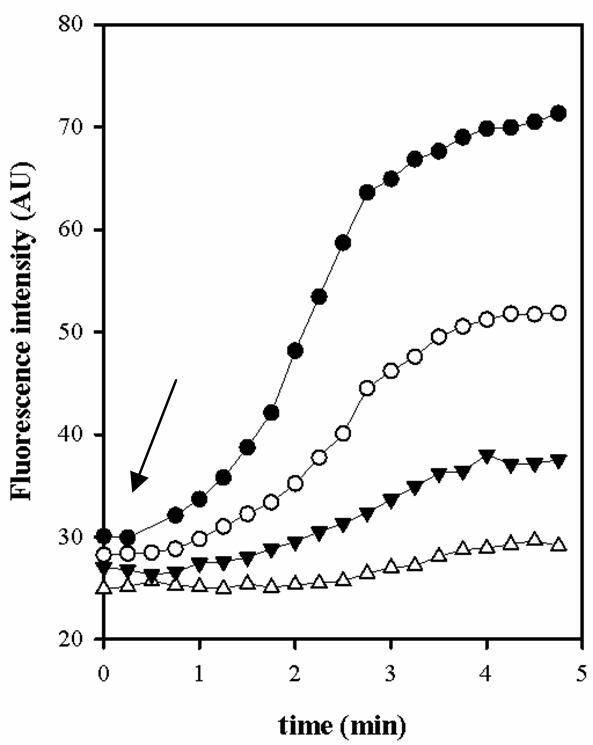
Figure 6. Fluorescence intensity of E. faecalis cells loaded with BCECF at different pHext. pHext: 6.0 (- Δ -), 6.5 (- ▼ -), 7.0 (- ○ -) and 7.5 (- ● -). The arrow indicates a pulse of glucose (Glc) 5 mM.
E. faecalis, unlike other species of BAL, are unable to maintain a ΔpH with the more acidic external media when the cells are not energized. These results demonstrate that for E. faecalis in very acidic conditions, when pH below 5.5, it is necessary to use different probes. In those conditions we used the CDCFD fluorescent probe in our experiment (Espariz et al., 2011).
Recipes
- BCECF stock solution
10 mM BCECF in dimethyl sulfoxide (DMSO) anhydrous
Stored at -20 °C
- 50 mM potassium phosphate (KPi) buffer
Mix 1 M K2HPO4 and 1 M KH2PO4 to a desired final pH (4.5, 5.8, 6.0, 6.5, 7.0 or 7.5) and dilute 20x with distilled H2O
mMRS: Original composition without glucose, ammonium citrate, and Tween 80
mMRSCitCa: mMRS supplemented with 30mM sodium citrate and 10mM chloride calcium
mMRSGlc: mMRS supplemented 0.5% (wt/vol) glucose
mMRSGal: mMRS supplemented 0.5% (wt/vol) galactose
The medium was adjusted to pH 6.0
Acknowledgments
This work was supported by grants from the Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT) and CONICET (PICT2010-1828 and PIP/2012). F.S. is a fellow of ANPCyT, P. M. is a fellow of CONICET, S. H. A. and C. M. are Career Investigators of the same institution.
References
- Breeuwer, P., Drocourt, J., Rombouts, F. M. and Abee, T. (1996). A novel method for continuous determination of the intracellular pH in bacteria with the internally conjugated fluorescent probe 5 (and 6-)-carboxyfluorescein succinimidyl ester. Appl Environ Microbiol 62(1): 178-183.
- Espariz, M., Repizo, G., Blancato, V., Mortera, P., Alarcon, S. and Magni, C. (2011). Identification of malic and soluble oxaloacetate decarboxylase enzymes in Enterococcus faecalis. FEBS J 278(12): 2140-2151.
- Molenaar, D., Abee, T. and Konings, W. N. (1991). Continuous measurement of the cytoplasmic pH in Lactococcus lactis with a fluorescent pH indicator. Biochim Biophys Acta 1115(1): 75-83.
- Mortera, P., Pudlik, A., Magni, C., Alarcon, S. and Lolkema, J. S. (2013). Ca2+-citrate uptake and metabolism in Lactobacillus casei ATCC 334. Appl Environ Microbiol 79(15): 4603-4612.
- Repizo, G. D., Blancato, V. S., Mortera, P., Lolkema, J. S. and Magni, C. (2013). Biochemical and genetic characterization of the Enterococcus faecalis oxaloacetate decarboxylase complex. Appl Environ Microbiol 79(9): 2882-2890.
- Zuljan, F. A., Repizo, G. D., Alarcon, S. H. and Magni, C. (2014). alpha-Acetolactate synthase of Lactococcus lactis contributes to pH homeostasis in acid stress conditions. Int J Food Microbiol 188: 99-107.
Article Information
Copyright
© 2015 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Mortera, P., Zuljan, F., Magni, C. and Alarcón, S. H. (2015). Loading of Cells with Fluorescent Probe to Study Intracellular Acid-base Homeostasis in Lactic Acid Bacteria. Bio-protocol 5(2): e1380. DOI: 10.21769/BioProtoc.1380.
Category
Microbiology > Microbial metabolism > Other compound
Cell Biology > Cell-based analysis > Ion analysis
Cell Biology > Cell metabolism > Other compound
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link









