- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Quantification of Extracellular Ammonium Concentrations and Transporter Activity in Yeast Using AmTrac Fluorescent Sensors
Published: Vol 5, Iss 1, Jan 5, 2015 DOI: 10.21769/BioProtoc.1372 Views: 11915
Reviewed by: Belen SanzAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Live-cell Imaging by Super-resolution Confocal Live Imaging Microscopy (SCLIM): Simultaneous Three-color and Four-dimensional Live Cell Imaging with High Space and Time Resolution
Kazuo Kurokawa and Akihiko Nakano
Sep 5, 2020 5983 Views
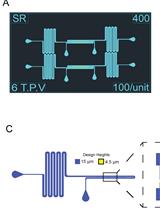
Fabrication of Microfluidic Devices for Continuously Monitoring Yeast Aging
Richard O’Laughlin [...] Nan Hao
Aug 5, 2023 1449 Views

Imaging the Entire Sexual Life Cycle of the Budding Yeast Saccharomyces cerevisiae Using a Microfluidic Platform
Taylor Kennedy [...] Orlando Argüello-Miranda
Dec 5, 2025 1477 Views
Abstract
AmTracs are the first example of “activity sensors”, since they report the activity of ammonium transporters by means of fluorescence readout in vivo (De Michele et al., 2013). AmTracs are based on a single fluorescent protein, a circularly permuted GFP (cpGFP), inserted into the cytosolic loop connecting the two pseudosymmetrical halves of the Arabidopsis and yeast plasma membrane ammonium transporters AtAMTs and ScMEP (Figure 1). Recently, FRET-based activity sensors for nitrate and peptide transporters have also been developed (Ho et al., 2014). Since transporter activity directly depends on the availability of substrate, AmTracs measure extracellular ammonium concentrations. Several versions of AmTrac exist, with different fluorescence intensity (FI) responses and affinities for ammonium, and based on different ammonium transporters (AmTrac: AtAMT1;3; AmTrac1;2: At AMT1;2; MepTrac: ScMEP2). Currently, the most useful AmTrac versions are probably AmTrac-GS (bright, with Km of 50 µM) and AmTrac-100 (a high capacity version with Km of 100 µM). The protocol for measuring ammonium concentrations in yeast cells at the fluorimeter is the same for all versions.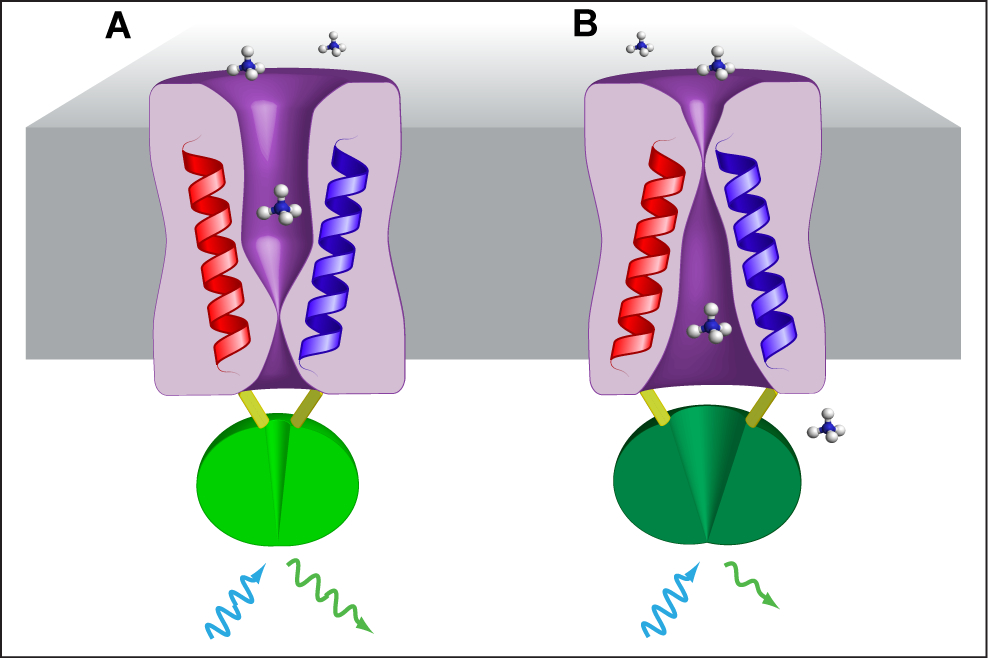
Figure 1. Model of AmTrac/MepTrac sensors and AMT transport mechanism. We propose that AMT switches between at least two distinct states during transport of ammonium: An outward, open state A and an inward, open state B. The movement of TMH-V (red) and TMH-VI (blue) is transmitted to the connecting loop, affecting the inserted cpGFP (green) and resulting in a change in fluorescence emission.
Materials and Reagents
- Yeast strain, with auxotrophic selection marker
Note: We used ura- strains 23344c: MATa ura3; or 31019b: MATa ura3 mep1Δ mep2 Δ::LEU2 mep3 Δ::KanMX2 (Marini et al., 1997). Strain 31019b lacks the endogenous ammonium transporters MEP1-3 and therefore can grow on ammonium as sole nitrogen source only when transformed with vectors harboring functional ammonium transporters. AmTrac should work in any transformable strain. - Vector pDR-F’-GW containing AmTrac under control of strong PMA promoter and ADH terminator, and with the selection marker URA3
Note: Vectors with many different AmTrac- and MepTrac versions are available through Addgene (www.addgene.org). An empty vector can be used as control.- pDR-AmTrac (Plasmid, catalog number: 47770 ) (original sensor, Km 55=µM, ΔF/F~30%)
- pDR-AmTrac-IS (Plasmid, catalog number: 47766 ) [variant with enhanced fluorescence and response (500% FI with respect to AmTrac), ΔF/F~40%]
- pDR-AmTrac1;2 (Plasmid, catalog number: 47765 ) (AmTrac based on AtAMT1;2)
- pDR-AmTrac-LS (Plasmid, catalog number: 47767 ) [variant with enhanced fluorescence and response (400% FI with respect to AmTrac), ΔF/F~50%]
- pDR-AmTrac-GS (Plasmid, catalog number: 47769 ) [variant with enhanced fluorescence and response (850% FI with respect to AmTrac), ΔF/F~40%]
- pDR-AmTrac-100 (Plasmid, catalog number: 47771 ) (high capacity sensor, Km=100 µM)
- pDR-Meptrac (Plasmid, catalog number: 47768 ) (sensor based on ScMEP2)
- pDR-Meptrac-H194E (Plasmid, catalog number: 47764 ) [high capacity, pH insensitive and pseudohyphal growth-impaired Meptrac variant (Boeckstaens et al. 2008)]
- pDf1-GW (Plasmid, catalog number 36026 ) (empty vector)
Note: It is useful to remove the GW cassette since it contains a ccdB suicidal gene.
- pDR-AmTrac (Plasmid, catalog number: 47770 ) (original sensor, Km 55=µM, ΔF/F~30%)
- Ammonium chloride (EMD Millipore, catalog numer: AX1270-7 )
- Yeast nitrogen base w/o amino acids w/o ammonium sulfate (Difco, catalog number: 233520 )
- Glucose (Fluka Analytical, catalog number: 49159 )
- Proline (L-Proline) (Sigma-Aldrich, catalog number: PO380 )
- Agar (Sigma-Aldrich, catalog number: A1296 )
- Arginine (L-Arginine monohydrochloride) (Sigma-Aldrich, catalog number: A5131 )
- MES (Sigma-Aldrich, catalog number: M2933 )
- NaOH (Sigma-Aldrich, catalog number: S5881 )
- Glycerol (BDH, catalog number: 1172-1LP )
- MilliQ water
- 100x proline solution (see Recipes)
- 50x arginine solution (see Recipes)
- SD medium (see Recipes)
- Washing buffer (see Recipes)
- Resuspension buffer (see Recipes)
Equipment
- Petri dishes (plastic, any size/brand)
- Sterile plastic tips for pipettes (1 ml and 200 µl sizes)
- 50 ml sterile plastic tubes (like Falcon®, any brand)
- Tube rack (any brand)
- Surgical tape (3 M Micropore, catalog numer: 1535-5 )
- 96 well ELISA microplate (flat bottom) (Greiner Bio-One GmbH, catalog numer: 650 101 )
Multichannel (8) pipette (for 50 µl) (Sartorious, catalog numer: 725240 ) - Reservoir with 12 compartments (VWR international, catalog numer: 80092-466 )
- Autoclave (tuttnauer Brinkmann, model: 3850 E )
- Orbital shaker, with temperature and velocity control (New Brunswick Scientific, model: innova 44 )
- Incubator for 28-30 °C plate incubation (VWR International)
- Sterile hood (Heraeus Holding)
- Centrifuge with swinging rotor for 50 ml tubes (Beckman Coulter, model: Allegra 25R )
- Fluorimeter for 96 well plates (for example: Tecan Trading AG, Safire or M1000 )
Procedure
- Transform yeast with the pDR-F’ vector containing the desired AmTrac. We routinely use the lithium acetate method (Gietz et al., 1992). Briefly, we add about 100 ng plasmid to a PCR tube containing 100 µl of the transformation solution (containing PEG, lithium acetate, salmon sperm DNA and yeast cells) and heat-shock at 42 °C for 13 min. The first time the reader might also transform with an empty vector, to measure background fluorescence.
- Plate the transformation on selective synthetic dextrose minimal medium (SD), with 1 mM arginine (from a filter-sterilized 50x stock dissolved in water) as sole nitrogen source. Wrap the plate in plastic cling wrap to prevent dehydration.
- Incubate the plate (upside down to prevent condensation) at 28-30 °C for 3 days.
- In the sterile hood, pick a colony with a sterile 1 ml pipette and place it in a 50 ml tube containing 5 ml SD medium, supplemented with 0.1% proline (from a filter-sterilized 100x stock dissolved in water).
- Tape the opening of the tube with surgical tape to prevent contamination, still allowing gas exchange.
- Place the tube in a rack in the orbital shaker.
- Grow for 36-48 h at 30 °C, 230 rpm. The culture should at least be turbid (OD600 >0.5). Even at saturation, the sensor still works. Avoid overgrowth though, since cells will start to die and degrade the sensors. In that case, start a new inoculum.
- Remove the tip with forceps, and centrifuge at 3,000 rcf at RT for 5 min to pellet the cells.
- Discard the supernatant by inversion. The reader should see a whitish pellet.
- Resuspend the pellet in 20 ml washing buffer, RT. The reader can ease resuspension by pipetting up and down, or by pouring a few ml first, shaking vigorously, and then bringing to volume.
- Centrifuge as above (step 8).
- Wash the pellet a second time as in step 10, to remove any trace of growth medium and released ammonium.
- Resuspend the pellet in 5 ml resuspension buffer.
- Pipette 200 µl of the culture in a well of the microplate and measure OD600 using the spectrofluorometer (mode: absorbance). As blank, use 200 µl of resuspension buffer in another well.
- Adjust to OD600 = 0.5 by diluting with resuspension buffer in the 50 ml tube.
- Mix well and aliquot in the wells, 200 µl each.
- An excitation and emission spectrum should be recorded first to confirm the fluorescence properties of the cpGFP as part of the sensors (parameters: mode: fluorescence; read: bottom reading; excitation: Fix emission to 530 and record spectra from 350 - 510 nm; emission: fix the excitation to 480 nm and record the spectra from 500 - 600 nm; step size 5 nm: bandwidth: 7.5/7.5 nm; gain:100), see Figure 2A. With knowledge of the major excitation maximum the reader can decide on the excitation wavelength for the single point measurements.
- Read basal fluorescence. It should be the same for all wells (parameters: mode: fluorescence; read: bottom reading; excitation: 488 nm; emission: 513 nm; bandwidth: 7.5/7.5 nm; gain:100; shake: orbital, medium, 3 sec, initial and between readings).
- Since response is saturated above 1 mM for all variants, it can be useful to make a calibration curve with ammonium chloride ranging from 0 to 1 mM final concentration in the wells (0, 1 µM, 10 µM, 100 µM, 1 mM), see Figure 1B. It is useful to include negative controls, e.g. sodium chloride, to exclude artifacts and to include other salts to check for specificity. In that case, use the highest concentration (e.g. 1 or 10 mM), see Figure 2C.
- Add the treatment solutions (as 5x concentrated stocks, dissolved in water) to the compartments of the reservoir (Figure 3).
- Add 50 µl of each treatment solution or water control to each wells with a multichannel pipette, pipetting up and down for about 5 times for mixing the treatment with the cells. Set up at least three replicates per treatment.
- Response is immediate, within the time limits of the microplate reader. With the parameters as above, it takes about 1 min to read a full plate.
- Repeat reading as step 18 after 5 min from the treatment. We usually consider these latter responses, as they tend to be a bit stronger.
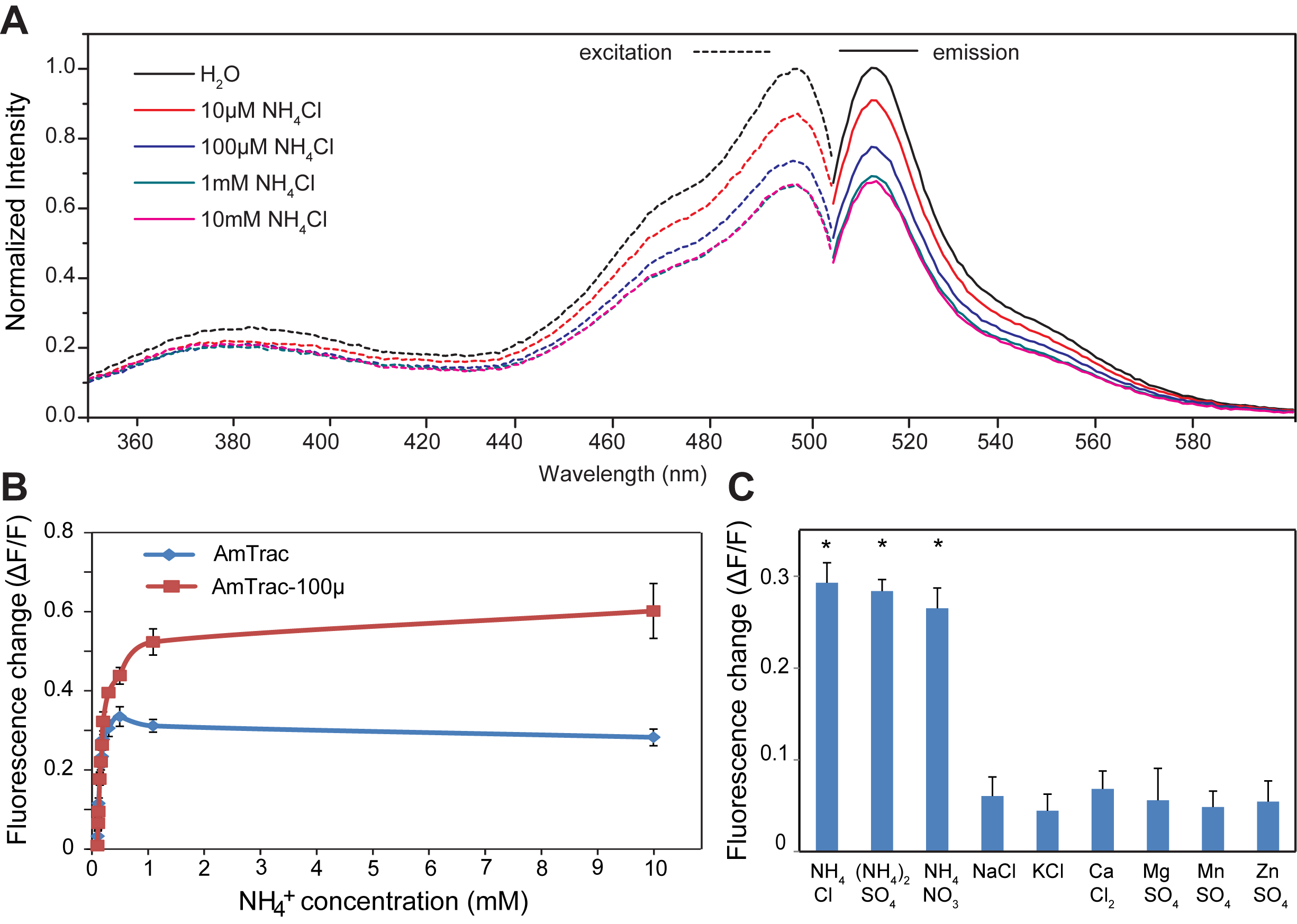
Figure 2. A. Excitation and emission spectra of AmTrac-GS treated with indicated ammonium chloride concentration. Data are normalized to the initial value. B. Response of AmTrac and AmTrac-100 to increasing concentrations of ammonium chloride. C. Specificity of AmTrac, the concentration of the indicated salts was 1 mM (De Michele et al., 2013).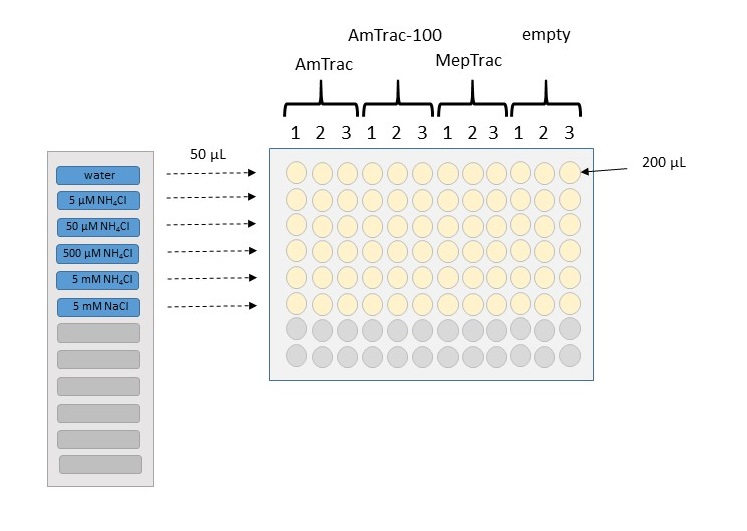
Figure 3. Schematic representation of the treatment procedure for building a calibration curve. Wells in a microplate are filled with 200 µl of yeast culture, resuspended in 50 mM MES buffer with 5% glycerol. Samples can be arranged in columns, with triplicates for each construct. Fluorescence from yeast transformed with the empty vector will be used as background value. After reading basal fluorescence, 50 µl of 5x stock treatment will be added to the wells, by using a multichannel pipette. In this example, we use a solvent control (water), four concentrations of increasing magnitude of NH4Cl, and a chloride control at the highest concentration.
Data analysis
- Subtract background fluorescence (from a yeast transformed with an empty vector) to all fluorescence values (spectra as well as single point measurements).
- AmTracs respond to ammonium by decreasing cpGFP fluorescence emission. For this reason, fluorescence of the water control shows always the highest value. Calculate ΔF, by subtracting the response to treatment to that of the water control (0 ammonium). Divide ΔF to the response to treatment (ΔF/F). By using ratios instead of absolute values, it is possible to compare different experiments, and in theory even using different dilutions (not just to OD600 = 0.5). However, it is good practice to maintain conditions as similar as possible.
- Calculate averages and errors from all replicates, for each treatment.
- Plot as histogram or line. Figure 2B shows the response of AmTrac and AmTrac-100 to increasing concentrations of ammonium chloride.
The reader can then use the calibration curve to quantify unknown ammonium concentration in a solution.
Recipes
- 100x proline solution
10 g L-Proline dissolved in 100 ml MilliQ water
pH ~ 5.5 - 50x arginine solution
50 mM L-Arginine monohydrochloride dissolved in MilliQ water
pH ~ 5.5 - SD medium
1.7 g/L yeast nitrogen base w/o amino acids w/o ammonium sulfate
30 g/L glucose
Autoclave, 120 °C, 30 min
When hand-warm, in a sterile hood, add proline from a 100x stock to a final concentration of 0.1%
The pH of the SD medium is ~ 4.2 and changes minimally upon the addition of proline to pH ~ 4.3
For solid medium, add 15 g/L agar before autoclaving
Add 1 mM arginine (final concentration) when medium is hand-warm before pouring plates. In case the SD medium does not solidify well, which may depend on the autoclave used, it is recommended to adjust the pH of the SD medium to pH 5.8 with NaOH before addition of agar and autoclaving. - Washing buffer
50 mM MES, bring to pH 6.0 with NaOH pellets - Resuspension buffer
50 mM MES, bring to pH 6.0 with NaOH pellets
Add 5% glycerol, to delay sedimentation of the cells
Acknowledgments
This work has been supported by grant MCB-1021677 by the National Science Foundation (Wolf B Frommer). The protocol has been adapted from De Michele et al. (2013).
References
- Boeckstaens, M., Andre, B. and Marini, A. M. (2008). Distinct transport mechanisms in yeast ammonium transport/sensor proteins of the Mep/Amt/Rh family and impact on filamentation. J Biol Chem 283(31): 21362-21370.
- De Michele, R., Ast, C., Loque, D., Ho, C. H., Andrade, S. L., Lanquar, V., Grossmann, G., Gehne, S., Kumke, M. U. and Frommer, W. B. (2013). Fluorescent sensors reporting the activity of ammonium transceptors in live cells. Elife 2: e00800.
- Gietz, D., St Jean, A., Woods, R. A. and Schiestl, R. H. (1992). Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res 20(6): 1425.
- Ho, C. H. and Frommer, W. B. (2014). Fluorescent sensors for activity and regulation of the nitrate transceptor CHL1/NRT1.1 and oligopeptide transporters. Elife 3: e01917.
- Marini, A. M., Soussi-Boudekou, S., Vissers, S. and Andre, B. (1997). A family of ammonium transporters in Saccharomyces cerevisiae. Mol Cell Biol 17(8): 4282-4293.
Article Information
Copyright
Ast et al. This article is distributed under the terms of the Creative Commons Attribution License (CC BY 4.0).
How to cite
Readers should cite both the Bio-protocol article and the original research article where this protocol was used:
- Ast, C., Frommer, W. B., Grossmann, G. and De Michele, R. (2015). Quantification of Extracellular Ammonium Concentrations and Transporter Activity in Yeast Using AmTrac Fluorescent Sensors. Bio-protocol 5(1): e1372. DOI: 10.21769/BioProtoc.1372.
- De Michele, R., Ast, C., Loque, D., Ho, C. H., Andrade, S. L., Lanquar, V., Grossmann, G., Gehne, S., Kumke, M. U. and Frommer, W. B. (2013). Fluorescent sensors reporting the activity of ammonium transceptors in live cells. Elife 2: e00800.
Category
Microbiology > Microbial metabolism > Nutrient transport
Cell Biology > Cell imaging > Live-cell imaging
Cell Biology > Cell imaging > Fluorescence
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link








