- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Enzymatic Reactions and Detection of C3 Cleavage Fragments
Published: Vol 4, Iss 16, Aug 20, 2014 DOI: 10.21769/BioProtoc.1205 Views: 11618
Reviewed by: Pia GiovannelliAnonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols

Measurement of CD74 N-terminal Fragment Accumulation in Cells Treated with SPPL2a Inhibitor
Rubén Martínez-Barricarte [...] Jean-Laurent Casanova
Jun 5, 2019 6981 Views
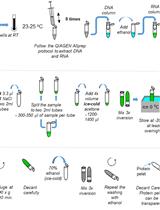
Capillary Nano-immunoassay for Quantification of Proteins from CD138-purified Myeloma Cells
Irena Misiewicz-Krzeminska [...] Norma C. Gutiérrez
Jun 20, 2019 6855 Views
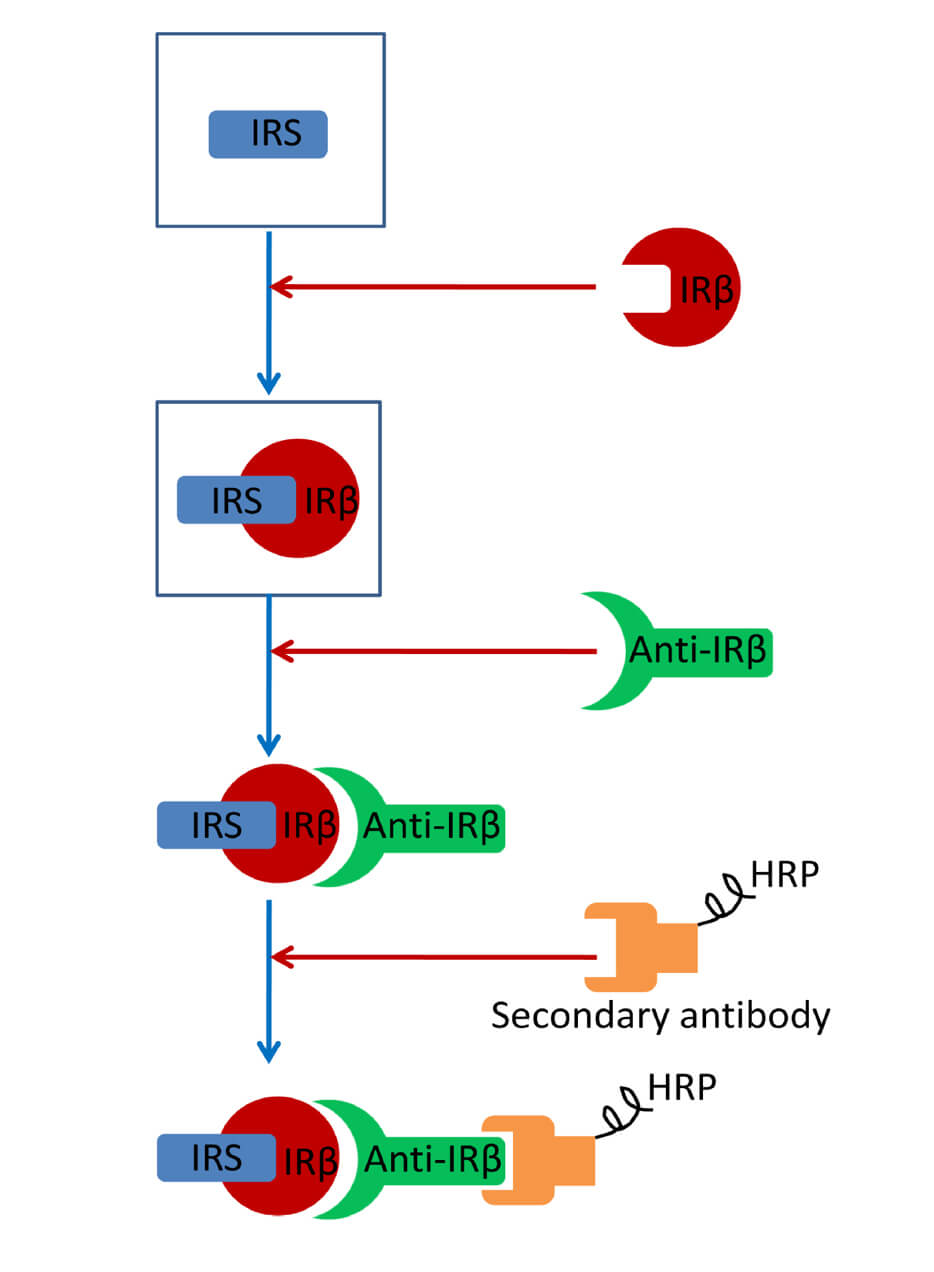
Far-western Blotting Detection of the Binding of Insulin Receptor Substrate to the Insulin Receptor
Jinghua Peng [...] Ling He
Feb 20, 2023 2492 Views
Abstract
The complement component C3 is the major effector molecule of the complement system. C3 circulates in the blood and interstitial fluids as pro-enzyme and is activated by enzymatic cleavage into a C3a portion, a classic anaphylatoxin that functions as chemoattractant and immune cell activator, and the C3b portion, the body’s most potent opsonin. C3 cleavage is in most cases mediated by an enzyme complex called the C3 convertase. However, it is now becoming increasingly clear that the cleavage of C3 by a range of ‘single’ proteases into bioactive C3a and C3b fragments is of high physiological significance. Here, we describe a protocol for the enzymatic cleavage of human C3 by the serine protease cathepsin L and the detection of the cleavage products C3a and C3b by western blotting as an example for this kind of enzymatic reactions.
Keywords: Complement systemMaterials and Reagents
- Purified human C3 (Complement Technology, catalog number: A113 )
- Purified human C3b (Complement Technology, catalog number: A114 )
- Purified human C3a (Complement Technology, catalog number: A118 )
- Recombinant human cathepsin L (CTSL) (R&D Systems, catalog number: 952-CY-010 )
- 1 M Dithiothreitol (DTT) (Life Technologies, catalog number: P2325 )
- Tris acetate gels 3-8% or equivalent (Life Technologies, NuPAGE® Novex®, catalog number: WG1603BOX )
- 20x NuPAGE Tris acetate running buffer (Life Technologies, catalog number: LA0041 )
- Protein size standard (Life Technologies, catalog number: LC5800 )
- 5x protein sample reducing loading buffer (Thermo Fisher Scientific, catalog number: 39000 )
- Nitrocellulose membranes (Life Technologies, catalog number: IB301002 )
- 5% dried milk powder in 1x PBS (Marvel)
- Tween 20 (Sigma-Aldrich, catalog number: P5927 )
- Rabbit anti-C3d antibody recognizing the intact and cleaved C3 α-chain (Abcam, catalog number: ab17453 )
- Rabbit anti-C3 antibody recognizing the C3 β-chain (MyBioSource, catalog number: MBS857324 )
- Mouse anti-C3 antibody recognizing the C3a portion within the uncleaved C3 α-chain (Abcam, catalog number: ab36385 )
- Mouse anti-C3a neoepitope antibody recognizing only cleaved C3a (Abcam, catalog number: ab11873 )
- Secondary antibodies to rabbit and mouse Ig conjugated to Horseradish peroxidase (HRP) (GE Healthcare, catalog numbers: RPN4301 and NA9310 , respectively)
- ClarityTM Western ECL substrate (Bio-Rad Laboratories, catalog number: 170-5060 )
- NaCl (Sigma-Aldrich, catalog number S7653 )
- KCl (Sigma-Aldrich, catalog number P9333 )
- Na2HPO4 (Sigma-Aldrich, catalog number S7907 )
- KH2PO4 (Sigma-Aldrich, catalog number P5655 )
- MES (free acid) (Sigma-Aldrich, catalogue number M0164 )
- Brij35 (Thermo Fisher Scientific, Pierce, catalog number: 28316 )
- NaOH (Sigma-Aldrich, catalog number S8045 )
- Tris base (Sigma-Aldrich, catalog number: T1503 )
- HCl (Sigma-Aldrich, catalog number: 258148 )
- 10x phosphate buffered saline (PBS) (see Recipes)
- 0.5 M 2- (N-morpholino) ethanesulfonic acid (MES) buffer (see Recipes)
- 0.05 M Tris (hydroxymethyl) aminomethan (THAM) buffer (see Recipes)
Equipment
- Centrifuge (Eppendorf, model: 5427 R )
- 37 °C, 5% CO2 cell culture incubator
- Heating block (Eppendorf, catalog number: 5382000031 )
- Reaction tubes (1.5 ml) (Eppendorf, catalog number: 0030125177 )
- Pipettes of several volume sizes (for example, Eppendorf)
- Power pack for gel electrophoresis (Bio-Rad Laboratories, catalog number: 164-5070 )
- XCell4 SureLockTM Midi-Cell gel running tank (Bio-Rad Laboratories, catalog number: WR0100 )
- Western blotting transfer equipment (Life Technologies, iBlot®, catalog number: IB21001 )
- Western blot visualization machine Chemi Doc MP imaging system (Bio-Rad Laboratories, catalog number: 170-8280 )
- Platform rocker (Stuart, catalog number: R11876-01 )
- Cold room
- Ice bucket
Software
- ImageLab software 4.1 (Bio-Rad Laboratories)
Procedure
- Enzymatic cleavage of C3 by cathepsin L (CTSL)
All reagents are kept on ice at all times unless otherwise stated.- C3 preparation: Human serum-purified C3 is diluted from stock (1 mg/ml) in 1x PBS to a final concentration of 6 μg/ml in 1.5 ml Eppendorf tubes. For example, 3 μl of stock C3 (1 mg/ml) is diluted it in 500 μl 1x PBS to obtain the 6 μg/ml final concentration (see Note 1). This amount of diluted C3 stock is sufficient for 10 enzymatic CTSL cleavage reactions.
- Activated cathepsin L (CTSL) preparation (see Note 1): For the activation of CTSL in MES/Brij 35 DTT buffer, DTT (1 M stock) is first diluted 1/200 in 50 mM MES buffer to 5 mM final concentration to generate the activation buffer. MES buffer provides pH of about 6.15 that is optimal for CTSL, while DTT and Brij 35 activate the enzyme. CTSL (stock 200 ng/µl) is then diluted to a final concentration of 40 ng/µl (1: 5) in the activation buffer in 1.5 ml Eppendorf tubes and incubated 15 min on ice without shaking to activate the enzyme (see Note 2). Calculate in advance how many reactions are required for an experiment (see Note 3) and activate the appropriate amount of CTSL.
- For a reaction with a ratio of about 1: 1 of C3: CSTL, 50 μl of diluted C3 stock (see step A1) are mixed with 38 μl 1x PBS and 12 μl of activated CTSL (see step A2).
- At this step it is recommended to also prepare appropriate control ‘reactions’:
- 50 µl of C3 diluted stock + 50 µl 1x PBS into an Eppendorf tube, and
- A reaction mixture of 12 μl MES buffer/DTT without CTSL + 38 µl of 1x PBS + 50 μl of C3 diluted stock as control reaction to assess the effect of the CTSL activation buffer without the enzyme on C3.
- In addition, similar diluted stocks of 250 ng/100 µl in 1x PBS of C3a and 250 ng/100 µl 1x PBS C3b are prepared and will be used as ‘cleavage controls’.
- 50 µl of C3 diluted stock + 50 µl 1x PBS into an Eppendorf tube, and
- All reaction mixtures (including controls) are incubated for 30 min (yields complete C3 cleavage) or desired times points at 37 °C in a heating block (alternatively, in a water bath).
- Samples are centrifuged at 400 x g for 1 min at room temperature.
- Twenty-five microliter of 5x protein sample reducing buffer is added to all samples and the samples then boiled at 95 °C for 3-4 min in preparation for subsequent Western blotting analysis (see section B).
- Samples are centrifuged at 400 x g for 1 min at room temperature.
- Samples can also be stored at this point for later analysis at -80 °C.
- C3 preparation: Human serum-purified C3 is diluted from stock (1 mg/ml) in 1x PBS to a final concentration of 6 μg/ml in 1.5 ml Eppendorf tubes. For example, 3 μl of stock C3 (1 mg/ml) is diluted it in 500 μl 1x PBS to obtain the 6 μg/ml final concentration (see Note 1). This amount of diluted C3 stock is sufficient for 10 enzymatic CTSL cleavage reactions.
- Detection of C3 enzymatic cleavage products by Western blotting
- Twenty five microliter of the reaction samples from step A8 [(C3 + CTSL) and controls (appropriately diluted stocks of C3a, C3b, C3 (all three control proteins are first diluted to 6 µg/ml in 1x PBS and then further diluted 1: 1 in 1x PBS) as well as C3 + MES buffer/DTT as described under step A4b)] are loaded along with the protein size marker (10 µl) onto a 3-8% Tris acetated gradient polyacrylamide gel (see Note 4) that had been assembled with 1x NuPAGE Tris acetate running buffer into the Western blotting chamber according to the manufacturer’s protocol (see Note 5).
- The loaded protein samples are separated in the gel at 150 V for 60 min or until the front of the dye in the protein sample buffer reaches the bottom end of the gel.
- The separated proteins are then transferred onto a nitrocellulose membrane using dry transfer in the Life Technologies iBlot transfer system according to the manufacturer’s protocol. This particular machine does not allow for custom settings and we use the 20 V and 5’ 30’’ transfer conditions (programme 3 on the iBlot machine). If you use another transfer devise, please follow the manufacturer’s instructions.
- The nitrocellulose membrane is then placed into a container with enough 5% milk in 1x PBS supplemented with 0.1% Tween 20 to cover the membrane safely and blocked for 1-2 h at room temperature on a rocking shaker (low rotation).
- Primary rabbit anti-C3 antibody and rabbit anti-C3d are diluted together at a concentration of 1 μg/ml each in 5% dry milk powder in 1x PBS with 0.1% Tween 20 and incubated with the membrane overnight at 4 °C on a shaker (see Note 6). Ensure that pH remains 7.4 after addition of the milk powder, otherwise adjust accordingly.
- The membrane is washed 3 times using 50 ml 1x PBS with 0.1% Tween 20 for 15 min each at room temperature on a shaker.
- Secondary anti-rabbit-HRP antibody is diluted in 5% dry milk powder in 1x PBS with 0.1% Tween 20 at 1:2,000 dilution (in enough volume to cover the membrane during shaking) and incubated with the membrane for 2 h at room temperature on a shaker (slow rotation).
- The membrane is washed again 3 times using 50 ml 1x PBS with 0.1% Tween 20 for 15 min each at room temperature on a shaker.
- The Western blot is developed using the ClarityTM Western ECL Substrate according to the manufacturer’s protocol. Briefly, the two substrate components in the ECL Kit are mixed at a ratio 1:1 and the membrane incubated in the mix for 5 min.
- Excess ECL detection fluid is removed from the membrane and the protein bands are then visualized using the Chemi Doc MP imaging system and ImageLab software (see Note 7).
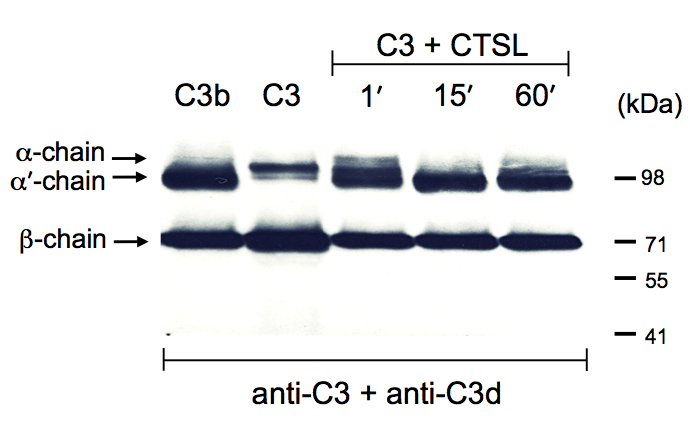
Figure 1. Cathepsin L cleaves C3 to generate C3b. Serum-purified C3 was incubated with activated Cathepsin L (CTSL) as per protocol for the indicated time points and the reactions terminated by addition of the protein reducing loading buffer. The reaction mixtures where then analysed by Western blotting for cleavage of the C3 α-chain with a mixture of two anti-C3 antibodies recognizing the non-processed α-chain of C3 and also the processed α’-chain of generated C3b (anti-C3d) and the β-chain in both C3 forms (anti-C3). Serum-purified C3 and C3b were loaded as control proteins. Data shown are representative of three (n=3) independently performed experiments.
- Twenty five microliter of the reaction samples from step A8 [(C3 + CTSL) and controls (appropriately diluted stocks of C3a, C3b, C3 (all three control proteins are first diluted to 6 µg/ml in 1x PBS and then further diluted 1: 1 in 1x PBS) as well as C3 + MES buffer/DTT as described under step A4b)] are loaded along with the protein size marker (10 µl) onto a 3-8% Tris acetated gradient polyacrylamide gel (see Note 4) that had been assembled with 1x NuPAGE Tris acetate running buffer into the Western blotting chamber according to the manufacturer’s protocol (see Note 5).
Notes
- Described here is the enzymatic cleavage of C3 by CTSL. If the C3 cleavage products of C3 by an alternative protease by a C3 convertase is to be assessed, please refer to the specific information in regards to activation buffers and suggested enzyme: substrate ratios for those reagents when setting up the experiment. Another note on the buffers used: MES is a buffer which has strongest buffering effect at pH 6.15 where cathepsin L is active. In order to preserve the integrity of papain-like cysteine proteases and to prevent autocatalytic degradation, it has become a general practice to reversibly inactivate stock solutions of cathepsins with methylmethanethiolsulfonate (MMTS) before storage. As a result, addition of a reducing agent such as DTT or an equivalent thiol is required to regenerate the active enzyme prior to use by cleaving the inhibiting S-S bond and freeing the catalytic centre of the enzyme for in vitro activation. Brij 35 concenration at 0.005% does increase cathepsin L activity after 10 min activation at least 20%.
- The efficiency of the CTSL activation reaction is variable among different enzyme preparations and vendors, but commonly only about 20%. We determined previously that a ratio of 1:1 - 1:2 of total C3: total CTSL results consistently in 100% C3 cleavage by CTSL within 30 min at 37 °C (Liszewski et al., 2013). These conditions have therefore been chosen in this protocol.
- Additional sample preparations may be required if, for example, the inhibition of CTSL-mediated C3 cleavage with inhibitory reagents including chemical inhibitors and blocking antibodies needs to be assessed. Please refer to Liszewski et al. (2013) for the nature and source of such reagents.
- It is important to use ‘low percentage’ polyacrylamide gels when the processing of the C3 α-chain is monitored as the molecular weight difference between the α-chain of C3 and the α’-chain of C3b (where the C3a portion is cleaved) is only about 8 kDa. Conversely, when the generation of the cleaved C3a is to be monitored (see Note 5), the usage of higher percentage gels (>12%) or gradient gels (4-20%) is required because of the low molecular weight of 8 kDa of C3a. For more details about size and gel appearance of C3 cleavage products please refer to Nilsson et al. (2011).
- This protocol uses the specific Life Technologies/Bio-RAd Midi gel Chamber, the iBlot dry transfer system and the Chemi Doc MP imaging system for different steps in the protocol. Of course, any other system (including semi-dry and wet transfer systems) can be used for the separation and visualization of enzymatic C3 cleavage by Western blotting.
- In this protocol (and in Figure 1), the detection of the processing of the α-chain of C3 into the α’-chain of C3b by CTSL is described and shown. For the detection of the cleaved C3a product, the usage of a ‘higher percentage’ or gradient gel is advised (see Note 3) and the antibodies in Step B5 are replaced with a mixture of primary mouse anti-C3 antibody recognizing the C3a portion within the uncleaved C3 α-chain (1:500) and mouse anti-C3a neoepitope antibody recognizing exclusively cleaved C3a (1:1,000).
- Detection of both, the CTSL-generated C3b and C3a at the same time can be achieved by using a silver staining protocol after gel separation of the C3 cleavage products in place of the Western blotting protocol (Liszewski et al., 2013). This has the advantage that all C3 cleavage fragments generated during the enzymatic reaction will be detected and not only those recognized by the specific antibodies chosen. However, silver staining is only suitable if purified reagents are used in in vitro rections as decribed here. To detect in vivo/ex vivo generated C3 cleavage by CTSL, for example in cell lysate preparations, silver staining is unsuitable.
Recipes
- 10x PBS
80 g of NaCl
20 g of KCl
14.4 g of Na2HPO4
2.4 g KH2PO4
Dissolved in 800 ml dH2O and adjust pH to 7.4 with pure HCl under stirring
Top up with dH2O to 1 L
- 0.5 M MES buffer
Add 97.6 g MES to 800 ml dH2O and dissolve
Adjust pH to 6.0 with 10 N NaOH and then top up to 1 L with dH2O
Add 0.5 ml of 10% Brij 35 solution to bring it to 0.005% final concentration
- 0.05 M Tris (pH 7.5) buffer (100 ml)
Mix 0.6 g of Tris base with 80 ml deionized (d) H2O
Adjust pH to 7.5 with HCl under stirring
Add dH2O to 100 ml
Stored at 4 °C
Acknowledgments
This protocol was developed by Claudia Kemper and validated by Martin Kolev. The protocol was originally published in Liszewski et al. (2013). This work was supported by the MRC Research Grant G1002165 (C.K.) the MRC Centre for Transplantation Grant MR/J006742/1, an EU-funded Innovative Medicines Initiative BTCURE (C.K.), and a Wellcome Trust Investigator Award (C.K).
References
- Liszewski, M. K., Kolev, M., Le Friec, G., Leung, M., Bertram, P. G., Fara, A. F., Subias, M., Pickering, M. C., Drouet, C., Meri, S., Arstila, T. P., Pekkarinen, P. T., Ma, M., Cope, A., Reinheckel, T., Rodriguez de Cordoba, S., Afzali, B., Atkinson, J. P. and Kemper, C. (2013). Intracellular complement activation sustains T cell homeostasis and mediates effector differentiation. Immunity 39(6): 1143-1157.
- Nilsson, U. R., Funke, L., Nilsson, B. and Ekdahl, K. N. (2011). Two conformational forms of target-bound iC3b that distinctively bind complement receptors 1 and 2 and two specific monoclonal antibodies. Ups J Med Sci 116(1): 26-33.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Kemper, C. and Kolev, M. (2014). Enzymatic Reactions and Detection of C3 Cleavage Fragments. Bio-protocol 4(16): e1205. DOI: 10.21769/BioProtoc.1205.
Category
Immunology > Host defense > General
Biochemistry > Protein > Activity
Biochemistry > Protein > Immunodetection > Western blot
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










