- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
Bacterial Fluorescent-dextran Diffusion Assay
Published: Vol 4, Iss 14, Jul 20, 2014 DOI: 10.21769/BioProtoc.1191 Views: 9741
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
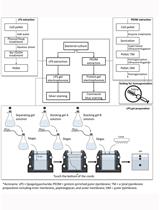
Extraction and Electrophoretic Analysis of Bacterial Lipopolysaccharides and Outer Membrane Proteins
Yue-Jia Lee and Thomas J. Inzana
Dec 20, 2021 5761 Views

Shipment of Cyanobacteria by Agarose Gel Embedding (SCAGE)—A Novel Method for Simple and Robust Delivery of Cyanobacteria
Phillipp Fink [...] Karl Forchhammer
Dec 5, 2024 1399 Views
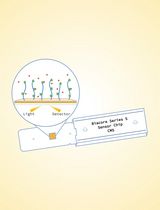
Surface Plasmon Resonance for the Interaction of Capsular Polysaccharide (CPS) With KpACE
Zhe Wang [...] Chao Cai
Jun 20, 2025 3554 Views
Abstract
Antimicrobial peptides are known to disrupt bacterial membranes allowing solutes to flow across the membrane in an unregulated manner resulting in death of the organism. Disrupting the bacterial membrane would thus perturb the cells osmotic balance resulting in an initial influx of the external aqueous buffer. We have designed an assay to investigate how antimicrobial peptide concentration affects the ability of fluorescently labelled dextran moieties of differing molecular weight and hydrodynamic radii to cross membranes of viable bacteria. This assay was used to show that diffusion of low and high molecular weight dextrans into bacteria was a function of antimicrobial peptide concentration (Sani et al., 2013).
Materials and Reagents
- Bacteria of interest [e.g. Staphylococcus aureus (S. aureus)] (Sani et al., 2013)
- Rhodamine-dextran (RD) of different molecular weights [e.g. 40 kDa RD molecular weight (RD-40) (Sigma-Aldrich, catalog number: 42874 )]
- Fluorescein-dextran (FD) of different molecular weights [e.g. 4.4 kDa FD (FD-4) (Sigma-Aldrich, catalog number: 46944 )]
- LIVE/DEAD® BacLightTM Bacterial Viability and Counting Kit (Life Technologies, catalog number: L34856 )
- Dulbecco’s Phosphate Buffered Saline (Dulbecco’s PBS) (Sigma-Aldrich, catalog number: D8537 )
- Peptide of interest (e.g. Maculatin 1.1) (Sani et al., 2013)
- Bacterial growth media [i.e., particular for the bacteria of interest e.g. S. aureus was grown as a batch culture in Luria broth (Thermo Fisher Scientific) (see Recipes)] (Sani et al., 2013)
Equipment
- Flow cytometer (e.g. Beckman Coulter, model: Quanta SC-MPL)
- 96 well plates (flat bottomed) (Thermo Fisher Scientific, Nunc®, catalog number: 12565210 )
- Transfer pipette
Procedure
- Stock preparation
- Prepare 3.2 mM solution of FD4 in Dulbecco’s PBS (12.8 mg FD4/ml) and 3.2 mM solution of RD-40 in Dulbecco’s PBS (128 mg RD 40/ml). Note that Dulbecco’s PBS is used as it has been found not to affect the membranes of a wide range of bacteria in this assay. 3.2 mM solutions have been optimized for this assay using S. aureus and may need slight optimization for other bacteria.
- Prepare 2.5 x 106 cells/ml solution of heat-killed (55 °C for 1.5 h) bacteria of interest in the bacterial growth media as a compensation control for non-specific FD4 and RD40 binding to the bacteria.
Note: Bacteria were counted, throughout, using the Quanta SC-MPL and the LIVE/DEAD® BacLightTM Bacterial Viability and Counting Kit. But other methods of bacterial counting using Petroff Hausser counting chamber or optical density estimation from growth curves and CFUs have been used.
- Prepare a 2.5 x 106 cells/ml solution of viable bacteria of interest in chilled bacterial growth media.
Note: harvesting and processing of the bacteria must be completed at 4 °C using media at 4 °C and the bacteria then stored for the assay at 4 °C.
- Prepare a stock solution of the peptide of interest (e.g. 1 mg/ml) in the bacterial growth media.
Note: Peptide may need to be initially solvated in DMSO to prevent precipitation in the bacterial growth media. The final DMSO concentration in the peptide stock solution needs to be ≤ 10% v/v, this concentration will equate to ≤ 5% v/v of DMSO in the initial assay dilution which does not affect the assay.
- Bacterial media conditioned 96 well plates; add 250 μl of the bacteria media/well and allow to sit for 16 h, 4 °C. Remove the media (aspiration) immediately prior to the assay.
- Prepare 3.2 mM solution of FD4 in Dulbecco’s PBS (12.8 mg FD4/ml) and 3.2 mM solution of RD-40 in Dulbecco’s PBS (128 mg RD 40/ml). Note that Dulbecco’s PBS is used as it has been found not to affect the membranes of a wide range of bacteria in this assay. 3.2 mM solutions have been optimized for this assay using S. aureus and may need slight optimization for other bacteria.
- Assay protocol
Critical point: Many bacteria can actively transport dextran, thus to inhibit active transport of dextran the assay must be completed at 4 °C or on ice. All solutions after being prepared must be chilled to 4 °C and kept at 4 °C or on ice. The viable bacteria need 30-40 min at 4 °C to inhibit active transport; this can easily be achieved if the harvested bacteria are centrifuged at 4 °C and pre-chilled media is used to resuspend the bacteria at 2.5 x 106 cells/ml which are then kept at 4 °C or on ice.- Prepare serial dilutions of the peptide in the media conditioned 96 well plates, e.g. 200 μl of the peptide stock solution is added to the first well and 100 μl is transferred to the second well containing 100 μl Dulbecco’s PBS and mixed (10-15 times, pipetting up/down). This transfer process in repeated from the second to the third well, etc., to form a double serial dilution.
Note: Gentle pipetting is required to avoid bubble formation and using the same tip in one serial dilution (high concentration to low) avoids peptide adherence to the tip, if tips are changed per dilution step.
- Add 10 μl of the 3.2 mM stock solution of FD4 and 10 μl of the 3.2 mM stock solution of RD40 to each peptide dilution. Or add 20 μl of each if using only one dextran.
- Add 80 μl of the 2.5 x 106 cells/ml of the bacteria of interest; gently mix with the transfer pipette and incubate at 4 °C for 60-90 min.
Note: Options at this point would be to have a series of the incubation times to look at the rate of dextran dye diffusion.
- Prepare non-specific dextran binding control: to three wells containing 100 μl Dulbecco’s PBS, 10 μl of the 3.2 mM stock solution of FD4 and 10 μl of the 3.2 mM stock solution of RD40, add 80 μl of the 2.5 x 106 cells/ml of the heat killed bacteria of interest and incubate at 4 °C for the equivalent time as in step B3 above.
- Prepare active transport/non-specific dextran binding control: To three wells containing 100 μl Dulbecco’s PBS, 10 μl of the 3.2 mM stock solution of FD4 and 10 μl of the 3.2 mM stock solution of RD40 add 80 μl of the 2.5 x 106 cells/ml of the viable bacteria of interest and incubate at 4 °C for the equivalent time as in step B3 above.
Note: These controls must have no peptide in the wells.
- Set up flow cytometer:
- Quanta SC-MPL flow cytometer was used and green fluorescence of FD-4 positive cells (525 ± 20 nm band-pass filter, typically FL1 channel) and the red fluorescence of the RD-40 positive cells (575 ± 15 nm band-pass filter, typically FL3 channel) were measured on a logarithmic scale and analysed simultaneously with the electrical volume (EV) and side scatter (SS) using a 488 nm laser.
- Other flow cytometers could be used which use forward and side scatter to discriminate bacterial cells in logarithmic scale mode.
- Fluorescent analysis gates (FD4, FD4/RD40 and RD40) were set to contain 1% of background fluorescence (defined as the non-specific adherence of the dextran molecules to heat-killed S. aureus cells and/or S. aureus cells harvested and maintained at 4 °C).
- Analysis of dye diffusion. Each test sample was kept at 4 °C or on ice for the analysis and was completed over 1 min, which typically captured 14,000 ± 2,000 counts/sample.
- Positive FD4 and RD40 bacteria were determined as being above background fluorescence and FD4-positive, FD4/RD40-dual positive and RD40-positive bacteria were plotted against peptide concentration.
- Prepare serial dilutions of the peptide in the media conditioned 96 well plates, e.g. 200 μl of the peptide stock solution is added to the first well and 100 μl is transferred to the second well containing 100 μl Dulbecco’s PBS and mixed (10-15 times, pipetting up/down). This transfer process in repeated from the second to the third well, etc., to form a double serial dilution.
Recipes
- Luria broth
10 g of tryptone/L
5 g of yeast extract/L
10 g NaCl
pH 7.5
Acknowledgments
None.
References
- Sani, M. A., Whitwell, T. C., Gehman, J. D., Robins-Browne, R. M., Pantarat, N., Attard, T. J., Reynolds, E. C., O'Brien-Simpson, N. M. and Separovic, F. (2013). Maculatin 1.1 disrupts Staphylococcus aureus lipid membranes via a pore mechanism. Antimicrob Agents Chemother 57(8): 3593-3600.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
O’Brien-Simpson, N. M., Pantarat, N., Walsh, K. A., Reynolds, E. C., Sani, M. and Separovic, F. (2014). Bacterial Fluorescent-dextran Diffusion Assay. Bio-protocol 4(14): e1191. DOI: 10.21769/BioProtoc.1191.
Category
Microbiology > Antimicrobial assay > Peptide assay
Microbiology > Microbial cell biology > Cell viability
Biochemistry > Carbohydrate > Polysaccharide
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










