- Submit a Protocol
- Receive Our Alerts
- Log in
- /
- Sign up
- My Bio Page
- Edit My Profile
- Change Password
- Log Out
- EN
- EN - English
- CN - 中文
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
- EN - English
- CN - 中文
- Home
- Protocols
- Articles and Issues
- For Authors
- About
- Become a Reviewer
The BhbA Enzyme Assay
Published: Vol 4, Iss 8, Apr 20, 2014 DOI: 10.21769/BioProtoc.1111 Views: 8830
Reviewed by: Anonymous reviewer(s)

Protocol Collections
Comprehensive collections of detailed, peer-reviewed protocols focusing on specific topics
Related protocols
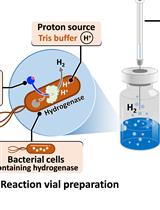
H2 Production from Methyl Viologen–Dependent Hydrogenase Activity Monitored by Gas Chromatography
Nuttavut Kosem
Dec 5, 2023 1769 Views
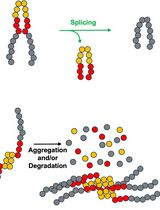
Monitoring Protein Stability In Vivo Using an Intein-Based Biosensor
John S. Smetana [...] Christopher W. Lennon
Apr 20, 2025 1584 Views
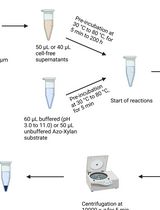
Endo-1,4-β-D-xylanase Assay Using Azo-Xylan and Variants Thereof
Luca Bombardi [...] Salvatore Fusco
Apr 20, 2025 1927 Views
Abstract
Reductive dehalogenation has been found primarily in anaerobic communities and is originally thought to rarely occur in aerobes. A reductive dehalogenase (BhbA) was characterized from an aerobic strain of Comamonas sp. 7D-2, which was isolated from a bromoxynil octanoate-contaminated soil sample collected in Jiangsu, China. BhbA catalyzes the reductive dehalogenation of bromoxynil and its derivative 3,5-dibromo-4-hydroxybenzoate under aerobic conditions. BhbA is membrane-associated and found to have the key features of anaerobic respiratory reductive dehalogenases. This protocol describes the method for enzyme analysis of the aerobic reductive dehalogenase (BhbA) in the membrane fraction.
Materials and Reagents
- Comamonas sp. 7D-2
- LB medium
- Substrate, 3,5-dibromo-4-hydroxybenzoate (DBHB) or 3-bromo-4-hydroxybenzoate (BHB) (Sigma-Aldrich)
- Electron donor, NADPH or NADH (Sangon Biotech, catalog numbers: Y4433000-100 mg and NB0642-1 g , respectively)
- Reaction inhibitor, sodium dithionite (Na2S2O4) (Sigma-Aldrich, catalog number: 7775-14-6 )
- Protein Quantification Kit (Sangon Biotech, catalog number: BE530-100 ml )
- Phosphate buffered saline (PBS) (Sambrook and Russell, 2001) (see Recipes)
- Mobile phase of HPLC (see Recipes)
Equipment
- 7 ml centrifuge tube
- Membrane filtration (pore size, 0.22 μm)
- Centrifuge
- HPLC (600 controller, Rheodyne 7725i manual injector and 2487 Dual λ Absorbance Detector) (Waters)
- Ultrasonic instrument
- Fast protein liquid chromatography
Procedure
- Comamonas sp. 7D-2 was inoculated in 100 ml LB medium (supplemented with 0.25 mM DBHB) and cultured at 30 °C for 14 h.
- Late exponential-phase cultures (approximately OD600=1.0) of strain 7D-2 were pelleted by centrifugation at 15,000 x g for 5 min, washed two times with PBS (pH 7.4), resuspended in PBS (containing 2 mM dithiothreitol) and then disrupted by sonication at 4 °C. BhbA was partially purified by fast protein liquid chromatography (Amersham Biosciences) at 4 °C (Chen et al., 2013; van de Pas et al., 1999).
- Protein concentrations were determined by the Bradford method (Bradford, 1976) with Protein Quantification Kit.
- A standard enzyme assay was performed under aerobic conditions in a 7 ml centrifuge tube containing 3.0 ml PBS, 0.1 mM DBHB (BHB) and 0.2 mM NADPH (NADH). The reaction was initiated by the addition of 30 μl of enzyme preparation and incubated at 30 °C for 60 min. Negative control without added enzyme was performed.
- The reaction was stopped by the addition of 15 μl sodium dithionite (1 M).
- The mixture was centrifuged at 16,000 x g for 10 min and filtered by membrane filtration; then, the DBHB concentration was analyzed using HPLC (Chen et al., 2013).
- One unit of BhbA activity was defined as the amount of BhbA that catalyzes the reduction of 1 nmol of DBHB per minute. Specific activity was expressed in units per milligram of protein. All assays were performed independently three times, and the means and standard deviations were calculated.
Recipes
- Phosphate buffered saline (PBS)
8.0 g NaCl
0.2 g KCl
1.44 g Na2HPO4
0.24 g KH2PO4 per liter of water
pH 7.4
- Mobile phase of HPLC
Acetonitrile: water: acetic acid (50: 50: 0.5/v:v:v)
Acknowledgments
This work was supported by grants from the Chinese National Science Foundation for Excellent Young Scholars (31222003), the Chinese National Natural Science Foundation (31070100), the Program for New Century Excellent Talents in University (NCET-12-0892), the Outstanding Youth Foundation of Jiangsu Province and the National Science and Technology Support Plan (2013AA102804). S. Zinder’s research on haloaromatic degradation was supported by funding by the DuPont Corporation.
References
- Bradford, M. M. (1976). A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72: 248-254.
- Chen, K., Huang, L., Xu, C., Liu, X., He, J., Zinder, S. H., Li, S. and Jiang, J. (2013). Molecular characterization of the enzymes involved in the degradation of a brominated aromatic herbicide. Mol Microbiol 89(6): 1121-1139.
- Sambrook, J., Russell, D. W. and Russell, D. W. (2001). Molecular cloning: a laboratory manual (3-volume set). Cold Spring Harbor Laboratory Press.
- van de Pas, B. A., Smidt, H., Hagen, W. R., van der Oost, J., Schraa, G., Stams, A. J. and de Vos, W. M. (1999). Purification and molecular characterization ofortho-chlorophenol reductive dehalogenase, a key rnzyme of halorespiration in Desulfitobacterium dehalogenans. J Biol Chem 274(29): 20287-20292.
Article Information
Copyright
© 2014 The Authors; exclusive licensee Bio-protocol LLC.
How to cite
Chen, K. and Jiang, J. (2014). The BhbA Enzyme Assay. Bio-protocol 4(8): e1111. DOI: 10.21769/BioProtoc.1111.
Category
Microbiology > Microbial biochemistry > Protein > Activity
Biochemistry > Protein > Activity
Do you have any questions about this protocol?
Post your question to gather feedback from the community. We will also invite the authors of this article to respond.
Share
Bluesky
X
Copy link










